Week 2 HW: DNA Read, Write & Edit
Part 1: Benchling & In-silico Gel Art
See this week’s lab protocol “Gel Art: Restriction Digests and Gel Electrophoresis” for details. Overview:
- Make a free account at benchling.com
- Import the Lambda DNA.
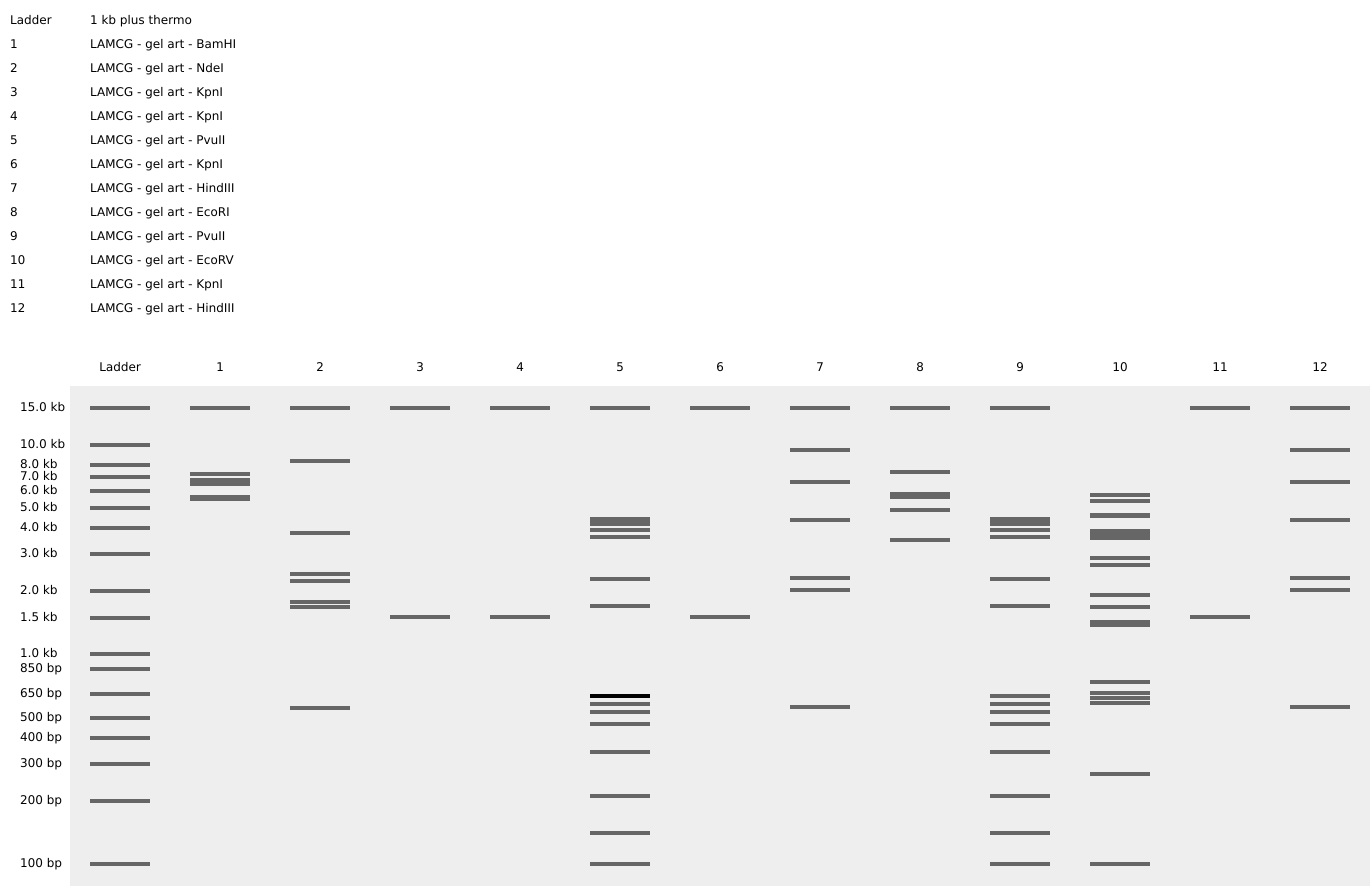
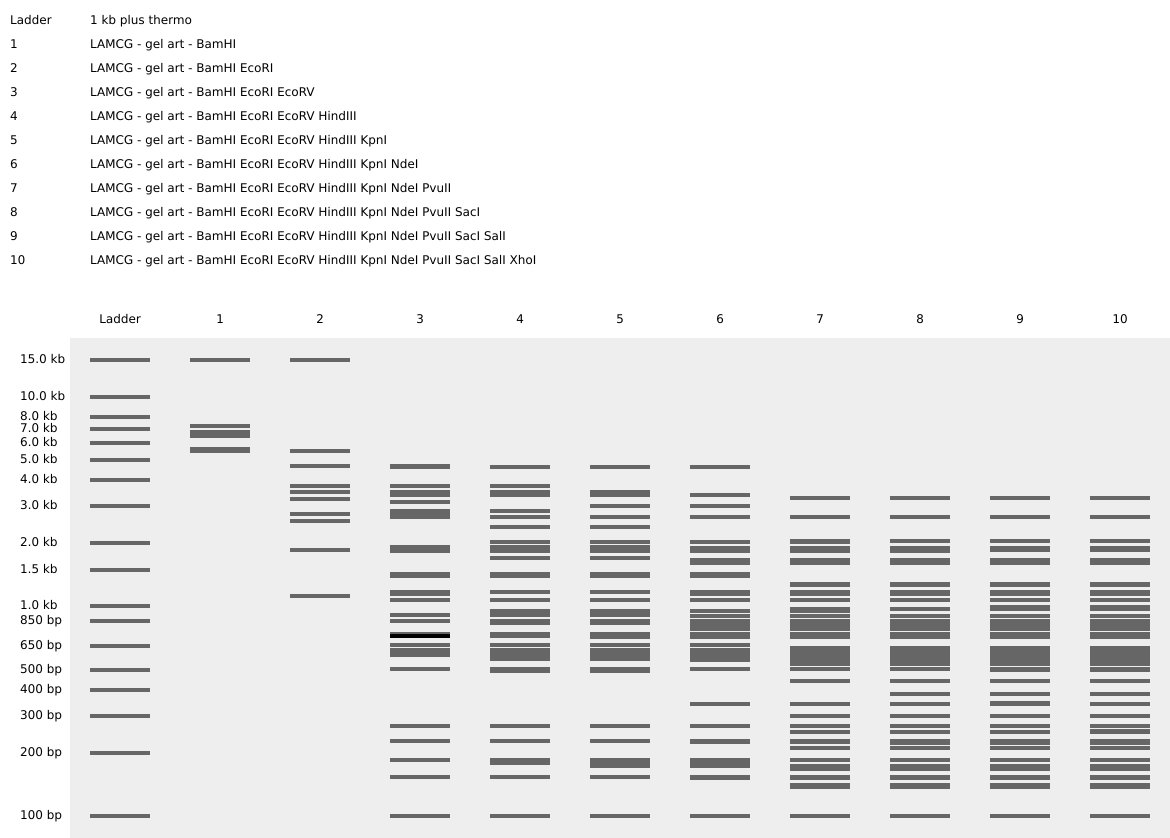
- Simulate Restriction Enzyme Digestion with the following Enzymes:
- EcoRI
- HindIII
- BamHI
- KpnI
- EcoRV
- SacI
- SalI
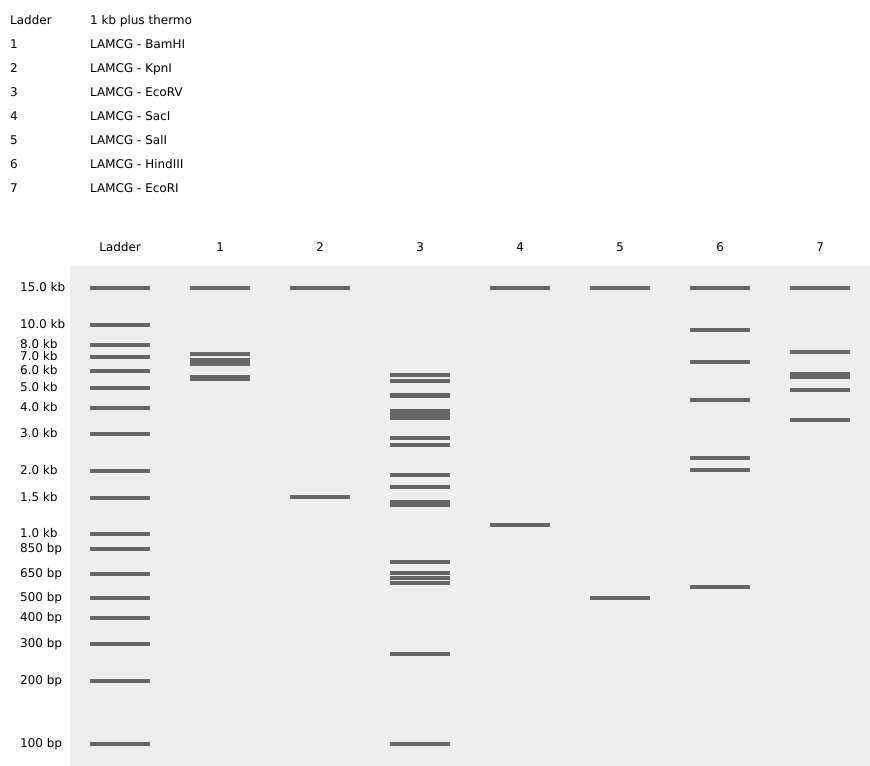

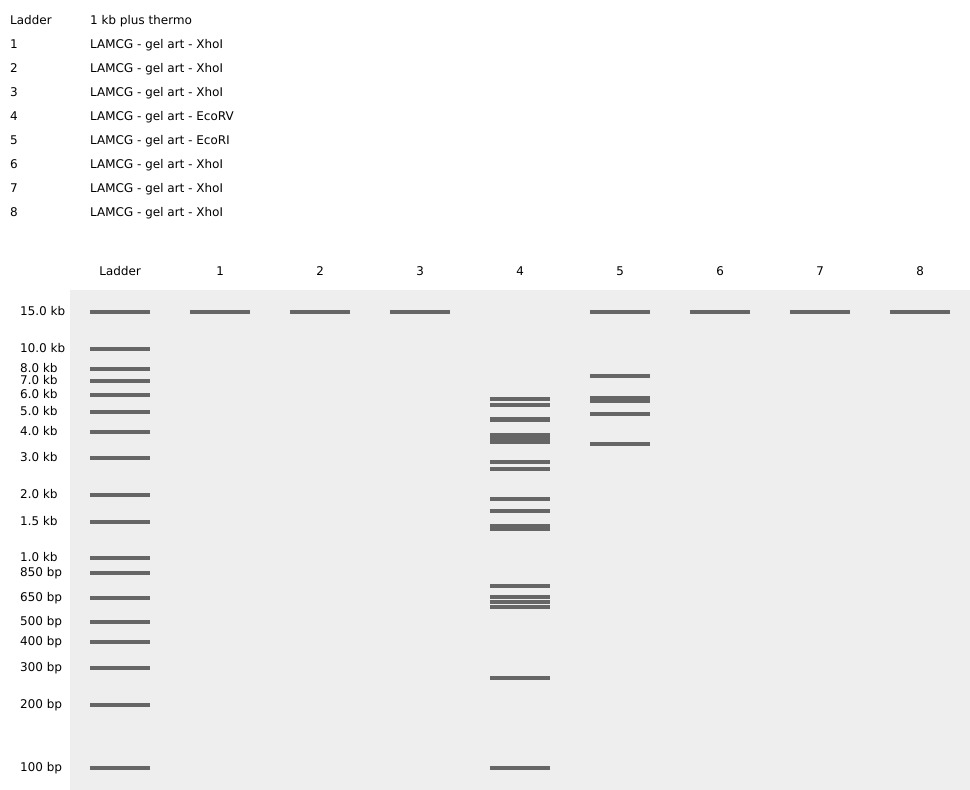
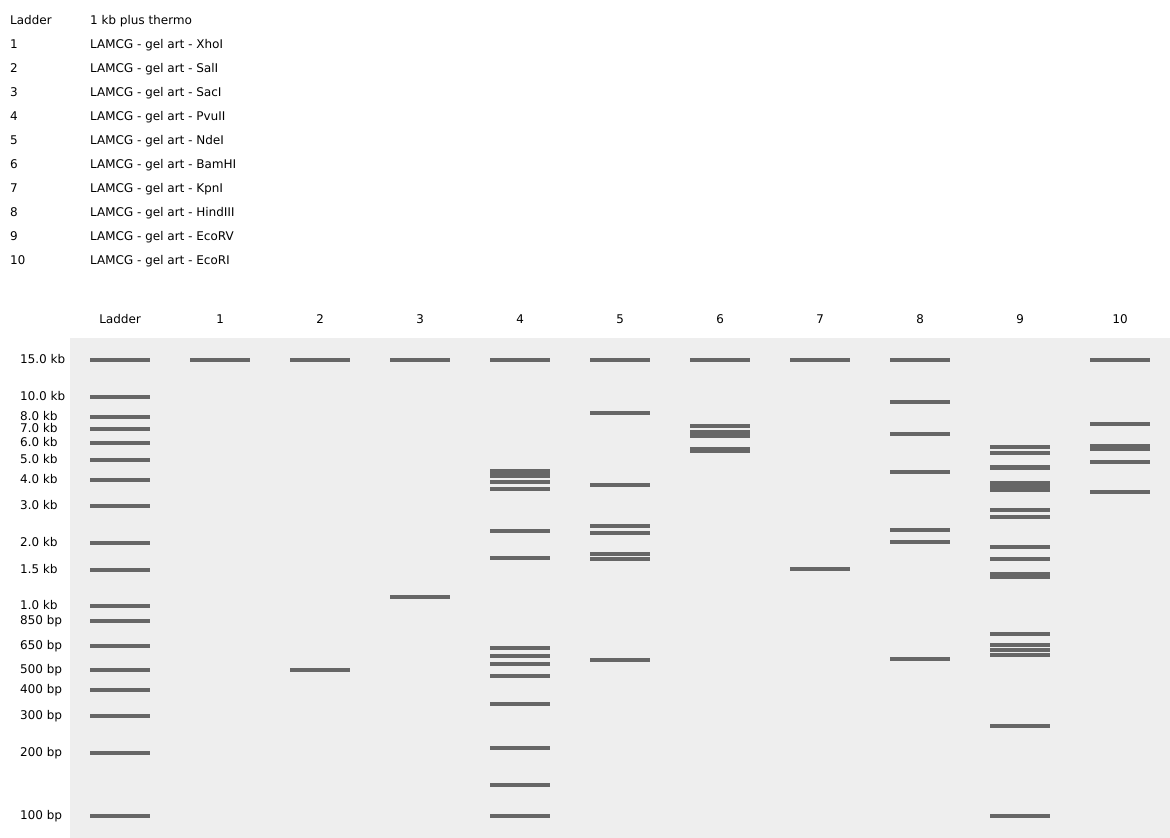
- Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks.
- You might find Ronan’s website a helpful tool for quickly iterating on designs!


Part 3: DNA Design Challenge
3.1. Choose your protein.
In recitation, we discussed that you will pick a protein for your homework that you find interesting. Which protein have you chosen and why? Using one of the tools described in recitation (NCBI, UniProt, google), obtain the protein sequence for the protein you chose.
