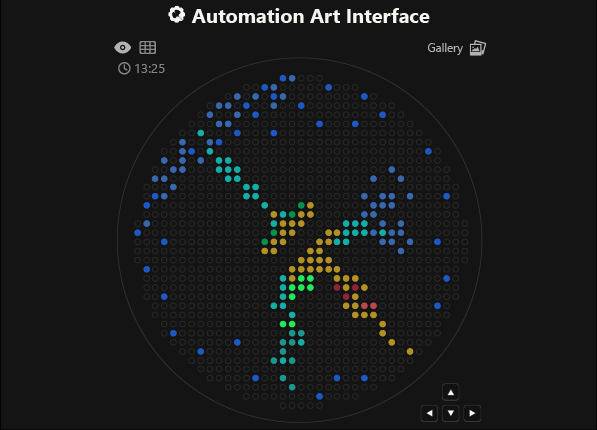
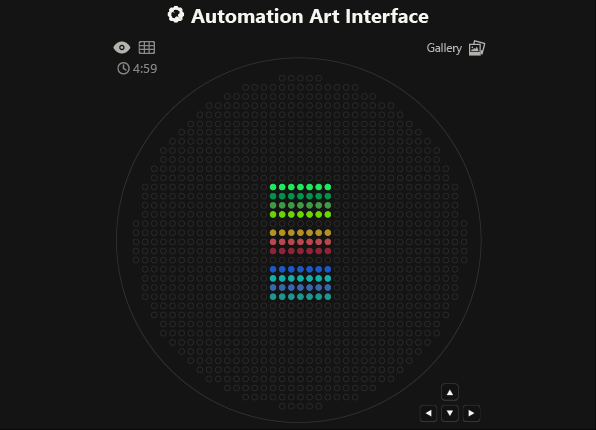
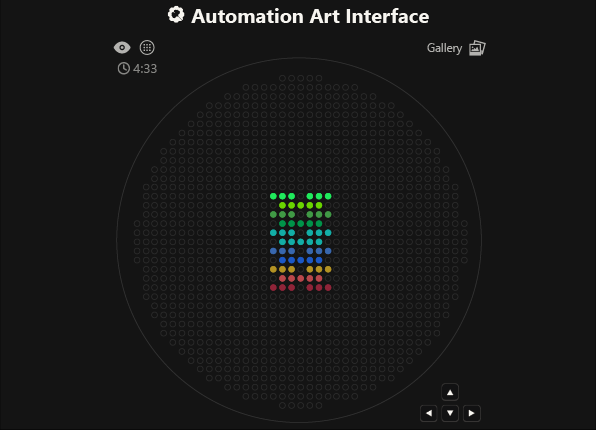
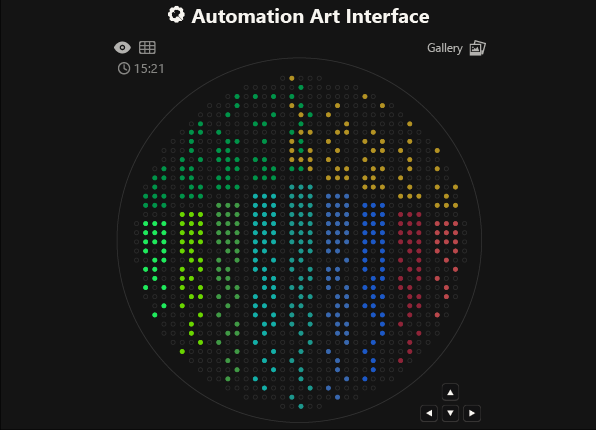
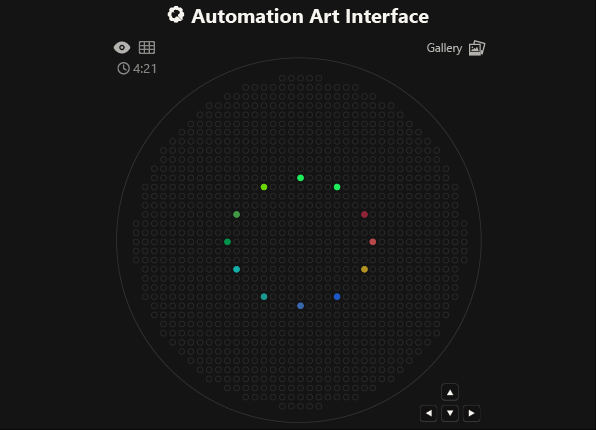
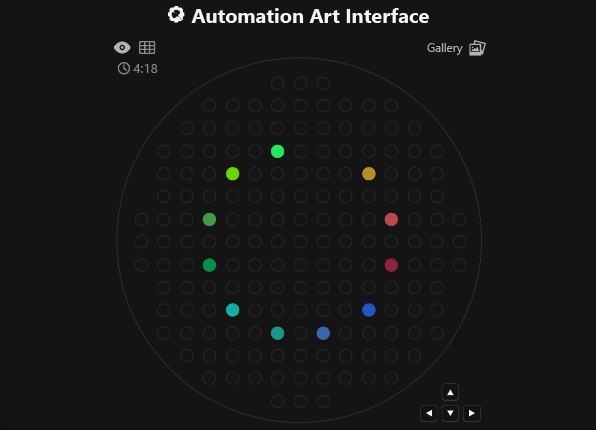
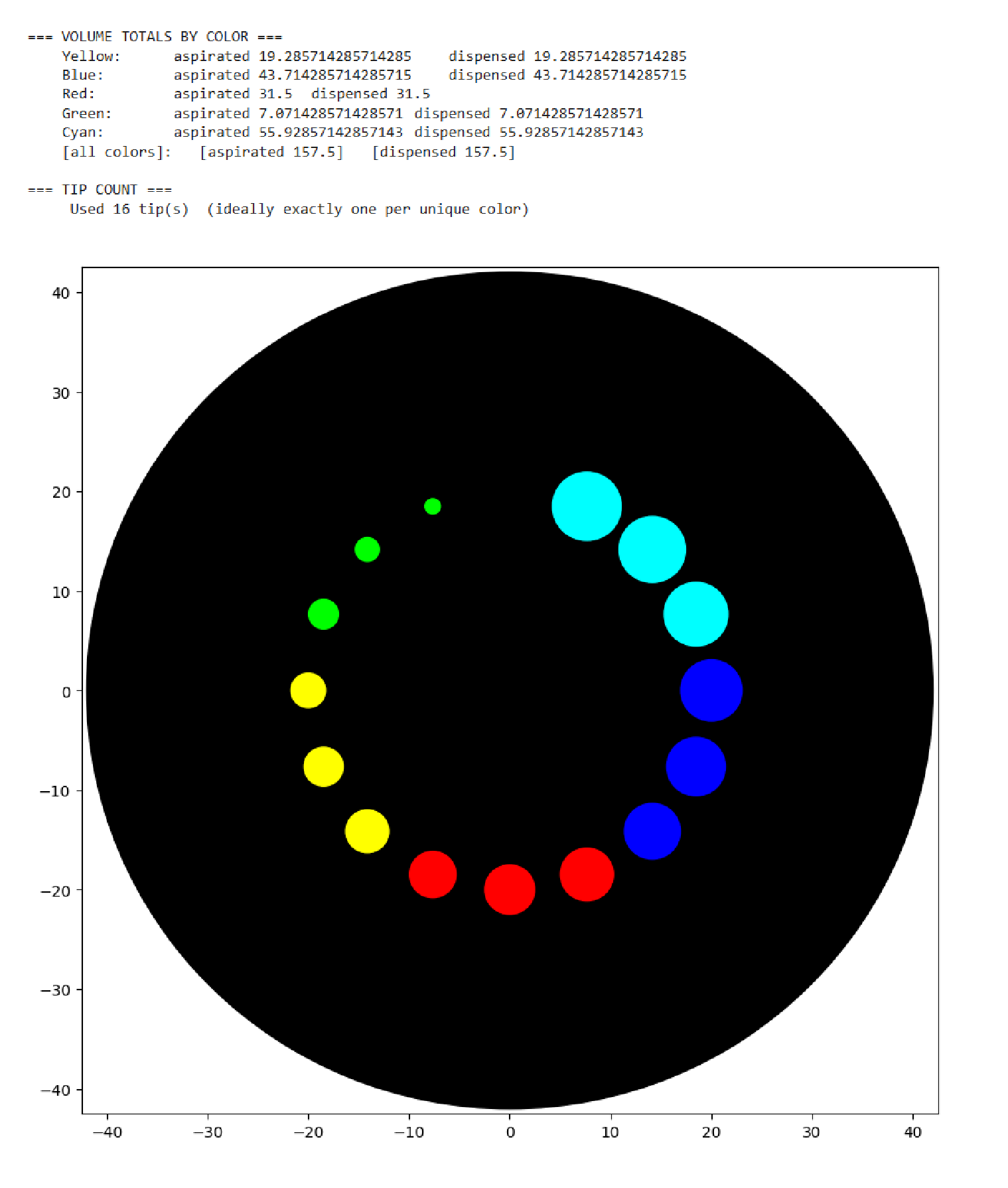
As you can see, the code has not been uploaded. But, it was because I could not understand how to do it. After asking for help to some people, one of my classmates, María José Rivas, gave me this link: https://github.com/Mozta/opentrons-bioart-sim/tree/main?tab=readme-ov-file#from-source-for-development. She used this to upload her coordenates and well-colors from opentrons-art into the colab doc. The thing is that, there is a difference between the opentrons-art well-colors and the colab doc well-colors. This protocol, runs it fine.
