Week 02 Homework: DNA Read, Write and Edit
Part 1: Benchling & In-silico Gel Art


Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
No lab access
Part 3: DNA Design Challenge


3.1. Choose your protein.


I chose Green Fluorescent Protein (GFP) because it is widely used in biotechnology as a visual reporter protein. Its ability to fluoresce green when exposed to blue light makes it an elegant example of how DNA sequences can encode observable biological functions. This is a random choice, I love green and this is HTGAA!


The 2008 Nobel Prize in Chemistry was awarded to Osamu Shimomura, Martin Chalfie, and Roger Y. Tsien for the discovery and development of Green Fluorescent Protein (GFP)
Protein Sequence: P42212|GFP_AEQVI Green fluorescent protein OS=Aequorea victoria MSKGEELFTGVVPILVELDGDVNGHKFSVSGEGEGDATYGKLTLKFICTTGKLPVPWPTLVTTLTYGVQCFSRYPDHMKQHDFFKSAMPEGYVQERTIFFKDDGNYKTRAEVKFEGDTLVNRIELKGIDFKEDGNILGHKLEYNYNSHNVYIMADKQKNGIKVNFKIRHNIEDGSVQLADHYQQNTPIGDGPVLLPDNHYLSTQSALSKDPNEKRDHMVLLEFVTAAGITHGMDELYK
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence. DNA Nucleotide Sequence 1 tacacacgaa taaaagataa caaagatgag taaaggagaa gaacttttca ctggagttgt 61 cccaattctt gttgaattag atggtgatgt taatgggcac aaattttctg tcagtggaga 121 gggtgaaggt gatgcaacat acggaaaact tacccttaaa tttatttgca ctactggaaa 181 actacctgtt ccatggccaa cacttgtcac tactttctct tatggtgttc aatgcttttc 241 aagataccca gatcatatga aacagcatga ctttttcaag agtgccatgc ccgaaggtta 301 tgtacaggaa agaactatat ttttcaaaga tgacgggaac tacaagacac gtgctgaagt 361 caagtttgaa ggtgataccc ttgttaatag aatcgagtta aaaggtattg attttaaaga 421 agatggaaac attcttggac acaaattgga atacaactat aactcacaca atgtatacat 481 catggcagac aaacaaaaga atggaatcaa agttaacttc aaaattagac acaacattga 541 agatggaagc gttcaactag