Week 02 HW: DNA READ, WRITE AND EDIT
Part 1: Benchling & In-silico Gel Art
This DNA gel art was designed in the style of Paul Vanouse’s Latent Figure Protocol. I chose to create the letter “P” as it is the initial of my name, Paula. To achieve this, I used Ronan’s website, which was a helpful tool for quickly iterating on the designs and determining the best enzyme combinations to form the silhouette of the letter.


Part 3: DNA Design Challenge
3.1 Choose your protein
Hydrophobin HFBI de Trichoderma reesei: I chose this protein because I will be participating in a summer research program at Aalto University focused on bio-based foams and mycelium-derived materials. Hydrophobins are proteins naturally produced by fungi and play an important role in fungal growth, particularly in modifying surface properties and mediating interactions at air–water interfaces. These characteristics are directly relevant to mycelium-based biomaterials, where fungal networks interact with substrates to form structured materials with tunable mechanical properties.
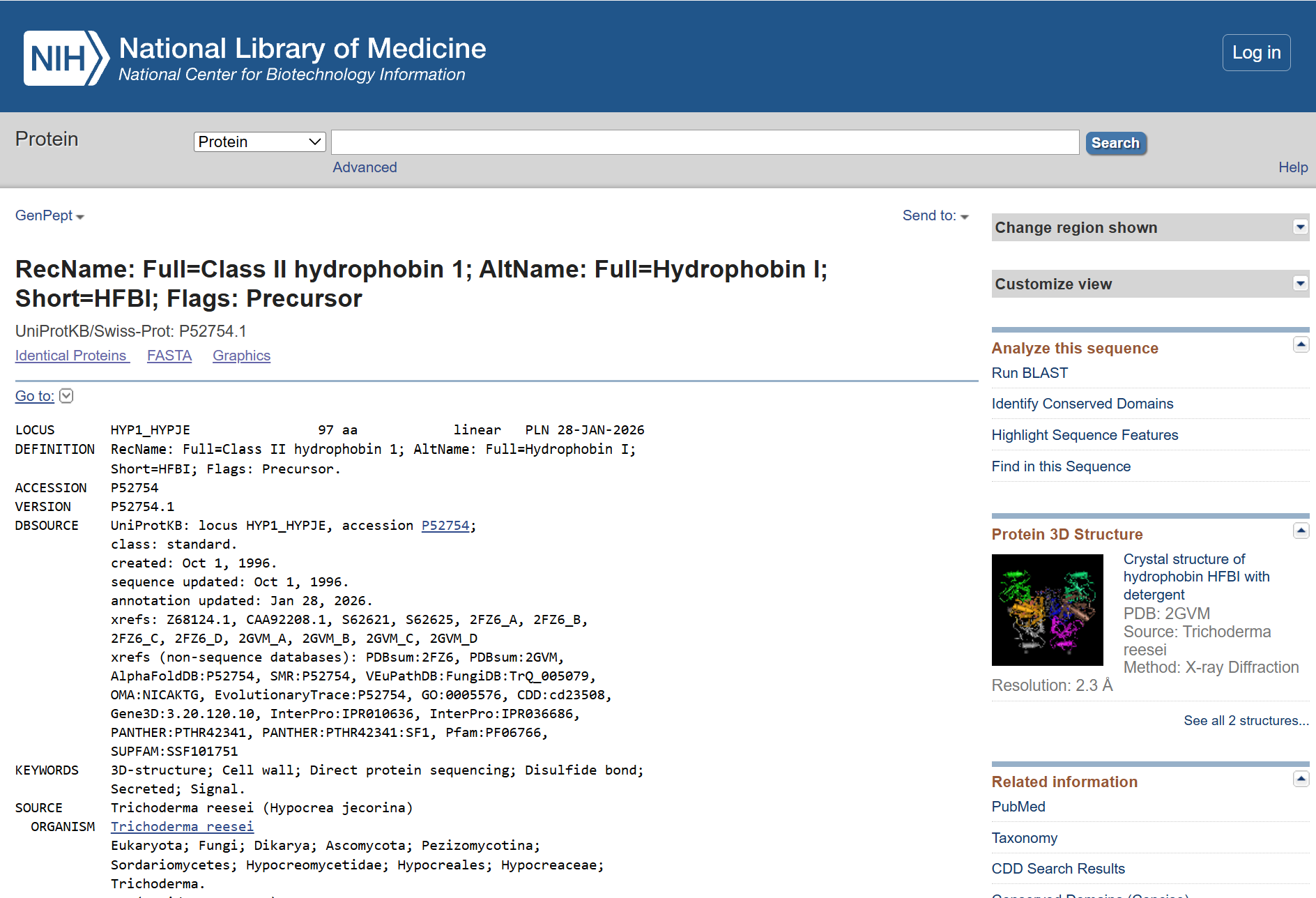
Hydrophobin HFBI de Trichoderma reesei AA sequence:
>sp|P52754.1|HYP1_HYPJE RecName: Full=Class II hydrophobin 1; AltName: Full=Hydrophobin I; Short=HFBI; Flags: Precursor
MKFFAIAALFAAAAVAQPLEDRSNGNGNVCPPGLFSNPQCCATQVLGLIGLDCKVPSQNVYDGTDFRNVC AKTGAQPLCCVAPVAGQALLCQTAVGA
3.2 Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence
Hydrophobin HFBI de Trichoderma reesei DNA sequence:
atgaaattttttgcgattgcggcgctgtttgcggcggcggcggtggcgcagccgctggaa gatcgcagcaacggcaacggcaacgtgtgcccgccgggcctgtttagcaacccgcagtgc