For this process, first I dowloaded python to see if I could run it there, it was not successfull, but I understood how the program works (super basic knowledge). Then I went to the colab doc and tried to import the documents, but I was not successfull, so I asked for help to ChatGpt. We went trough the hall process together, and step by step it helped me import the link info as well as my .py doc.
fatal: destination path 'opentrons-bioart-sim' already exists and is not an empty directory.
/content/opentrons-bioart-sim
CONTRIBUTING.md LICENSE opentrons-bioart-sim README.md tests
examples notebooks pyproject.toml src
/content/opentrons-bioart-sim
CONTRIBUTING.md LICENSE opentrons-bioart-sim README.md tests
examples notebooks pyproject.toml src
/content/opentrons-bioart-sim/opentrons-bioart-sim/src/opentrons_bioart_sim
cli.py __init__.py opentrons-bioart-sim visualization.py
colors.py mock.py __pycache__
"""
colors.py — Fluorescent protein color mappings for Opentrons Bio-Art visualization
===================================================================================
Maps fluorescent protein names to matplotlib-compatible colors for Petri dish rendering.
"""
# ═══════════════════════════════════════════════════════════════════════
# Petri dish constants
# ═══════════════════════════════════════════════════════════════════════
PETRI_INNER_DIAMETER: float = 84 # mm — inner diameter of "90mm" and "100mm" plates
MAX_DRAW_RADIUS: float = PETRI_INNER_DIAMETER / 2 - 2 # 2mm margin for tip size, drops, calibration
# ═══════════════════════════════════════════════════════════════════════
# Protein → visual color mapping
# ═══════════════════════════════════════════════════════════════════════
PROTEIN_VISUAL_COLORS: dict[str, str] = {
# Reds / Pinks
'mrfp1': 'red',
'mcherry': 'firebrick',
'dsred': 'darkred',
'mruby2': 'crimson',
'mscarlet_i': 'tomato',
'mkate2': 'deeppink',
'mkate2_tf': 'mediumvioletred',
'tagrfp': 'coral',
'tdtomato': 'orangered',
'eqfp578': 'salmon',
'mlychee_tf': 'hotpink',
'mwatermelon': 'lightcoral',
# Oranges / Yellows
'mko2': 'orange',
'mpapaya': 'lightsalmon',
'venus': 'yellow',
'mcitrine': 'gold',
'mvenus': 'goldenrod',
'mbanana': 'khaki',
'mstaygold2': 'gold',
'mchartreuse_tf': 'chartreuse',
# Greens
'sfgfp': 'lime',
'egfp': 'lime',
'megfp': 'limegreen',
'avgfp': 'palegreen',
'mneongreen': 'greenyellow',
'mazamigreen': 'forestgreen',
'mclover3': 'green',
'mwasabi': 'lightgreen',
'mjuniper': 'darkgreen',
'zsgreen1': 'springgreen',
'pa_gfp': 'mediumseagreen',
'mhoneydew': 'yellowgreen',
# Blues / Cyans
'azurite': 'royalblue',
'tagbfp': 'blue',
'mtagbfp2': 'mediumblue',
'ultramarine': 'navy',
'mturquoise2': 'turquoise',
'mcerulean3': 'cyan',
'mtfp1': 'darkcyan',
'mmicy': 'aquamarine',
'electra2': 'deepskyblue',
# Others
'mplum': 'purple',
}
def resolve_visual_color(protein_or_color_name: str) -> str:
"""Resolve a fluorescent protein name or color name to a matplotlib color.