cagaccatta tcaacaaaat actccaattg gcgatggccc 601 tgtcctttta ccagacaacc attacctgtc cacacaatct gccctttcga aagatcccaa 661 cgaaaagaga gaccacatgg tccttcttga gtttgtaaca gctgctggga ttacacatgg 721 catggatgaa ctatacaaat aaatgtccag acttccaatt gacactaaag tgtccgaaca 781 attactaaaa tctcagggtt cctggttaaa ttcaggctga gatattattt atatatttat 841 agattcatta aaattgtatg aataatttat tgatgttatt gatagaggtt attttcttat 901 taaacaggct acttggagtg tattcttaat tctatattaa ttacaatttg atttgacttg 961 ctcaaa
3.3. Codon optimization. ATGGTCTCAAAAGGTGAAGAATTGTTTACAGGTGTCGTACCTATACTTGTAGAACTCGATGGTGATGTTAATGGTCATAAATTTTCGGTCTCAGGAGAAGGTGAAGGAGACGCGACTTATGGTAAACTCACTTTAAAATTCATATGTACAACTGGTAAATTACCTGTTCCATGGCCGACTTTAGTGACAACGTTGACGTATGGTGTTCAATGTTTTAGTCGTTATCCTGATCATATGAAACAACATGATTTCTTTAAAAGTGCAATGCCTGAGGGTTATGTTCAAGAACGGACGATTTTCTTTAAAGATGATGGGAATTACAAAACTCGCGCAGAAGTCAAATTTGAAGGAGACACACTGGTAAATCGTATAGAACTTAAAGGTATTGACTTTAAAGAAGATGGAAATATTTTAGGTCATAAACTTGAATACAATTTGAACTCCCATAATGTCTACATAATGGCAGACAAACAGAAGAATGGAATAAAAGTTAATTTTAAAATACGCCATAATATTGAAGATGGTTCGGTCCAACTGGCAGATCATTATCAACAGAATACTCCAATTGGAGATGGTCCAGTCTTGTTACCAGATAATCATTATCTCAGTACTCAATCAGCGCTCTCTAAGGATCCAAATGAAAAGCGTGACCATATGGTGTTGCTCGAATTTGTTACAGCGGCAGGCATTACATTAGGAATGGATGAATTATATAAATAG
3.4. You have a sequence! Now what? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both. I read about how fluorescent green protein is used in molecular biology to identify and track protein movement and gene expression. I am not yet ready to describe how a DNA sequence can be transcribed and translated into this protein. The best I can do here “in my own words” is to go back to the central dogma / flow of genetic information: DNA >RNA> Protein This is what I think happened in this week’s HW
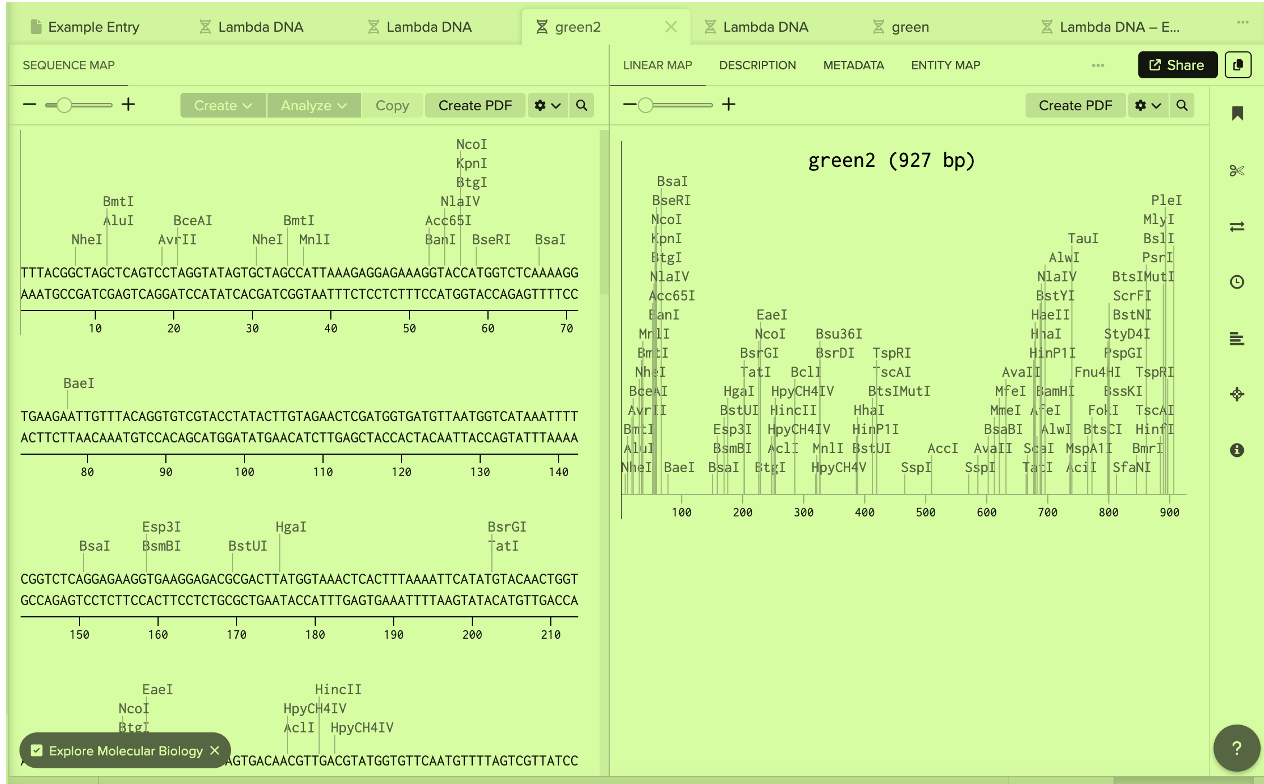

