Week 2 HW: DNA Read, Write & Edit
Lecture Prep
Professor Jacobson
DNA polymerase has a raw error rate of ~1:104 nucleotides, which improves to ~1:107 with proofreading. The human genome is ~3*109 base pairs, so many errors would occur per replication without correction. Biology resolves this through mismatch repair and other DNA repair pathways, reducing the final error rate.
An average human protein ~300 amino acids can be encoded by different DNA sequences due to genetic code degeneracy. In practice, most sequences do not work because of codon usage bias, mRNA secondary structure, GC content constraints, regulatory signal interference, effects on translation speed and protein folding, and cellular toxicity.
Dr. LeProust
The most commonly used method is solid-phase phosphoramidite chemical synthesis.
Each nucleotide-addition step is slightly imperfect, and these small errors accumulate over many cycles, leading to low yield and high error rates for long oligos.
At 2000bp, cumulative coupling inefficiency and chemical side reactions cause the full-length product yield to approach zero, making direct synthesis impractical; intead, genes are assembled from shorter oligos using enzymatic methods.
George Church
The 10 amino acids essential for all animals are methionine, threonine, tryptophan, phenylalanine, leucine, isoleucine, valine, histidine, arginine, and lysine (Berg, JM et al., 2019). Animals lack the metabolic pathways