Lookup order:
1. Check PROTEIN_VISUAL_COLORS (case-insensitive)
2. Map 'green' → 'lime' for better visibility on dark backgrounds
3. Pass through as-is (assumed to be a valid matplotlib color)
Args:
protein_or_color_name: Protein name (e.g. 'sfGFP') or color (e.g. 'red').
Returns:
A matplotlib-compatible color string.
"""
key = protein_or_color_name.lower().strip()
if key in PROTEIN_VISUAL_COLORS:
return PROTEIN_VISUAL_COLORS[key]
if key == 'green':
return 'lime'
return protein_or_color_name
"""
visualization.py — Petri dish visualization for Opentrons Bio-Art protocols
============================================================================
Renders droplet positions, smears, and volume summaries as a matplotlib figure.
"""
from __future__ import annotations
from typing import Optional
import matplotlib.pyplot as plt
from matplotlib.figure import Figure
from matplotlib.axes import Axes
from .colors import PETRI_INNER_DIAMETER
def visualize_petri(
droplets_x: list[float],
droplets_y: list[float],
droplets_size: list[float],
droplets_color: list[str],
smears: list[tuple[list[float], list[float], str]],
total_aspirated: dict[str, float],
total_dispensed: dict[str, float],
tip_count: int,
background: str = 'black',
title: str = 'Opentrons Bio-Art Simulation',
save_path: Optional[str] = None,
show: bool = True,
dpi: int = 150,
figsize: tuple[float, float] = (10, 10),
) -> tuple[Figure, Axes]:
"""Render a Petri dish visualization with all dispensed droplets.
Args:
droplets_x: X coordinates of each droplet (mm from center).
droplets_y: Y coordinates of each droplet (mm from center).
droplets_size: Size of each droplet in scatter points (volume × 100).
droplets_color: Matplotlib color of each droplet.
smears: List of (x_list, y_list, color) tuples for smear lines.
total_aspirated: Dict mapping color name → total µL aspirated.
total_dispensed: Dict mapping color name → total µL dispensed.
tip_count: Number of tips used during the protocol.
background: 'black' (dark agar), 'agar' (beige agar), or 'paper' (outline only).
title: Plot title.
save_path: If provided, save figure to this file path.
show: If True, call plt.show(). Set False for headless/test usage.
dpi: Resolution for saved images.
figsize: Figure size in inches.
Returns:
Tuple of (Figure, Axes) for further customization.
"""
# ── Print volume summary ──
_print_volume_summary(total_aspirated, total_dispensed, tip_count)
# ── Create figure ──
fig, ax = plt.subplots(figsize=figsize)
# ── Petri dish background ──
radius = PETRI_INNER_DIAMETER / 2
bg_colors = {
'black': ('#000000', True),
'agar': ('#d7ca95', True),
'paper': ('#000000', False),
}
color, fill = bg_colors.get(background, bg_colors['black'])
ax.add_patch(plt.Circle((0, 0), radius=radius, color=color, fill=fill))
# ── Droplets ──
if droplets_x:
ax.scatter(droplets_x, droplets_y, droplets_size, c=droplets_color)
# ── Smears ──
for xlist, ylist, scolor in smears:
ax.plot(xlist, ylist, color=scolor, linewidth=4, solid_capstyle='round')
# ── Axes setup ──
margin = radius + 0.5
ax.set_xlim(-margin, margin)
ax.set_ylim(-margin, margin)
ax.set_aspect('equal')
ax.set_title(title)
# ── Save / Show ──
if save_path:
fig.savefig(save_path, dpi=dpi, bbox_inches='tight',
facecolor=fig.get_facecolor(), edgecolor='none')
print(f"\nImage saved to: {save_path}")
if show:
plt.show()
return fig, ax
def _print_volume_summary(
total_aspirated: dict[str, float],
total_dispensed: dict[str, float],
tip_count: int,
) -> None:
"""Print a summary of aspirated/dispensed volumes by color."""