Part 4: Prepare a Twist DNA Synthesis Order: Design the full machine (Expression Cassette?) that makes bacteria glow.
4.1. Create a Twist account and a Benchling account
4.2. Build Your DNA Insert Sequence Let’s make a sequence that will make E. coli glow fluorescent green under UV light by constitutively (always) expressing sfGFP (a green fluorescent protein):
Promoter TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGC RBS CATTAAAGAGGAGAAAGGTACC Start Codon ATG Coding Sequence GTCTCAAAAGGTGAAGAATTGTTTACAGGTGTCGTACCTATACTTGTAGAACTCGATGGTGATGTTAATGGTCATAAATTTTCGGTCTCAGGAGAAGGTGAAGGAGACGCGACTTATGGTAAACTCACTTTAAAATTCATATGTACAACTGGTAAATTACCTGTTCCATGGCCGACTTTAGTGACAACGTTGACGTATGGTGTTCAATGTTTTAGTCGTTATCCTGATCATATGAAACAACATGATTTCTTTAAAAGTGCAATGCCTGAGGGTTATGTTCAAGAACGGACGATTTTCTTTAAAGATGATGGGAATTACAAAACTCGCGCAGAAGTCAAATTTGAAGGAGACACACTGGTAAATCGTATAGAACTTAAAGGTATTGACTTTAAAGAAGATGGAAATATTTTAGGTCATAAACTTGAATACAATTTGAACTCCCATAATGTCTACATAATGGCAGACAAACAGAAGAATGGAATAAAAGTTAATTTTAAAATACGCCATAATATTGAAGATGGTTCGGTCCAACTGGCAGATCATTATCAACAGAATACTCCAATTGGAGATGGTCCAGTCTTGTTACCAGATAATCATTATCTCAGTACTCAATCAGCGCTCTCTAAGGATCCAAATGAAAAGCGTGACCATATGGTGTTGCTCGAATTTGTTACAGCGGCAGGCATTACATTAGGAATGGATGAATTATATAAA 7x His Tag CATCACCATCACCATCATCAC Stop Codon TAA Terminator CCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA
4.3. On Twist, Select The “Genes” Option 4.4. Select “Clonal Genes” option 4.5. Import your sequence I had to stop here because Twist would not accept my .fasta file. When I loaded it kept adding.txt 4.6. Choose Your Vector No can do
Part 5: DNA Read/Write/Edit
5.1 DNA Read


What DNA would you want to sequence (e.g., read) and why?
The Smell of Renewal: mapping post-fire soil chemistry + culture as a recovery marker
I would want to sequence DNA from soil bacteria called Streptomyces that produce geosmin, a key molecule behind petrichor (the earthy smell after rain). I’m interested in this because after a major fire, soil ecosystems change dramatically, and I want to understand what microbial communities survive, return, or disappear during recovery.
Why sequence it? The big “why” is that my home burned down in the Palisades fire last year. As I recover from the trauma of the fire, I find myself deeply drawn to the repair and restoration of my neighborhood — both physically and emotionally. I am particularly interested in sequencing soil bacteria such as Streptomyces, which produce geosmin, a key molecule behind petrichor — the earthy smell after rain.
Sequencing would allow me to identify which Streptomyces species or strains are present in post-fire soil, compare them to soil from unaffected areas, and observe how the microbial “signature” of a burned landscape changes over time. This could support environmental monitoring by tracking soil recovery and ecosystem health after disaster.
Because petrichor is strongly tied to emotional memory and renewal, understanding the biology behind it could connect ecological recovery with human recovery after disaster.
(CHAT GPT: How would I do this?)To study these microbial communities, I would use 16S rRNA gene sequencing, a common method for identifying and comparing bacterial species in environmental samples. By extracting DNA from soil and sequencing this conserved bacterial marker gene, I could determine which Streptomyces strains are present and how their abundance changes over time following a fire.
What I learned this week
- Understood the Central Dogma in a functional way.
- Learned what promoters and RBS actually do.
- Codon-optimized a gene.
- Wrestled with file formats (real-world friction).
- Designed a sequencing project grounded in lived experience.
- Named 16S rRNA as a method.
- Connected six diverse interdisciplinary areas of inquiry
Fire Soil Microbes Memory Recovery Design Biology