to synthesize these amino acids and must be taken through a dietary sources. This constraint underlies the concept of the “lysine contingency”, which highlights lysine as a particularly limiting amino acid in many cereal-based diets, since staple crops such as maize, rice, and wheat are deficient in lysine relative to animal nutritional requirements (Galili G, 2002). As a result, growth and health in animals can be constrained by lysine availability even when total protein intake is sufficient. The lysine contingency thus illustrates how molecular-level biochemical limitations can shape global food systems, ecological dependencies, and nutritional outcomes, and why biotechnological interventions that enhance lysine availability, such as microbial lysine production or high-lysine crops can have disproportionate impacts on food security and animal productivity.
This assignment was developed with the assistance of an AI language model (ChatGPT, OpenAI) for brainstorming, structuring responses, and editing for clarity.
Part 1: Benchling & In-silico Gel Art
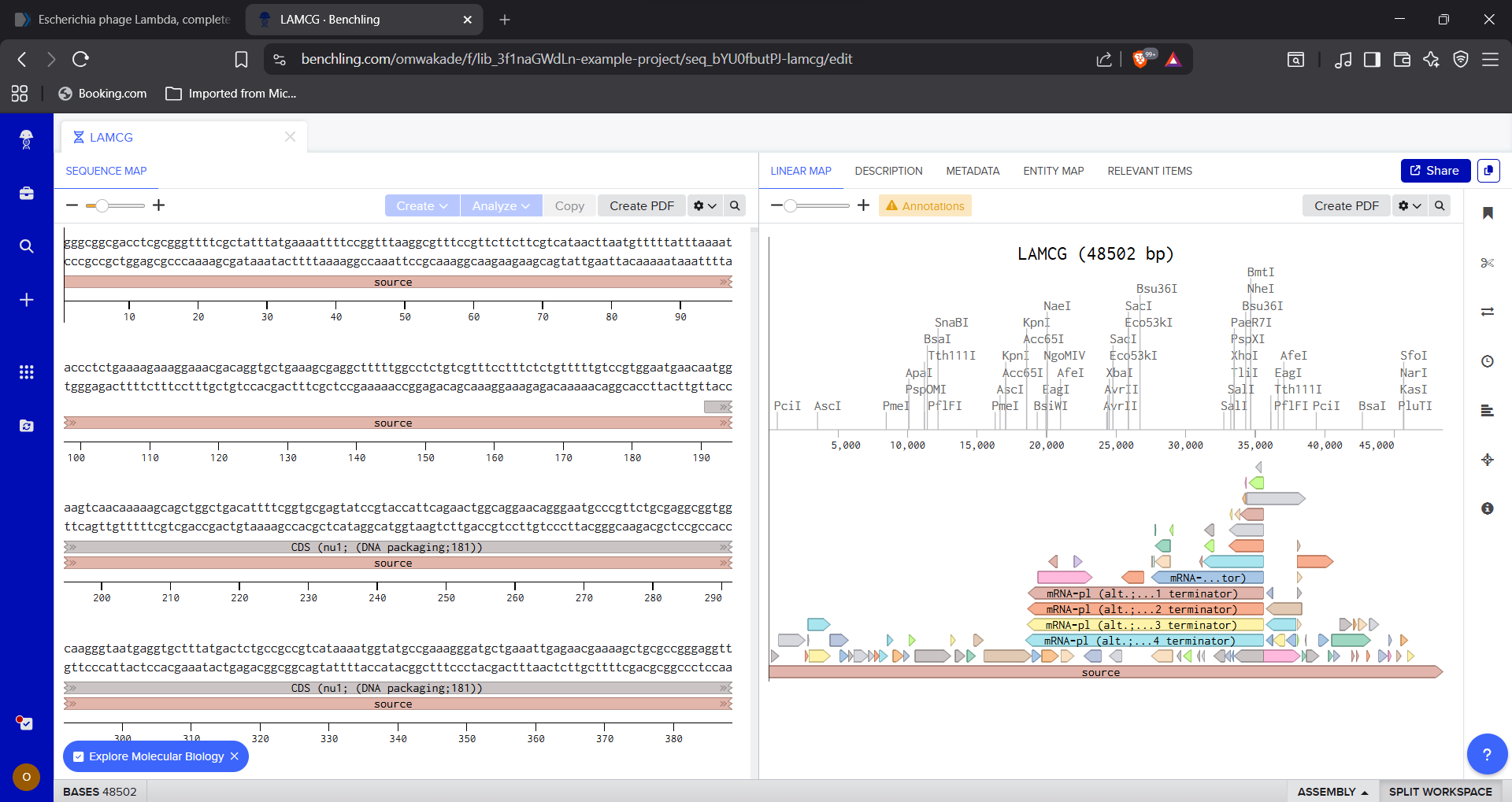
1.1 Importing E.coli phage Lambda Sequence in Benchling
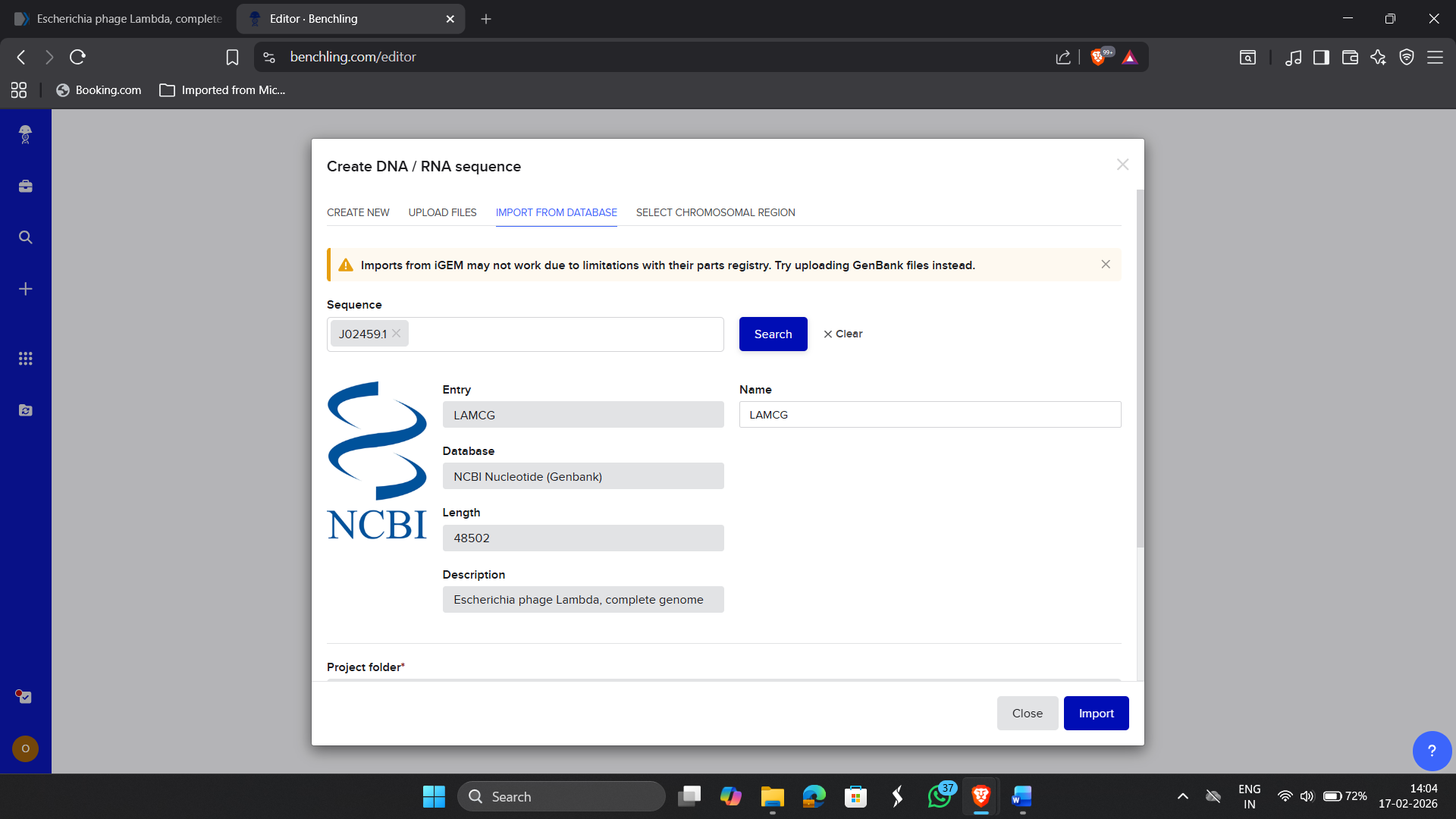
 | 
 |
|---|
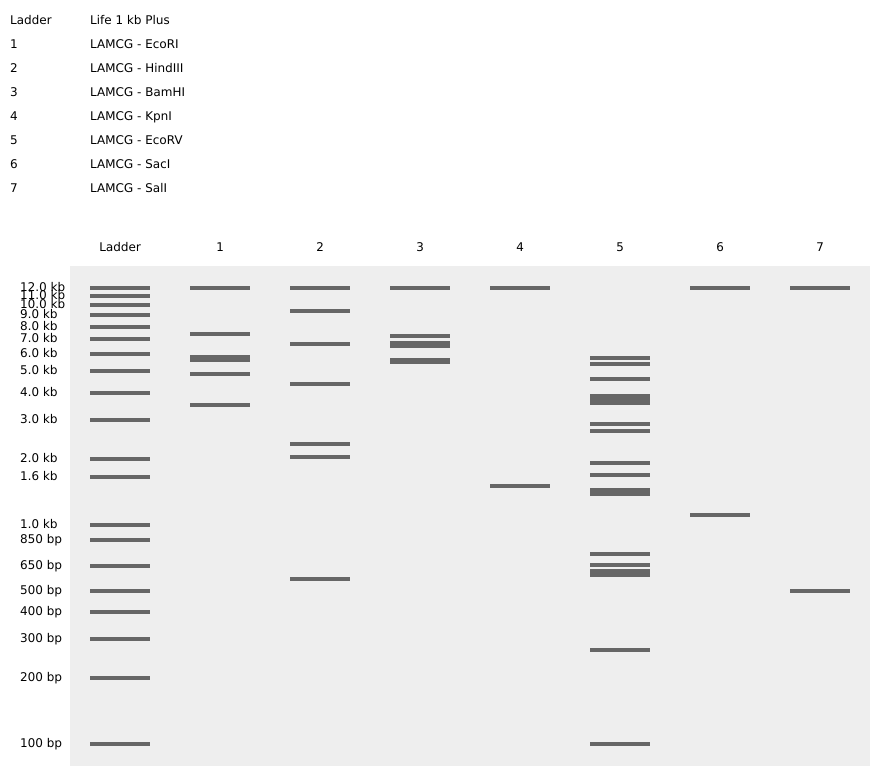
1.2 Restriction Enzyme Digestion
- Simulation of Restriction digestion with
- EcoRI
- HindIII
- BamHI
- KpnI
- EcoRV
- SacI
- SalI
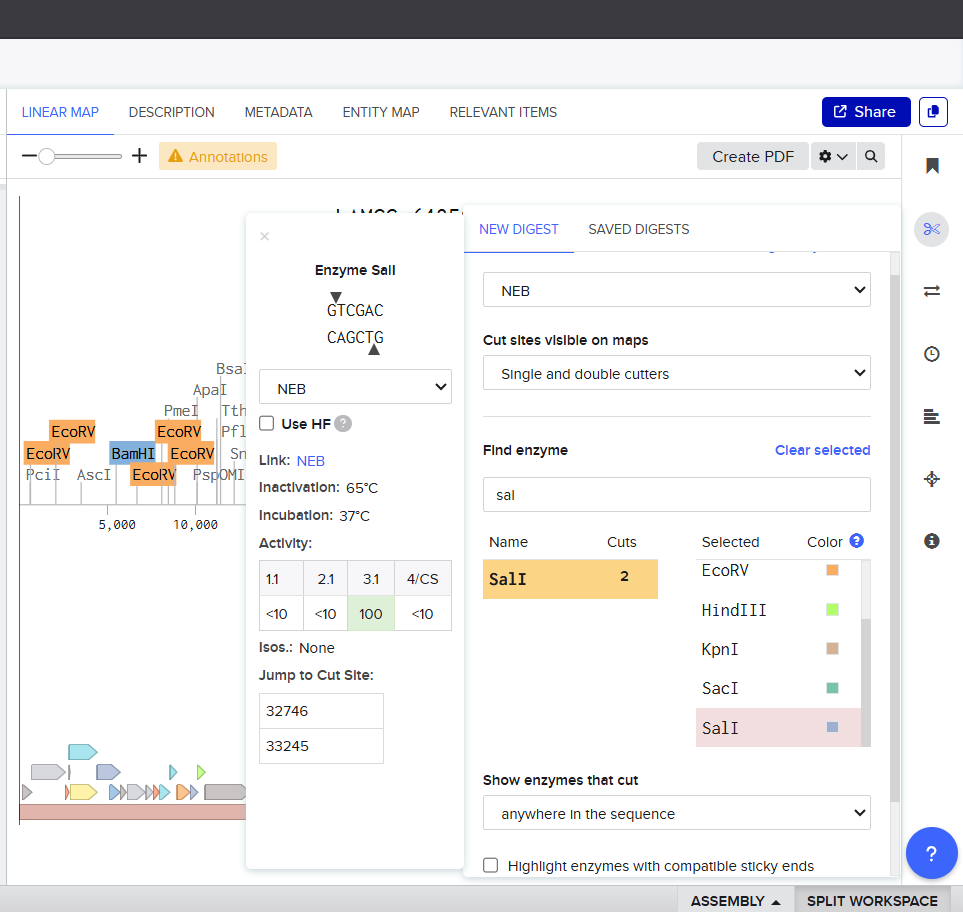