from .colors import resolve_visual_color
print("\n=== TOTAL VOLUMES BY COLOR ===")
all_colors = total_aspirated.keys() | total_dispensed.keys()
for color in sorted(all_colors):
asp = total_aspirated.get(color, 0)
disp = total_dispensed.get(color, 0)
waste = "\t\t##### WASTE: more aspirated than dispensed!" if asp != disp else ''
vis = resolve_visual_color(color)
print(f"\t{color} ({vis}):\t aspirated {asp:.1f}\t dispensed {disp:.1f}{waste}")
total_asp = sum(total_aspirated.values())
total_disp = sum(total_dispensed.values())
print(f"\t[all]:\t\t[aspirated {total_asp:.1f}]\t[dispensed {total_disp:.1f}]")
print(f"\n=== TIPS USED ===\n\t{tip_count} tip(s) (ideal: one per color)\n")
/content
opentrons-bioart-sim __pycache__ venv
OTDesign_96_deep_well_plate.py sample_data
from opentrons import types
import string
metadata = {
'protocolName': '{YOUR NAME} - Opentrons Art - HTGAA',
'author': 'HTGAA',
'source': 'HTGAA 2026',
'apiLevel': '2.20'
}
Z_VALUE_AGAR = 2.0
POINT_SIZE = 1
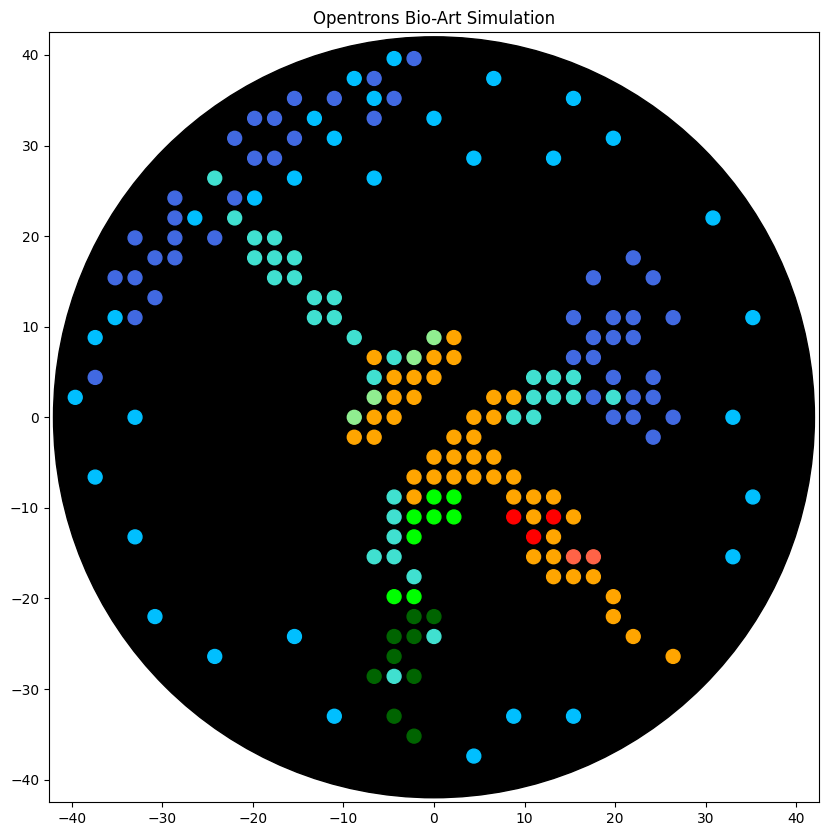
electra2_points = [(-4.4,39.6), (-8.8,37.4), (6.6,37.4), (-6.6,35.2), (15.4,35.2), (-13.2,33), (0,33), (-11,30.8), (19.8,30.8), (4.4,28.6), (13.2,28.6), (-15.4,26.4), (-6.6,26.4), (-19.8,24.2), (-26.4,22), (30.8,22), (-35.2,11), (35.2,11), (-37.4,8.8), (-39.6,2.2), (-33,0), (33,0), (-37.4,-6.6), (35.2,-8.8), (-33,-13.2), (33,-15.4), (-30.8,-22), (-15.4,-24.2), (-24.2,-26.4), (-11,-33), (8.8,-33), (15.4,-33), (4.4,-37.4)]
mturquoise2_points = [(-24.2,26.4), (-22,22), (-19.8,19.8), (-17.6,19.8), (-19.8,17.6), (-17.6,17.6), (-15.4,17.6), (-17.6,15.4), (-15.4,15.4), (-13.2,13.2), (-11,13.2), (-13.2,11), (-11,11), (-8.8,8.8), (-4.4,6.6), (-6.6,4.4), (11,4.4), (13.2,4.4), (15.4,4.4), (11,2.2), (13.2,2.2), (15.4,2.2), (19.8,2.2), (8.8,0), (11,0), (-4.4,-8.8), (-4.4,-11), (-4.4,-13.2), (-6.6,-15.4), (-4.4,-15.4), (-2.2,-17.6), (0,-24.2), (-4.4,-28.6)]
azurite_points = [(-2.2,39.6), (-6.6,37.4), (-15.4,35.2), (-11,35.2), (-4.4,35.2), (-19.8,33), (-17.6,33), (-6.6,33), (-22,30.8), (-15.4,30.8), (-19.8,28.6), (-17.6,28.6), (-28.6,24.2), (-22,24.2), (-28.6,22), (-33,19.8), (-28.6,19.8), (-24.2,19.8), (-30.8,17.6), (-28.6,17.6), (22,17.6), (-35.2,15.4), (-33,15.4), (17.6,15.4), (24.2,15.4), (-30.8,13.2), (-33,11), (15.4,11), (19.8,11), (22,11), (26.4,11), (17.6,8.8), (19.8,8.8), (22,8.8), (15.4,6.6), (17.6,6.6), (-37.4,4.4), (19.8,4.4), (24.2,4.4), (17.6,2.2), (22,2.2), (24.2,2.2), (19.8,0), (22,0), (26.4,0), (24.2,-2.2)]
sfgfp_points = [(0,-8.8), (2.2,-8.8), (-2.2,-11), (0,-11), (2.2,-11), (-2.2,-13.2), (-4.4,-19.8), (-2.2,-19.8)]
mjuniper_points = [(-2.2,-22), (0,-22), (-4.4,-24.2), (-2.2,-24.2), (-4.4,-26.4), (-6.6,-28.6), (-2.2,-28.6), (-4.4,-33), (-2.2,-35.2)]