tgcgcgacccaggtgctgggcctgattggcctggattgcaaagtgccgagccagaacgtg tatgatggcaccgattttcgcaacgtgtgcgcgaaaaccggcgcgcagccgctgtgctgc gtggcgccggtggcgggccaggcgctgctgtgccagaccgcggtgggcgcg
3.3. Codon optimization
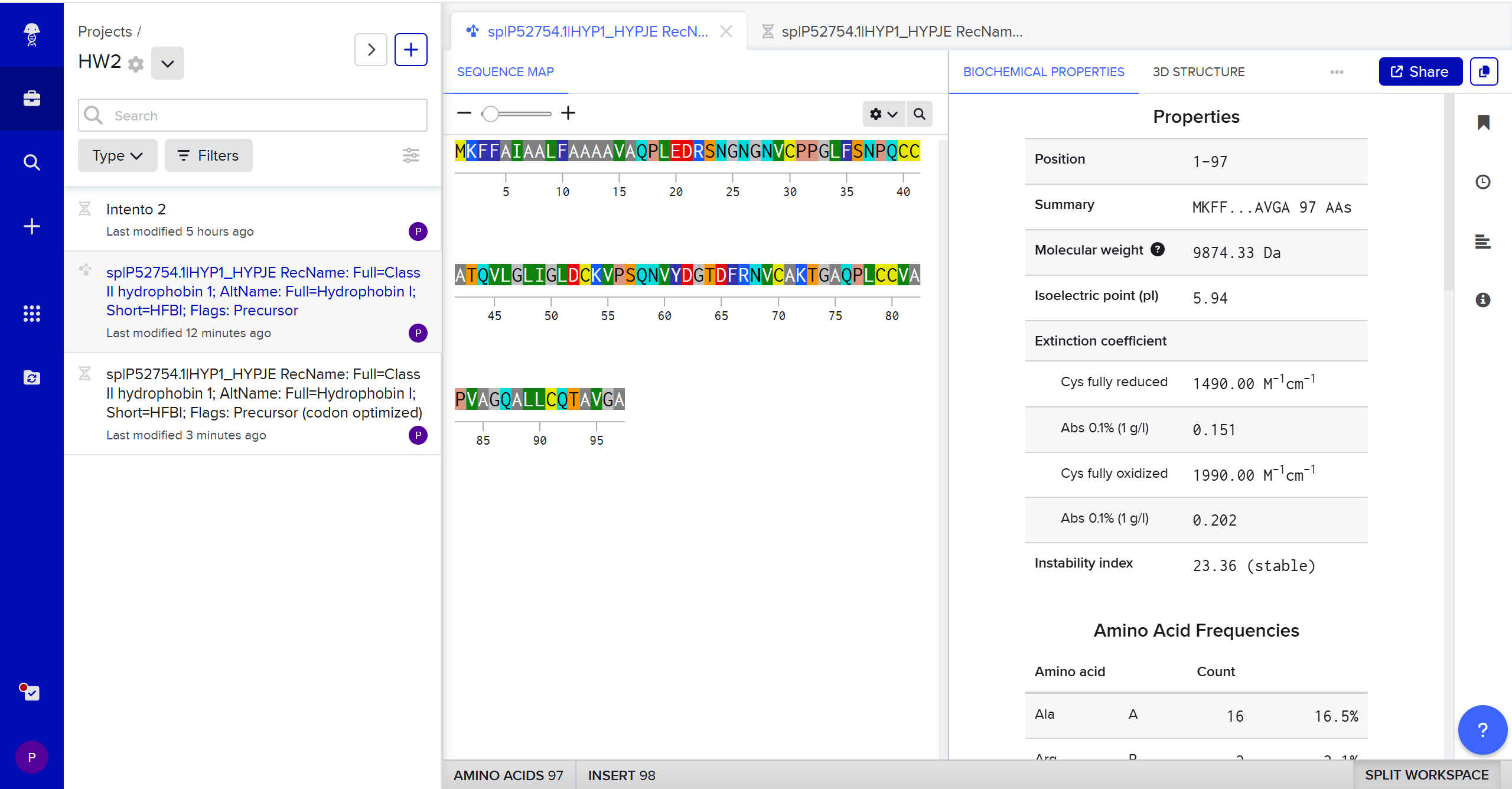
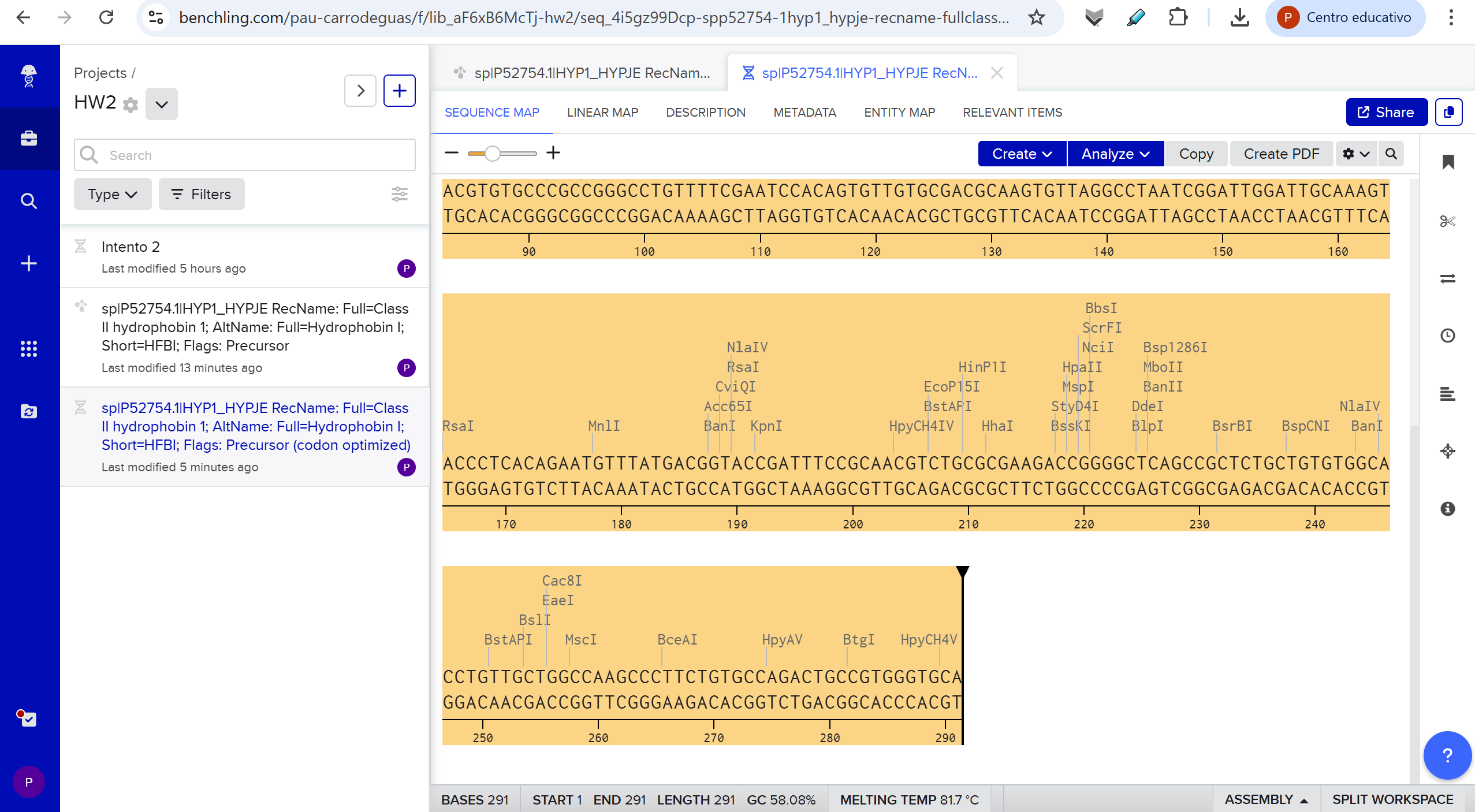
Hydrophobin HFBI de Trichoderma reesei with Codon-Optimization:
ATGAAATTTTTTGCCATTGCCGCGCTGTTCGCGGCGGCCGCGGTCGCACAGCCGCTGGAAGATCGTAGCAATGGCAACGGTAACGTGTGCCCGCCGGGCCTGTTTTCGAATCCACAGTGTTGTGCGACGCAAGTGTTAGGCCTAATCGGATTGGATTGCAAAGTACCCTCACAGAATGTTTATGACGGTACCGATTTCCGCAACGTCTGCGCGAAGACCGGGGCTCAGCCGCTCTGCTGTGTGGCACCTGTTGCTGGCCAAGCCCTTCTGTGCCAGACTGCCGTGGGTGCA
3.4. What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein.
Cell-Dependent Recombinant Protein Expression: After chemically synthesizing the DNA sequence encoding HFBI, the gene is inserted into a plasmid vector using DNA assembly methods such as Gibson Assembly or restriction enzyme-based cloning methods. Then, the recombinant plasmid is introduced into a host organism, which uses its own transcription and translation machinery to express the protein as the cells grow.
Cell-Free Protein Expression: In this method, instead of using living cells, the DNA sequence encoding HFBI is added to a reaction mixture containing ribosomes, enzymes, nucleotides, amino acids, and energy sources extracted from cells and transcription and translation occur directly in vitro [1]. Compared to in vivo techniques based on bacterial or tissue culture cells, in vitro protein expression is considerably faster because it does not require gene transfection, cell culture or extensive protein purification [2].