- Virtual Gel image


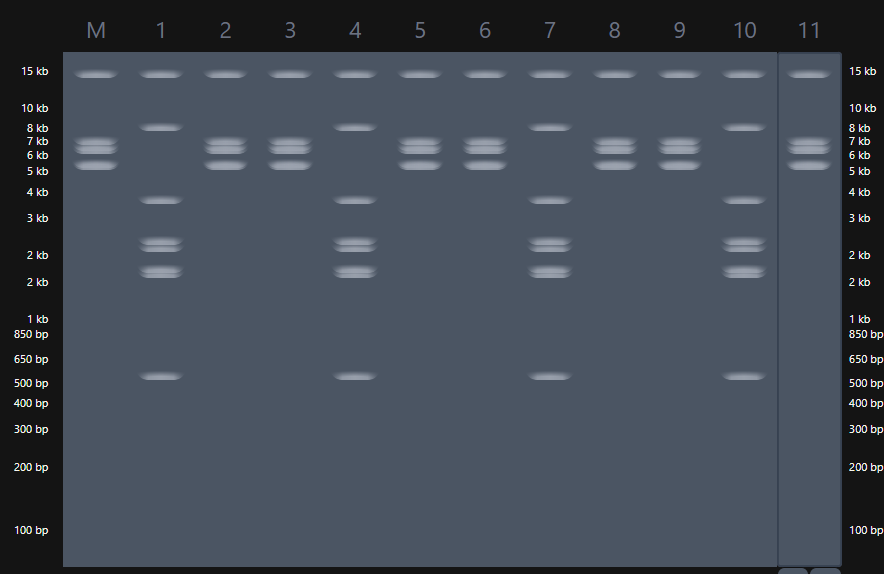
1.3 Gel image iteration using Ronan’s website

 | 
 | 
 |
|---|
1.4 Iteration of Classwork
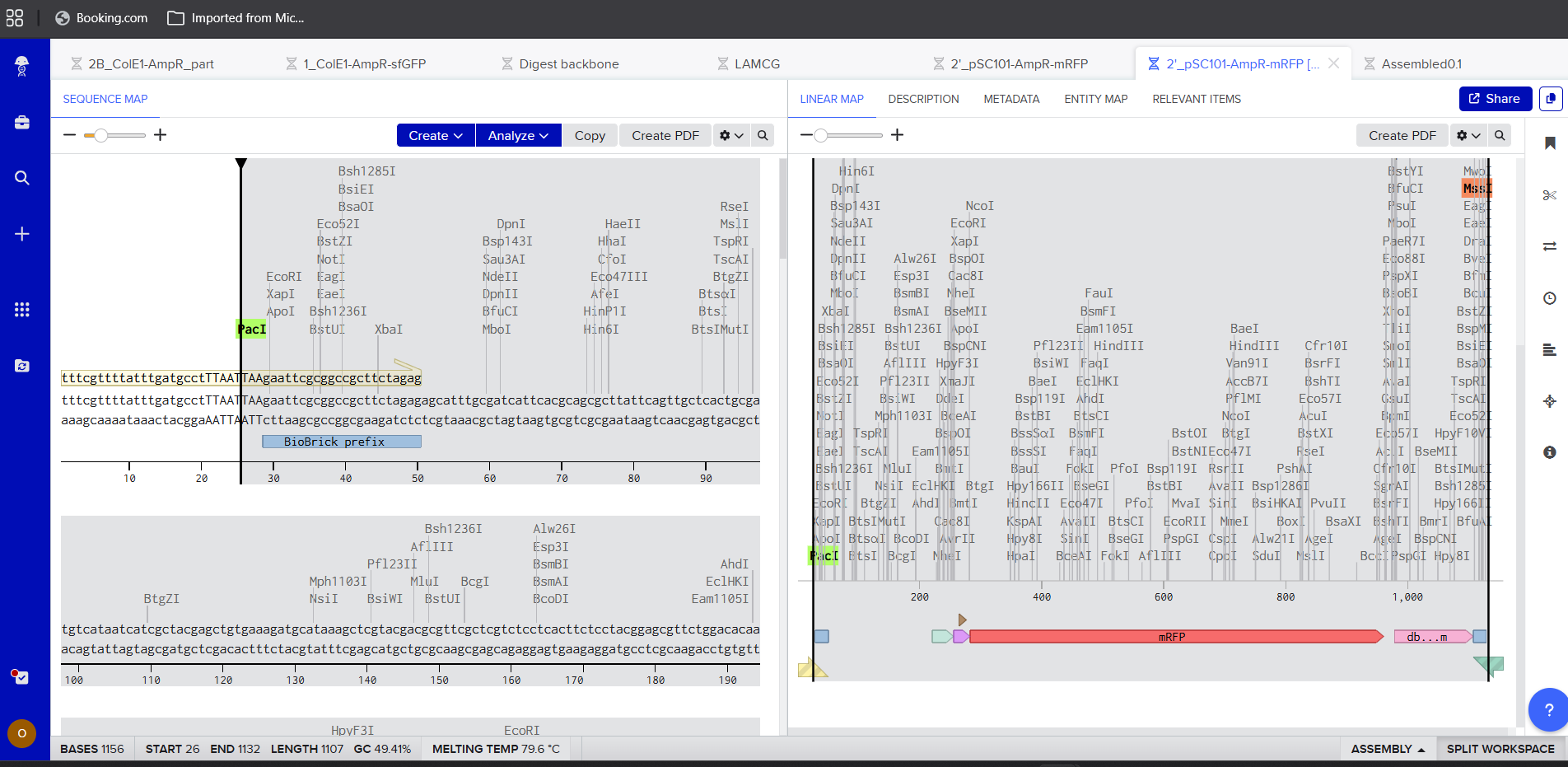
- PCR using Benchling
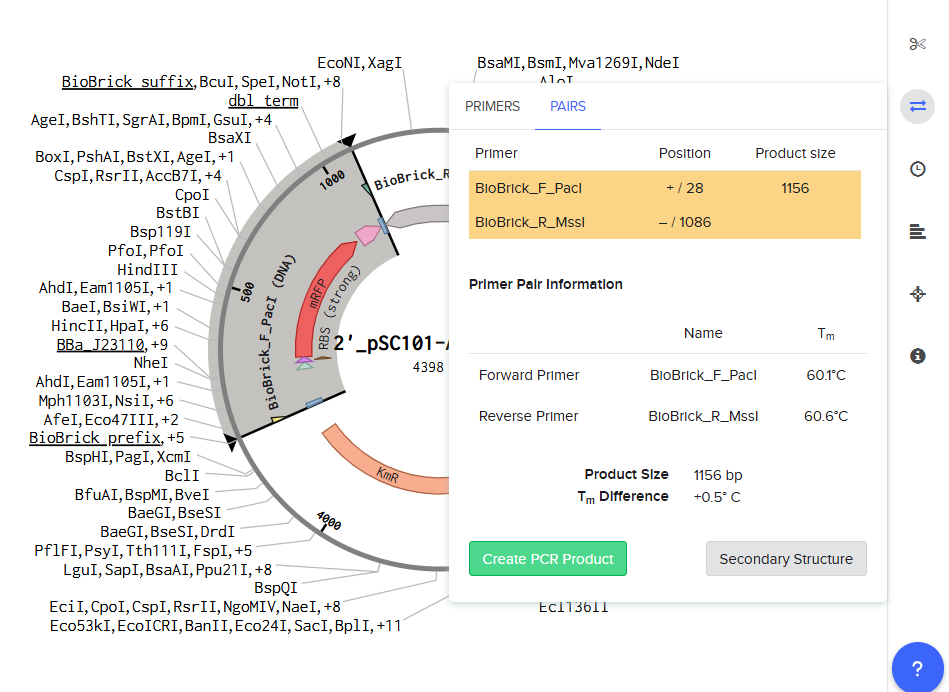
 | 
 |
|---|
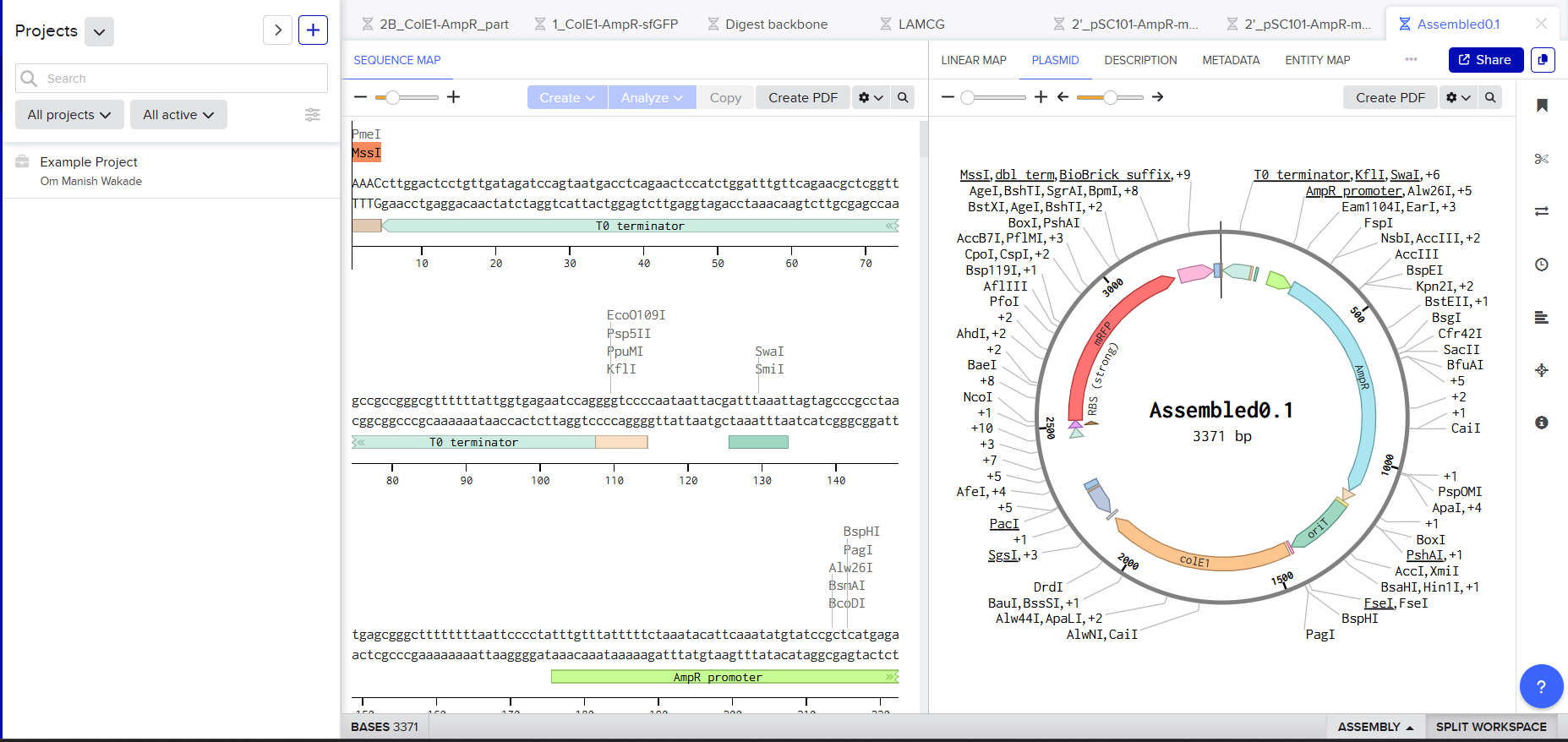
- Ligation using Benchling
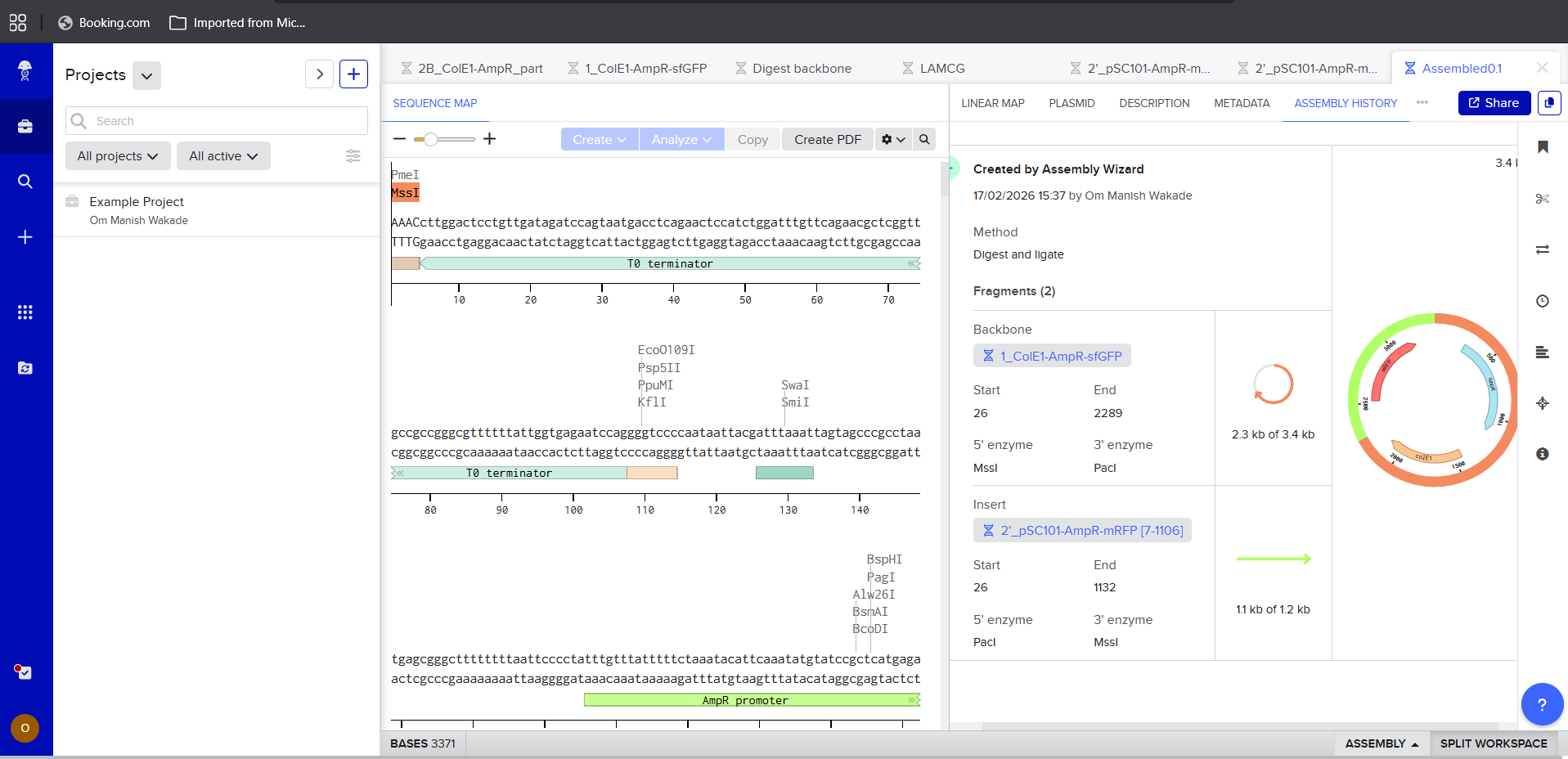
 | 
 |
|---|
Part 2: DNA Design Challenge
2.1 Protein Selection
As a protein Chitin Synthase 3 (CHS3) was chosen because it is responsible for synthesizing chitin, a major polysaccharide in fungal cell walls. Engineering bacteria to produce chitin has exciting applications in Biomaterial, Synthetic Biology, and Sustainable Biopolymer Production.