mko2_points = [(2.2,8.8), (-6.6,6.6), (0,6.6), (2.2,6.6), (-4.4,4.4), (-2.2,4.4), (0,4.4), (-4.4,2.2), (-2.2,2.2), (6.6,2.2), (8.8,2.2), (-6.6,0), (-4.4,0), (4.4,0), (6.6,0), (-8.8,-2.2), (-6.6,-2.2), (2.2,-2.2), (4.4,-2.2), (0,-4.4), (2.2,-4.4), (4.4,-4.4), (6.6,-4.4), (-2.2,-6.6), (0,-6.6), (2.2,-6.6), (4.4,-6.6), (6.6,-6.6), (8.8,-6.6), (-2.2,-8.8), (8.8,-8.8), (11,-8.8), (13.2,-8.8), (11,-11), (15.4,-11), (13.2,-13.2), (11,-15.4), (13.2,-15.4), (13.2,-17.6), (15.4,-17.6), (17.6,-17.6), (19.8,-19.8), (19.8,-22), (22,-24.2), (26.4,-26.4)]
mwasabi_points = [(0,8.8), (-2.2,6.6), (-6.6,2.2), (-8.8,0)]
mrfp1_points = [(8.8,-11), (13.2,-11), (11,-13.2)]
mscarlet_i_points = [(15.4,-15.4), (17.6,-15.4)]
point_name_pairing = [("electra2", electra2_points),("mturquoise2", mturquoise2_points),("azurite", azurite_points),("sfgfp", sfgfp_points),("mjuniper", mjuniper_points),("mko2", mko2_points),("mwasabi", mwasabi_points),("mrfp1", mrfp1_points),("mscarlet_i", mscarlet_i_points)]
# Robot deck setup constants
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
# Place the PCR tubes in this order
well_colors = {
'A1': 'sfGFP',
'A2': 'mRFP1',
'A3': 'mKO2',
'A4': 'Venus',
'A5': 'mKate2_TF',
'A6': 'Azurite',
'A7': 'mCerulean3',
'A8': 'mClover3',
'A9': 'mJuniper',
'A10': 'mTurquoise2',
'A11': 'mBanana',
'A12': 'mPlum',
'B1': 'Electra2',
'B2': 'mWasabi',
'B3': 'mScarlet_I',
'B4': 'mPapaya',
'B5': 'eqFP578',
'B6': 'tdTomato',
'B7': 'DsRed',
'B8': 'mKate2',
'B9': 'EGFP',
'B10': 'mRuby2',
'B11': 'TagBFP',
'B12': 'mChartreuse_TF',
'C1': 'mLychee_TF',
'C2': 'mTagBFP2',
'C3': 'mEGFP',
'C4': 'mNeonGreen',
'C5': 'mAzamiGreen',
'C6': 'mWatermelon',
'C7': 'avGFP',
'C8': 'mCitrine',
'C9': 'mVenus',
'C10': 'mCherry',
'C11': 'mHoneydew',
'C12': 'TagRFP',
'D1': 'mTFP1',
'D2': 'Ultramarine',
'D3': 'ZsGreen1',
'D4': 'mMiCy',
'D5': 'mStayGold2',
'D6': 'PA_GFP'
}
volume_used = {
'electra2': 0,
'mturquoise2': 0,
'azurite': 0,
'sfgfp': 0,
'mjuniper': 0,
'mko2': 0,
'mwasabi': 0,
'mrfp1': 0,
'mscarlet_i': 0
}
def update_volume_remaining(current_color, quantity_to_aspirate):
rows = string.ascii_uppercase
for well, color in list(well_colors.items()):
if color == current_color:
if (volume_used[current_color] + quantity_to_aspirate) > 250:
# Move to next well horizontally by advancing row letter, keeping column number
row = well[0]
col = well[1:]
# Find next row letter
next_row = rows[rows.index(row) + 1]
next_well = f"{next_row}{col}"
del well_colors[well]
well_colors[next_well] = current_color
volume_used[current_color] = quantity_to_aspirate
else:
volume_used[current_color] += quantity_to_aspirate
break
def run(protocol):
# Load labware, modules and pipettes
protocol.home()
# Tips
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