[Example from our group homework, you may notice the particular format — The example below came from UniProt]
THYROGLOBULINE (CANIS LUPUS FAMILIARIS)
The world of proteins is so vast that choosing a single protein has been a profound task.
To follow the same path as week 1 HW, let´s start with Thyroglobulin, a very complex and specialized protein that is key to the generation of T3 and T4 hormones; in other words, it is a hormone protein. Because of its complexity, specificity, and its work with DNA, it is a modern protein. Some interesting facts about Thyroglobuline are: its size, it is very big in comparison to other proteins, it only functions in the thyroid gland, it is prone to being attacked to inmune system´s cells when something is not working well, and it does not accept errors in its process. If we compared it to the Actin protein, we could understand that Actin is a simpler protein that achieves a general action and that it is present in all eukaryotic forms since early life on Earth. Actin is the protein in charge of the formation of the cytoskeleton, motility, and shape of cells, among many other functions. The interesting fact about Actin is that it can allow errors to occur, in contrast to Thyroglobuline, which is very precise.
In the exercise bellow I will develop Thyroglobuline for Canis lupus familiaris.
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
The Central Dogma discussed in class and recitation describes the process in which DNA sequence becomes transcribed and translated into protein. The Central Dogma gives us the framework to work backwards from a given protein sequence and infer the DNA sequence that the protein is derived from. Using one of the tools discussed in class, NCBI or online tools (google “reverse translation tools”), determine the nucleotide sequence that corresponds to the protein sequence you chose above.
[Example: Get to the original sequence of phage MS2 L-protein from its genome phage MS2 genome - Nucleotide - NCBI]
3.3. Codon optimization.
Once a nucleotide sequence of your protein is determined, you need to codon optimize your sequence. You may, once again, utilize google for a “codon optimization tool”. In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
[Example from Codon Optimization Tool | Twist Bioscience while avoiding Type IIs enzyme recognition sites BsaI, BsmBI, and BbsI]
Codon optimization is needed to make the codon sequence of the original host be read or expressed in the codon sequence of the organism that will produce it later, without modifying the amino acid sequence. In the case of research, it is necessary to have a bank of protein that will be analyzed and tested; in that way, it is not sustainable to always get it from the original host for many reasons: budget, quantity, ethics, etc.