Amino acid sequence of CHS3 from UniProt
>sp|P29465|CHS3_YEAST Chitin synthase 3 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=CHS3 PE=1 SV=3 MTGLNGDDPDDYYLNLNQDEESLLRSRHSVGSGAPHRQGSLVRPERSRLNNPDNPHFYYA QKTQEQMNHLDVLPSSTGVNPNATRRSGSLRSKGSVRSKFSGRETDSYLLQDMNTTDKKA SVKISDEGVAEDEFDKDGDVDNFEESSTQPINKSIKPLRKETNDTLSFWQMYCYFITFWA PAPILAFCGMPKKERQMAWREKVALISVILYIGAIVAFLTFGFTKTVCSSSKLRLKNNEV STEFVVINGKAYELDTSSRSGIQDVEVDSDTLYGPWSDAGKDASFLFQNVNGNCHNLITP KSNSSIPHDDDNNLAWYFPCKLKNQDGSSKPNFTVENYAGWNCHTSKEDRDAFYGLKSKA DVYFTWDGIKNSSRNLIVYNGDVLDLDLLDWLEKDDVDYPVVFDDLKTSNLQGYDLSLVL SNGHERKIARCLSEIIKVGEVDSKTVGCIASDVVLYVSLVFILSVVIIKFIIACYFRWTV ARKQGAYIVDNKTMDKHTNDIEDWSNNIQTKAPLKEVDPHLRPKKYSKKSLGHKRASTFD LLKKHSSKMFQFNESVIDLDTSMSSSLQSSGSYRGMTTMTTQNAWKLSNENKAVHSRNPS TLLPTSSMFWNKATSSPVPGSSLIQSLDSTIIHPDIVQQPPLDFMPYGFPLIHTICFVTC YSEDEEGLRTTLDSLSTTDYPNSHKLLMVVCDGLIKGSGNDKTTPEIALGMMDDFVTPPD EVKPYSYVAVASGSKRHNMAKIYAGFYKYDDSTIPPENQQRVPIITIVKCGTPAEQGAAK PGNRGKRDSQIILMSFLEKITFDERMTQLEFQLLKNIWQITGLMADFYETVLMVDADTKV FPDALTHMVAEMVKDPLIMGLCGETKIANKAQSWVTAIQVFEYYISHHQAKAFESVFGSV TCLPGCFSMYRIKSPKGSDGYWVPVLANPDIVERYSDNVTNTLHKKNLLLLGEDRFLSSL MLKTFPKRKQVFVPKAACKTIAPDKFKVLLSQRRRWINSTVHNLFELVLIRDLCGTFCFS MQFVIGIELIGTMVLPLAICFTIYVIIFAIVSKPTPVITLVLLAIILGLPGLIVVITATR WSYLWWMCVYICALPIWNFVLPSYAYWKFDDFSWGDTRTIAGGNKKAQDENEGEFDHSKI KMRTWREFEREDILNRKEESDSFVA
2.2 Reverse Translate: Protein Sequence –> DNA Sequence
DNA sequence was obtained using NCBI for CHS3 protein.
Nucleotide sequence of CHS3 from NCBI
lcl|NM_001178371.1_cds_NP_009579.1_1 [gene=CHS3] [locus_tag=YBR023C] [db_xref=SGD:S000000227,GeneID:852311] [protein=chitin synthase CHS3] [protein_id=NP_009579.1] [location=1..3498] [gbkey=CDS] ATGACCGGCTTGAATGGAGATGATCCTGATGACTACTATCTGAACCTTAATCAAGATGAAGAGTCTCTACTTAGGTCAAGACACAGTGTCGGCTCAGGAGCACCTCATAGACAAGGCTCTTTAGTGCGGCCCGAAAGAAGCCGACTGAACAATCCTGATAATCCACATTTTTATTATGCGCAGAAAACGCAGGAGCAGATGAATCACCTGGATGTTTTACCATCAAGTACCGGTGTAAACCCAAATGCAACTCGTCGGAGTGGCTCCCTGCGCTCCAAAGGCTCAGTGAGAAGCAAATTTAGTGGCCGCGAAACGGATAGCTATCTTTTACAAGATATGAATACTACTGACAAGAAGGCTTCCGTTAAAATAAGTGATGAAGGTGTTGCGGAAGACGAATTTGATAAAGATGGTGATGTGGACAATTTCGAAGAAAGCTCCACGCAGCCCATAAATAAGTCTATCAAACCATTAAGAAAGGAAACGAATGATACATTGTCATTTTGGCAGATGTACTGTTATTTCATTACGTTTTGGGCACCTGCTCCAATTCTTGCTTTCTGCGGGATGCCAAAGAAGGAAAGACAAATGGCGTGGAGAGAAAAGGTTGCTTTAATTTCTGTCATCTTGTACATTGGTGCGATTGTGGCTTTCCTGACTTTTGGTTTCACTAAAACCGTTTGTAGTAGTTCGAAACTACGTTTGAAAAACAACGAAGTATCAACAGAATTTGTCGTAATTAACGGTAAGGCTTATGAATTGGATACTTCCTCGCGTTCCGGTATACAAGACGTTGAAGTAGATTCAGACACCCTTTATGGGCCCTGGTCAGATGCTGGTAAAGATGCTTCGTTCTTGTTTCAAAATGTGAATGGTAACTGTCATAACCTTATAACTCCAAAGAGTAATTCTTCCATTCCCCATGACGATGATAATAATTTAGCATGGTATTTTCCTTGTAAGTTAAAGAATCAAGATGGCTCTTCGAAGCCAAACTTCACAGTTGAAAATTACGCAGGATGGAACTGTCATACGTCTAAAGAAGATAGGGACGCATTTTACGGTTTAAAGTCGAAAGCTGATGTTTACTTCACTTGGGATGGTATAAAGAACTCTTCTAGAAACTTGATTGTTTATAATGGCGACGTTTTGGATTTAGATCTTCTTGATTGGTTGGAAAAGGATGACGTTGACTATCCCGTTGTATTCGATGACTTGAAGACTTCAAATTTACAAGGTTATGATCTTTCGTTGGTTTTGTCAAATGGGCATGAAAGAAAAATTGCGAGATGTTTGAGCGAAATTATTAAAGTTGGTGAAGTAGACTCCAAAACCGTCGGTTGTATTGCCTCTGATGTCGTTTTGTATGTTTCTCTGGTATTTATTCTTTCAGTGGTGATAATTAAATTCATAATTGCCTGCTACTTCCGTTGGACTGTAGCTAGGAAACAAGGTGCATATATCGTGGACAATAAAACAATGGATAAACACACAAACGATATCGAGGATTGGTCTAATAATATTCAAACAAAAGCTCCTCTAAAGGAAGTAGATCCTCATTTGAGGCCAAAGAAATACTCAAAAAAGTCGTTGGGACACAAGCGTGCTTCAACCTTTGACTTGCTGAAAAAACACAGCTCCAAAATGTTTCAATTTAACGAATCTGTGATAGATCTAGACACCTCCATGAGCAGTTCACTACAATCTTCTGGTTCATACAGAGGAATGACAACAATGACCACTCAAAATGCTTGGAAACTCTCGAATGAAAACAAAGCTGTACATTCCCGTAATCCATCTACTTTGTTGCCTACATCCTCGATGTTTTGGAATAAAGCGACTTCCTCTCCTGTACCAGGATCATCGCTGATTCAGAGTCTTGATTCGACGATTATACATCCCGATATCGTTCAACAACCACCACTGGATTTTATGCCATACGGGTTCCCATTGATTCATACTATCTGTTTTGTTACTTGTTATTCTGAGGATGAAGAGGGTTTAAGAACCACTTTAGACTCTCTTTCTACCACAGATTATCCAAATTCCCATAAACTACTGATGGTTGTTTGTGATGGTTTAATTAAGGGCTCGGGCAACGATAAGACTACTCCAGAGATAGCGTTAGGAATGATGGACGACTTTGTCACCCCACCTGATGAAGTTAAACCTTACTCCTATGTGGCAGTGGCATCAGGCTCTAAAAGACACAATATGGCCAAGATATATGCGGGTTTTTACAAATATGACGATTCTACAATTCCACCAGAAAATCAACAACGTGTCCCAATCATTACAATTGTGAAGTGCGGTACTCCTGCAGAGCAGGGGGCCGCCAAACCCGGTAACAGAGGTAAGCGTGATTCTCAAATTATTCTGATGTCCTTTTTAGAAAAAATAACATTTGATGAAAGAATGACTCAATTGGAATTTCAGCTTTTAAAAAATATTTGGCAGATTACGGGGCTAATGGCAGACTTCTACGAAACGGTACTTATGGTTGATGCTGATACTAAAGTCTTTCCCGATGCTTTAACTCATATGGTCGCTGAAATGGTTAAAGATCCTTTGATTATGGGTCTTTGTGGTGAGACCAAGATCGCTAATAAGGCACAATCTTGGGTAACTGCAATTCAAGTGTTTGAGTACTATATTTCGCATCATCAGGCTAAAGCTTTTGAATCTGTCTTCGGTTCGGTAACTTGTTTGCCGGGATGTTTCTCAATGTATCGTATAAAATCTCCTAAAGGTTCAGATGGTTATTGGGTACCTGTATTGGCAAATCCAGATATTGTTGAAAGATATTCGGATAATGTTACAAACACTTTGCATAAGAAGAACTTATTATTACTTGGTGAAGATAGATTTTTATCTTCATTAATGTTAAAGACTTTCCCTAAGAGAAAGCAAGTATTTGTTCCAAAAGCTGCTTGTAAAACTATTGCCCCTGATAAATTCAAAGTCTTACTTTCCCAGCGTCGAAGATGGATTAATTCTACGGTACATAACCTTTTTGAATTAGTTCTAATCAGAGACTTATGTGGCACTTTCTGTTTTTCCATGCAATTTGTGATTGGTATTGAATTGATTGGTACTATGGTACTGCCGTTAGCCATTTGCTTTACTATTTATGTCATTATTTTTGCCATTGTATCAAAACCTACACCCGTAATCACTTTAGTTTTACTGGCAATTATTCTTGGTCTGCCCGGCTTAATTGTTGTTATAACTGCTACGAGATGGTCGTACCTATGGTGGATGTGCGTATATATTTGTGCTTTGCCTATTTGGAATTTCGTACTACCTTCATATGCGTACTGGAAATTTGATGACTTCTCATGGGGTGATACGAGAACTATTGCGGGAGGTAATAAAAAGGCACAAGACGAGAATGAAGGTGAATTTGATCACTCAAAGATTAAAATGAGGACATGGAGGGAATTTGAAAGGGAAGATATTCTCAATCGGAAGGAGGAAAGTGACTCCTTCGTTGCATAG
2.3 Codon optimization
Codon optimization using Vector Builder
ATGACCGGCCTGAATGGCGATGATCCGGATGACTATTATCTGAACCTGAACCAGGATGAAGAAAGCCTGCTGCGCAGCCGCCACAGCGTGGGTAGCGGCGCGCCGCATCGTCAGGGCAGCCTGGTGCGTCCGGAACGCAGCCGCTTAAACAACCCGGATAATCCGCATTTTTACTACGCGCAGAAAACGCAGGAACAGATGAACCACCTGGATGTGCTGCCGAGCAGCACCGGCGTGAACCCAAATGCCACACGTCGCAGTGGCAGCCTGCGCAGTAAAGGATCAGTTCGCAGCAAATTTAGCGGCCGTGAAACCGATAGCTACCTGCTGCAGGATATGAACACCACCGATAAAAAAGCCAGCGTGAAAATTAGCGACGAAGGCGTGGCGGAAGATGAATTTGATAAAGATGGCGACGTGGATAACTTTGAGGAAAGCAGCACCCAGCCGATTAATAAAAGCATTAAACCGCTGCGTAAAGAAACCAATGATACCCTGTCCTTTTGGCAGATGTATTGCTACTTTATTACCTTTTGGGCCCCGGCCCCGATCCTGGCGTTTTGCGGCATGCCGAAAAAGGAACGCCAGATGGCCTGGCGCGAAAAAGTCGCGCTGATTAGCGTGATTCTGTATATTGGCGCCATTGTGGCCTTTCTGACCTTTGGTTTTACGAAAACCGTGTGTAGCAGCAGCAAACTGCGCCTGAAAAATAACGAAGTGAGCACCGAATTTGTGGTCATTAATGGCAAAGCGTATGAACTGGATACCAGCAGCCGCAGCGGCATTCAGGATGTGGAAGTGGATAGCGATACCCTGTATGGCCCGTGGAGCGACGCGGGCAAAGATGCCAGCTTCCTGTTTCAGAATGTGAATGGCAACTGCCATAACCTGATTACCCCTAAAAGCAACAGCAGCATTCCGCATGATGACGATAACAATCTCGCGTGGTATTTTCCGTGTAAACTGAAAAACCAGGACGGATCGAGCAAACCGAACTTTACCGTGGAAAACTACGCCGGCTGGAATTGCCACACCAGCAAAGAAGACCGTGATGCGTTTTATGGTCTGAAAAGCAAAGCCGATGTGTACTTTACCTGGGATGGTATTAAAAACAGCTCGCGCAACCTGATTGTGTATAATGGCGATGTGCTGGACCTGGATCTGCTCGATTGGCTGGAAAAAGACGATGTAGATTATCCGGTGGTGTTCGACGATCTGAAAACCAGCAACCTGCAGGGCTACGATCTGAGCCTGGTGCTGAGCAACGGCCATGAACGCAAAATCGCCCGCTGCCTGAGCGAAATCATTAAAGTGGGCGAAGTTGATAGTAAAACCGTCGGCTGCATTGCCTCTGATGTGGTGCTGTATGTGAGCCTGGTGTTCATTCTGAGCGTTGTGATTATCAAATTCATCATCGCGTGCTATTTTCGCTGGACCGTGGCGCGTAAACAGGGCGCCTATATTGTGGATAATAAAACCATGGATAAACATACCAACGATATTGAAGATTGGAGCAATAATATTCAGACCAAAGCGCCGCTGAAAGAAGTGGATCCGCACCTGCGCCCGAAAAAATATAGCAAAAAAAGCCTGGGCCATAAACGCGCCAGCACCTTTGATCTGCTGAAAAAACACAGTAGCAAAATGTTTCAGTTTAATGAAAGCGTTATTGATCTGGACACGAGCATGAGCAGCAGCCTGCAAAGCAGCGGTAGCTATCGCGGTATGACCACCATGACCACCCAGAACGCCTGGAAACTGAGCAACGAAAATAAAGCGGTGCATAGCCGCAATCCGAGCACCCTGCTGCCGACCAGCAGCATGTTTTGGAATAAAGCCACCAGCAGCCCGGTTCCGGGCAGCAGCCTGATCCAGAGCCTGGATAGCACCATTATTCACCCGGATATTGTGCAGCAACCGCCGCTGGATTTTATGCCGTATGGTTTCCCGCTGATTCATACCATTTGCTTCGTGACCTGCTACAGCGAAGATGAGGAGGGCCTGCGCACCACCCTGGATAGCCTGTCTACGACCGATTACCCGAATAGCCACAAACTGCTGATGGTGGTGTGCGATGGTCTGATTAAAGGCAGCGGCAACGATAAAACCACCCCGGAAATTGCACTGGGCATGATGGATGATTTCGTGACCCCGCCGGATGAAGTTAAACCGTATAGCTATGTTGCCGTTGCAAGCGGCAGCAAACGCCATAACATGGCCAAAATCTATGCGGGCTTCTACAAATACGATGATTCAACCATTCCGCCGGAAAACCAGCAGCGCGTGCCGATCATTACCATTGTGAAATGCGGCACCCCGGCGGAACAGGGCGCCGCGAAACCGGGCAATCGCGGTAAACGCGATAGCCAGATCATTCTGATGAGCTTTCTGGAAAAAATCACCTTTGATGAACGTATGACCCAGCTGGAATTTCAGTTACTGAAGAACATTTGGCAGATTACGGGCCTGATGGCGGATTTCTATGAAACGGTGCTGATGGTTGATGCGGATACCAAAGTGTTCCCGGATGCCCTGACCCATATGGTGGCCGAAATGGTGAAGGATCCGCTGATTATGGGCCTGTGCGGCGAAACCAAAATTGCCAATAAAGCGCAGAGCTGGGTCACCGCGATTCAGGTGTTTGAATACTATATCAGCCATCATCAGGCGAAAGCGTTTGAAAGCGTGTTTGGCAGCGTTACCTGCCTGCCGGGCTGCTTTTCAATGTACCGTATCAAAAGCCCGAAAGGCAGCGATGGCTATTGGGTGCCGGTGCTGGCCAATCCGGATATTGTCGAACGTTATAGCGATAATGTGACCAACACCCTGCATAAAAAAAACCTGCTGCTGTTGGGCGAAGATCGCTTCCTGAGCTCACTGATGCTGAAAACCTTTCCGAAACGCAAACAGGTTTTTGTGCCGAAAGCGGCGTGTAAAACCATTGCCCCGGATAAATTTAAAGTGCTGCTGAGCCAGCGCCGTCGCTGGATCAACAGCACCGTGCATAATCTGTTTGAACTGGTGCTGATTCGTGATCTGTGTGGCACCTTCTGCTTTAGCATGCAGTTCGTGATTGGCATTGAACTGATTGGCACCATGGTTCTGCCGCTGGCGATTTGCTTTACCATTTACGTGATTATTTTTGCGATTGTGTCAAAACCGACCCCGGTCATTACCCTGGTCCTGCTGGCGATTATTCTGGGCCTGCCGGGCCTGATTGTGGTGATTACAGCCACCCGCTGGAGTTACCTGTGGTGGATGTGCGTGTATATTTGCGCGCTGCCGATTTGGAACTTTGTGCTGCCGTCGTATGCGTATTGGAAATTTGATGACTTCTCATGGGGCGACACCCGCACCATTGCGGGCGGCAACAAAAAAGCGCAGGATGAAAACGAAGGCGAATTTGATCATAGCAAAATTAAAATGCGCACGTGGCGTGAATTTGAACGTGAAGATATTCTGAATCGCAAAGAAGAAAGCGATAGCTTTGTGGCCTAA
2.4 You have a Sequence! Now What?
The optimized DNA sequence can be synthesized by cloning into expression vector pET-28a(+)
Cell-Dependent Expression in E. coli
- Insert gene into plasmid under T7 promoter.
- Transform into BL21(DE3).
- Add IPTG to induce T7 RNA polymerase.
- DNA –> mRNA (Transcription).
- mRNA –> Protein (Translation at ribosome).
- Chitin syntase produced in cytoplasm or membrane
Central Dogma Flow
graph LR; DNA(CHS3)-->mRNA-->ChitinSyntheseProteins
Part 3: Prepare a Twist DNA Synthesis Order