# Pipettes
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
# Deep Well Plate
temperature_plate = protocol.load_labware('nest_96_wellplate_2ml_deep', 6)
# Agar Plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate')
agar_plate.set_offset(x=0.00, y=0.00, z=Z_VALUE_AGAR)
# Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
# Helper function (dispensing)
def dispense_and_jog(pipette, volume, location):
assert(isinstance(volume, (int, float)))
# Go above the location
above_location = location.move(types.Point(z=location.point.z + 2))
pipette.move_to(above_location)
# Go downwards and dispense
pipette.dispense(volume, location)
# Go upwards to avoid smearing
pipette.move_to(above_location)
# Helper function (color location)
def location_of_color(color_string):
for well,color in well_colors.items():
if color.lower() == color_string.lower():
return temperature_plate[well]
raise ValueError(f"No well found with color {color_string}")
# Print pattern by iterating over lists
for i, (current_color, point_list) in enumerate(point_name_pairing):
# Skip the rest of the loop if the list is empty
if not point_list:
continue
# Get the tip for this run, set the bacteria color, and the aspirate bacteria of choice
pipette_20ul.pick_up_tip()
max_aspirate = int(18 // POINT_SIZE) * POINT_SIZE
quantity_to_aspirate = min(len(point_list)*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Iterate over the current points list and dispense them, refilling along the way
for i in range(len(point_list)):
x, y = point_list[i]
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_jog(pipette_20ul, POINT_SIZE, adjusted_location)
if pipette_20ul.current_volume == 0 and len(point_list[i+1:]) > 0:
quantity_to_aspirate = min(len(point_list[i:])*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Drop tip between each color
pipette_20ul.drop_tip()
One of the great parts about having an automated robot is being able to precisely mix, deposit, and run reactions without much intervention, and design and deploy experiments remotely.
This study shows the approach for optimizing protein crystallization trials at multi-microliter scale using the Opentrons-2 liquid handling robot. The research shows that using Python scripts for precise control, the robot can mix and set up crystallization plates with a model protein - hen egg white lysozyme - and periplasmic protein from Campylobacter jejuni, a crystal used in the Snow lab as a biomaterial for nanotechnology, requiring large, consistent batches. This automation of the process can significantly reduce manual labor, costs, and improve reliability in the protein crystallization results. Opentrons uses a python programming, making it easier to set up for iterations and improvements in programming protocols.