In the case of canine Thyroglobulin, the experiment will be based on the question: how to produce canine Thyroglobulin that could be used as one component of an implant of a thyroid gland? The cell organism that will produce the protein later will be CHO (Chinese hamster ovary cells), a mammal cell that has the capacity to perform complex processes to produce proteins as specialized as thyroglobulin.
3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
If I had to produce this from my DNA, I would have to use a codon-optimized sequence so that the protein could be interpreted by human cells. Although canine thyroglobuline and human thyroglobuline are not so different, and the DNA is not that different, it is necessary to make this step in order to have amino acids arranged in perfect order so that they can be read by mRNA. The technologies to do this would be:
cell-free methods: for producing the protein, specifically CHO cells that come from Chinese hamster ovaries. As this protein comes from a mammal, the cells for reproducing the protein need to be from the same group; it is not efficient to use bacterial cells like E.coli, for example, because the protein needs to fold in a specific way, and the differences between bacteria, mammal,s and plants make this process very different.
bioreactor: for scaling the production, avoiding cito-contamination, and giving the process a controlled atmosphere to fold and grow.
Part 4: Prepare a Twist DNA Synthesis Order
This is a practice exercise, not necessarily your real Twist order!
4.1. Create a Twist account and a Benchling account
Although the Benchling account was successfully created and used, the Twist account was not able to grant access.
4.2. Build Your DNA Insert Sequence
For example, let’s make a sequence that will make E. coli glow fluorescent green under UV light by constitutively (always) expressing sfGFP (a green fluorescent protein):
- In Benchling, select New DNA/RNA sequence
- Give your insert sequence a name and select DNA with a Linear topology (this is a linear sequence that will be inserted into a circular backbone vector of our choosing).
- Go through each piece of the given DNA sequences highlighted below (Promoter, RBS, Start Codon, Coding Sequence, His Tag, Stop Codon, Terminator) and paste the sequences into the Benchling file one after the other (replacing the coding sequence with your codon optimized DNA sequence of interest!). Each time you add a new piece of the sequence, make sure to annotate by right clicking over the sequence and creating an annotation that describes what each piece (e.g., Promoter, RBS, etc.) is (see image below).
For this part, I started by searching for the protein sequence in FASTA format, then I translated it to a DNA sequence and finally I made the codon optimization of the DNA sequence for e.coli.The results are bellow:
FASTA SEQUENCE:
REVERSE TRANSLATE:
CODON OPTIMIZATION FOR E.COLI
After that, I imported the sequence to Benchling, it looked like this:
And finally, I added each of the parts listed before:
- Promoter(e.g. BBa_J23106): TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGC
- RBS (e.g. BBa_B0034 with spacers for optimal expression): CATTAAAGAGGAGAAAGGTACC
- Start Codon: ATG
- Coding Sequence (your codon optimized DNA for a protein of interest, sfGFP for example): for this part I used the one listed before
- 7x His Tag (Let’s add a 7×His tag at the C-terminus of the protein to enable protein purification from E. coli): CATCACCATCACCATCATCAC
- Stop Codon: TAA
- Terminator (e.g. BBa_B0015): CCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA
The final result was this:
🍊Orange: promoter 🌾 Light orange: RBS 🌞 Yellow: Start codon 🍏 Green: coding sequence 💎 Sky blue: 7xHis Tag 🌞 Yellow: Stop codon 🎀 Pink: Terminator
Finally, this is the link of the construct:
- 4.3. On Twist, Select The “Genes” Option
- 4.4. Select “Clonal Genes” option
- 4.5. Import your sequence
- 4.6. Choose Your Vector
Part 5: DNA Read/Write/Edit
5.1 DNA Read (i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I would like to sequence the DNA of an ecuatorian tree called Cholán, or Tecoma stans.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why? Also answer the following questions:
- Is your method first-, second- or third-generation or other? How so?
- What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
- What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
- What is the output of your chosen sequencing technology?
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
(ii) What technology or technologies would you use to perform this DNA synthesis and why? Also answer the following questions:
- What are the essential steps of your chosen sequencing methods?
- What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
5.3 DNA Edit
(i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
(ii) What technology or technologies would you use to perform these DNA edits and why? Also answer the following questions:
- How does your technology of choice edit DNA? What are the essential steps?
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
- What are the limitations of your editing methods (if any) in terms of efficiency or precision?