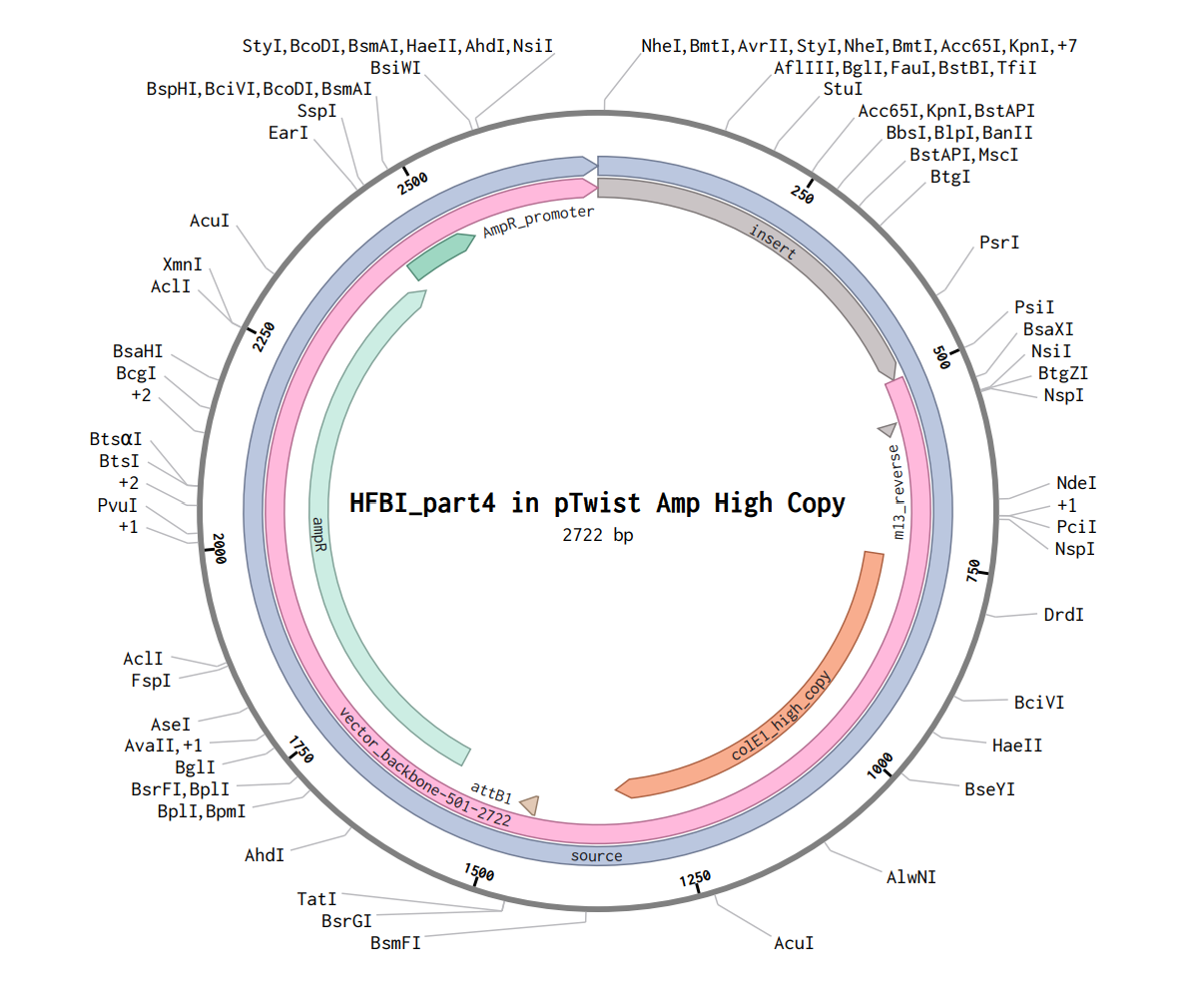
Part 4: Prepare a Twist DNA Synthesis Order
Build Your DNA Insert Sequence
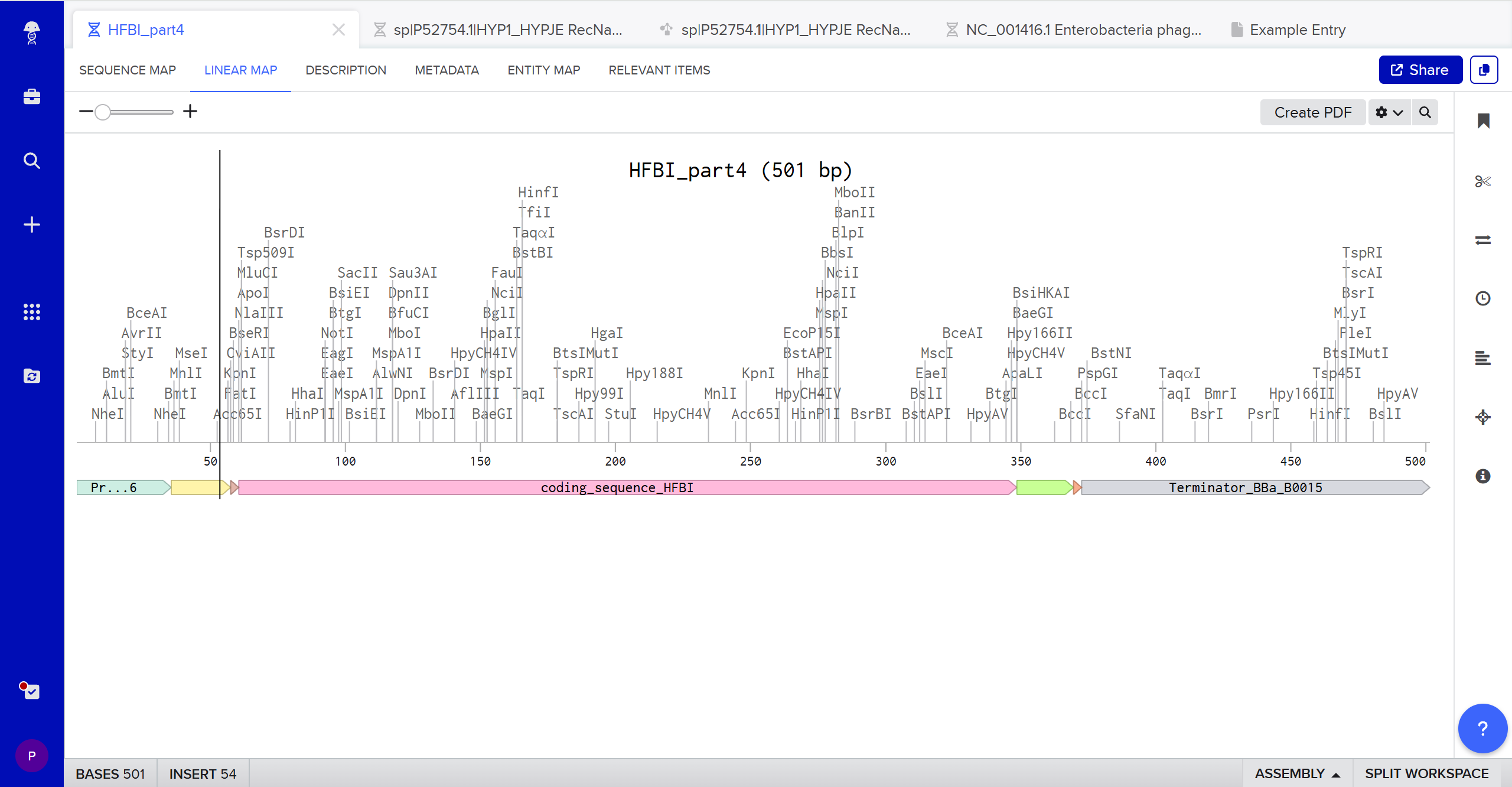

>HFBI_part4
TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGCCATTAAAGAGGAGAAAGGTACCATGAAATTTTTTGCCATTGCCGCGCTGTTCGCGGCGGCCGCGGTCGCACAGCCGCTGGAAGATCGTAGCAATGGCAACGGTAACGTGTGCCCGCCGGGCCTGTTTTCGAATCCACAGTGTTGTGCGACGCAAGTGTTAGGCCTAATCGGATTGGATTGCAAAGTACCCTCACAGAATGTTTATGACGGTACCGATTTCCGCAACGTCTGCGCGAAGACCGGGGCTCAGCCGCTCTGCTGTGTGGCACCTGTTGCTGGCCAAGCCCTTCTGTGCCAGACTGCCGTGGGTGCACATCACCATCACCATCATCACTAACCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA

Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence and why?
I would sequence the synthetic bispecific nanobody construct designed to bind CD44 and block IL-17 signaling. Sequencing would allow me to verify that the DNA was synthesized correctly, confirm the absence of mutations, and ensure the construct is suitable for expression in the probiotic host before experimental use.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
I would use next-generation sequencing (NGS), specifically Illumina sequencing, to analyze the engineered construct. Illumina platforms provide high accuracy, relatively low cost per base, and are well suited for short constructs such as nanobody genes as well as targeted panels of inflammatory genes.
1. Is your method first-, second- or third-generation or other? How so?
This method is considered second-generation sequencing because it relies on massively parallel sequencing of many short DNA fragments simultaneously after amplification on a flow cell.
2. What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
The input would be DNA extracted either from the engineered plasmid containing the nanobody gene.
Essential preparation steps:
- DNA extraction and purification
- Fragmentation
- Adapter ligation to both ends of DNA fragments
- PCR amplification to generate sufficient material
- Loading onto the sequencing flow cell
3. What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
Essential steps:
- DNA fragments bind to complementary oligos on a flow cell.
- Clonal amplification creates clusters of identical molecules.
- Fluorescently labeled nucleotides are incorporated one base at a time.
- A camera detects the fluorescent signal after each cycle.
The color detected corresponds to a specific base (A, T, C, or G), which allows the sequence to be reconstructed digitally. This process is called base calling.
4. What is the output of your chosen sequencing technology?
The output is a large dataset of short DNA reads in digital format (FASTQ files), which can then be aligned to reference sequences or assembled to confirm the construct sequence.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
I would synthesize a genetic construct encoding a bispecific nanobody that can anchor to CD44 receptors while simultaneously blocking IL-17 inflammatory signaling. This construct would be expressed in a probiotic Lactobacillus strain to create a localized therapeutic system for endometriosis.
The goal is to combine targeted binding with immune modulation to reduce inflammation and lesion growth without systemic side effects.
The DNA construct would include:
- Promoter for bacterial expression
- Secretion signal peptide
- Anti-CD44 nanobody domain
- Flexible linker
- Anti-IL-17 nanobody domain
- Terminator sequence
(ii) What technology or technologies would you use to perform this DNA synthesis and why? I would use phosphoramidite chemical DNA synthesis combined with gene assembly, such as the synthesis services provided by companies like Twist Bioscience.
This method allows precise control of nucleotide sequence and is scalable for custom gene design.
1. What are the essential steps of your chosen sequencing methods?
Essential steps:
- Chemical synthesis of short oligonucleotides
- Assembly of oligos into full gene fragments
- Error correction and cloning into plasmid vectors
- Sequence verification
- Delivery as plasmid DNA
3. What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
- Error rates increase with longer sequences
- Cost increases with length and complexity
- Repetitive or GC-rich regions can be difficult to synthesize
- Turnaround time can vary depending on design complexity
5.3 DNA Edit
(i) What DNA would you want to edit and why?
I would edit the genome of a probiotic Lactobacillus strain to stably express the therapeutic bispecific nanobody. Genome integration would improve stability compared to plasmid-based expression and reduce the need for antibiotic selection.
This could enable long-term therapeutic delivery directly at mucosal surfaces.
(ii) What technology or technologies would you use to perform these DNA edits and why?
I would use CRISPR-Cas9 genome editing because it allows precise insertion of DNA sequences into specific genomic locations with relatively high efficiency.
1. How does your technology of choice edit DNA? What are the essential steps?
CRISPR-Cas9 uses a guide RNA to direct the Cas9 enzyme to a specific DNA sequence. Cas9 creates a double-strand break at that location. The cell’s repair machinery then inserts the desired DNA sequence using a repair template provided by the researcher.
2. What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
Inputs would include:
- Guide RNA targeting the desired genomic site
- Cas9 enzyme or expression plasmid
- Donor DNA template containing the nanobody gene
- Host bacterial cells
3. What are the limitations of your editing methods (if any) in terms of efficiency or precision?
- Off-target edits may occur
- Editing efficiency can vary between organisms
- Delivery of CRISPR components into cells can be challenging
- Integration success may require screening multiple clones
Bibliografia
[1] “Cell-Free Protein Expression | NEB.” Accessed: Feb. 17, 2026. [Online]. Available: https://www.neb.com/en/applications/protein-expression/cell-free-protein-expression
[2] “Cell-Free Protein Expression - PE.” Accessed: Feb. 17, 2026. [Online]. Available: https://www.thermofisher.com/ht/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/cell-free-protein-expression.html