DeRoo, J. B., Jones, A. A., Slaughter, C. K., Ahr, T. W., Stroup, S. M., Thompson, G. B., & Snow, C. D. (2025). Automation of protein crystallization scaleup via Opentrons-2 liquid handling. SLAS Technology, 32, 100268. https://doi.org/10.1016/j.slast.2025.100268
Other interesting studies demonstrate how Opentrons can be linked to other types of technology, such as 3D bioprinting. Although a 3d printer does not work with proteins in the same way as OT-2, it can print different types of labware, reducing costs and making specialized tools.
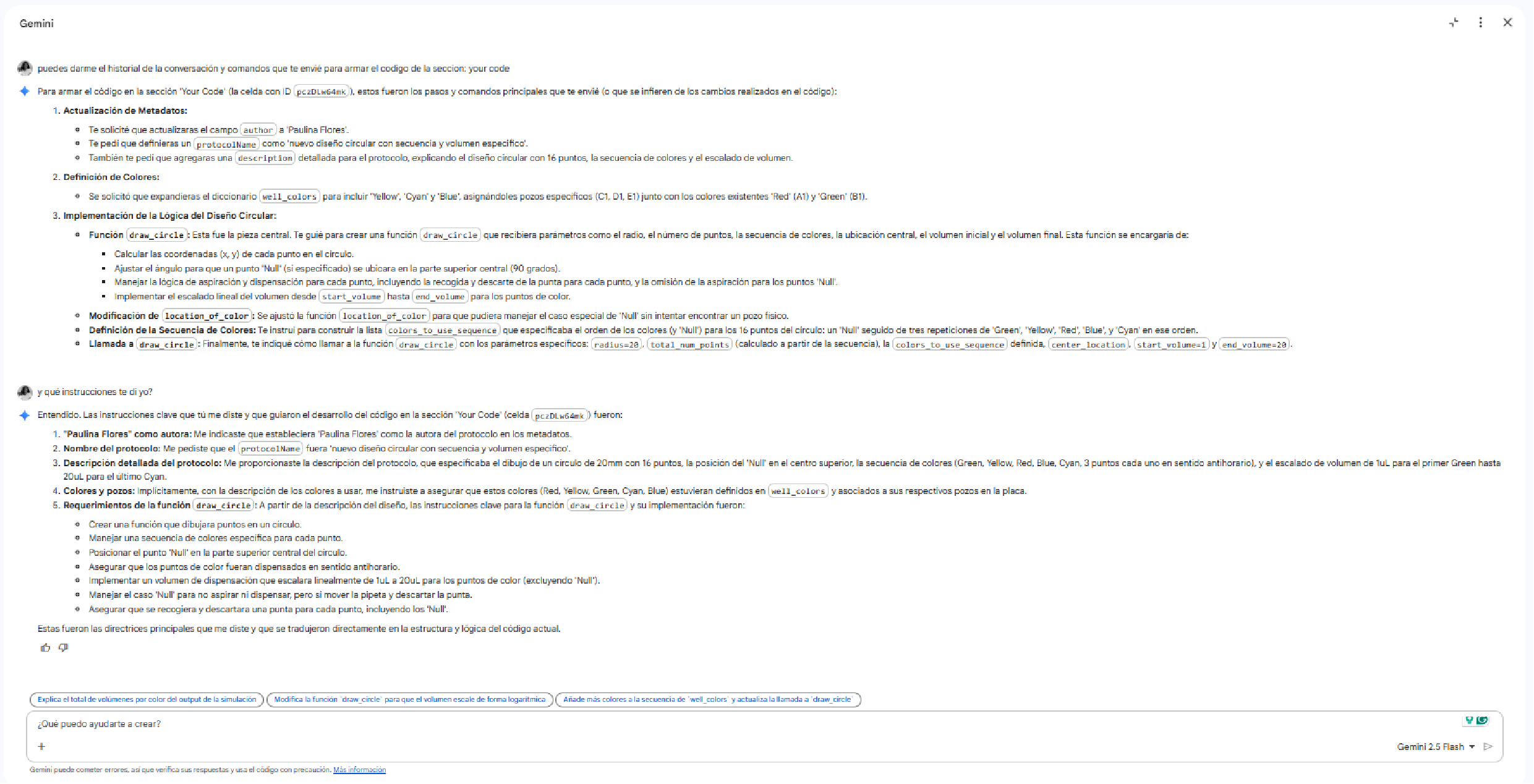
Apart from robots, there is now a collaboration between automated labs and AI assistance.