Week 4: Protein Design - part I
Part A. Conceptual Questions
Answer any NINE of the following questions from Shuguang Zhang: (i.e. you can select two to skip)
- How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
- Why do humans eat beef but do not become a cow, eat fish but do not become fish?
- Why are there only 20 natural amino acids?
- Can you make other non-natural amino acids? Design some new amino acids.
- Where did amino acids come from before enzymes that make them, and before life started?
- If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
- Can you discover additional helices in proteins?
- Why are most molecular helices right-handed?
- Why do β-sheets tend to aggregate?
- What is the driving force for β-sheet aggregation?
- Why do many amyloid diseases form β-sheets?
- Can you use amyloid β-sheets as materials?
- Design a β-sheet motif that forms a well-ordered structure.
Part B: Protein Analysis and Visualization
In this part of the homework, you will be using online resources and 3D visualization software to answer questions about proteins. Pick any protein (from any organism) of your interest that has a 3D structure and answer the following questions:
LUCIFERASE OF Pyrophorus plagiophthalamus
Luciferase is a protein/enzyme that generates bioluminescence by catalyzing oxidation of D-luciferine in the presence of ATP, oxygen, and MG+2. In the case of this particular insect, Pyrophorus plagiophthalmus, different isoforms of luciferasecan emit light ranging from green to orange, depending on which organs it expresses the gene. These variations of color arise from subtle structural variations in the enzyme´s active site that alter the electronic environment of the excited oxyluciferin intermediate. Click Beetle´s luciferase is a very stable protein in a wide range of pH range compared to other active luciferases. It is very common to use this enzyme for in vivo imaging applications, especially the red-emitting variants. They are also used as a biosensor to monitor gene expression and as a gene reporter.
I chose this particular protein because I am interested in analyzing how sound frequencies might influence bacterial protein expression, growth dynamics, or spatial organization. In this way, having luciferase as a biosensor is ideal; light emission provides a real-time, quantifiable readout.
Burbelo, P. D., Kisailus, A. E., & Peck, J. W. (2002). Detecting Protein-Protein Interactions Using Renilla Luciferase Fusion Proteins. BioTechniques, 33(5), 1044–1050. https://doi.org/10.2144/02335st05
How long is it? What is the most frequent amino acid?
For this part, I used Google Colab and did some research on Leucine. Luciferase of Pryphorus plagiophtalmus: has 543 amino acids, being the most frequent L (Leucina) that appears 56 times. Leucine is commonly known for being an amino acid that helps synthesize muscle proteins and supports tissue regeneration. In this case, its function is related to a hydrophobic nucleotide, correct protein folding, and formation of alpha helices.
How many protein sequence homologs are there for your protein?
According to Uniprot´s BLAST TOOL, it has 236 homologs. This means that there is a variety of similar proteins in the living realm. They might not be the same, but they share a very similar structure. These homologs can be orthologs and parologs. The second ones are proteins that can be found inside the insect’s body, but with very subtle variations in their structure.
Does your protein belong to any protein family?
Yes, it belongs to the luciferase proteins of insects. This type of protein needs ATP, d-luciferin, and oxygen to perform the oxidation process.
When was the structure solved? Is it a good quality structure? A good-quality structure is one with high resolution. Smaller the better (Resolution: 2.70 Å) :
This particular protein, Luciferase of Pyrophorus plagiophthalamus, is not in the bank information of RCSB, so I took the first luciferase structured in the bank which is: 1LCI Firefly luciferase from Photinus pyralis. Its structure was solved in 1997. The quality is 2.00 Å, which is a good quality.
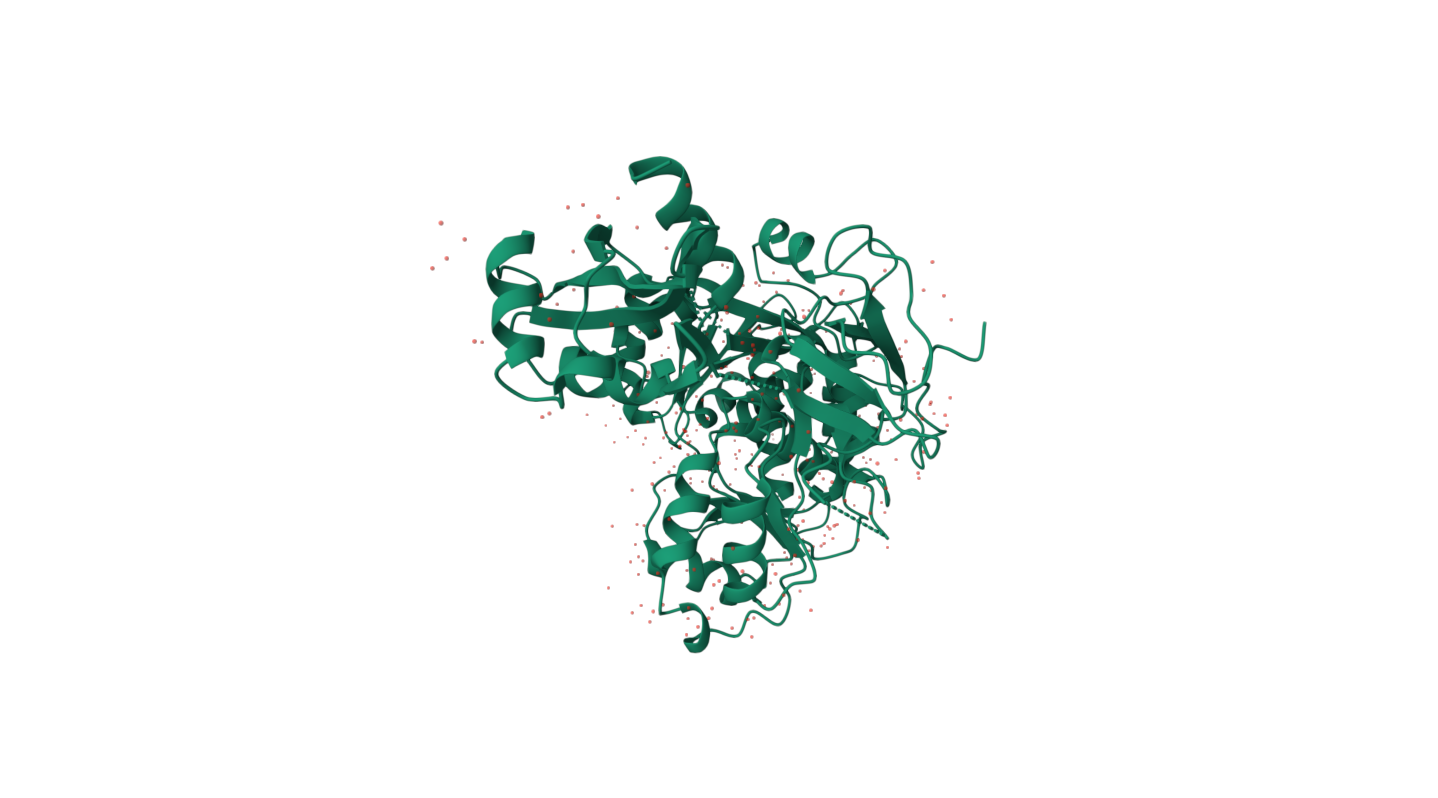
Are there any other molecules in the solved structure apart from protein?
There is a presence chrystallographic molecules of water (HOH), which stabilizes the protein and may participate in hydrogen bond formation. As this is the first protein from luciferase to be structured, it does not include other types of components, besides the protein and water.
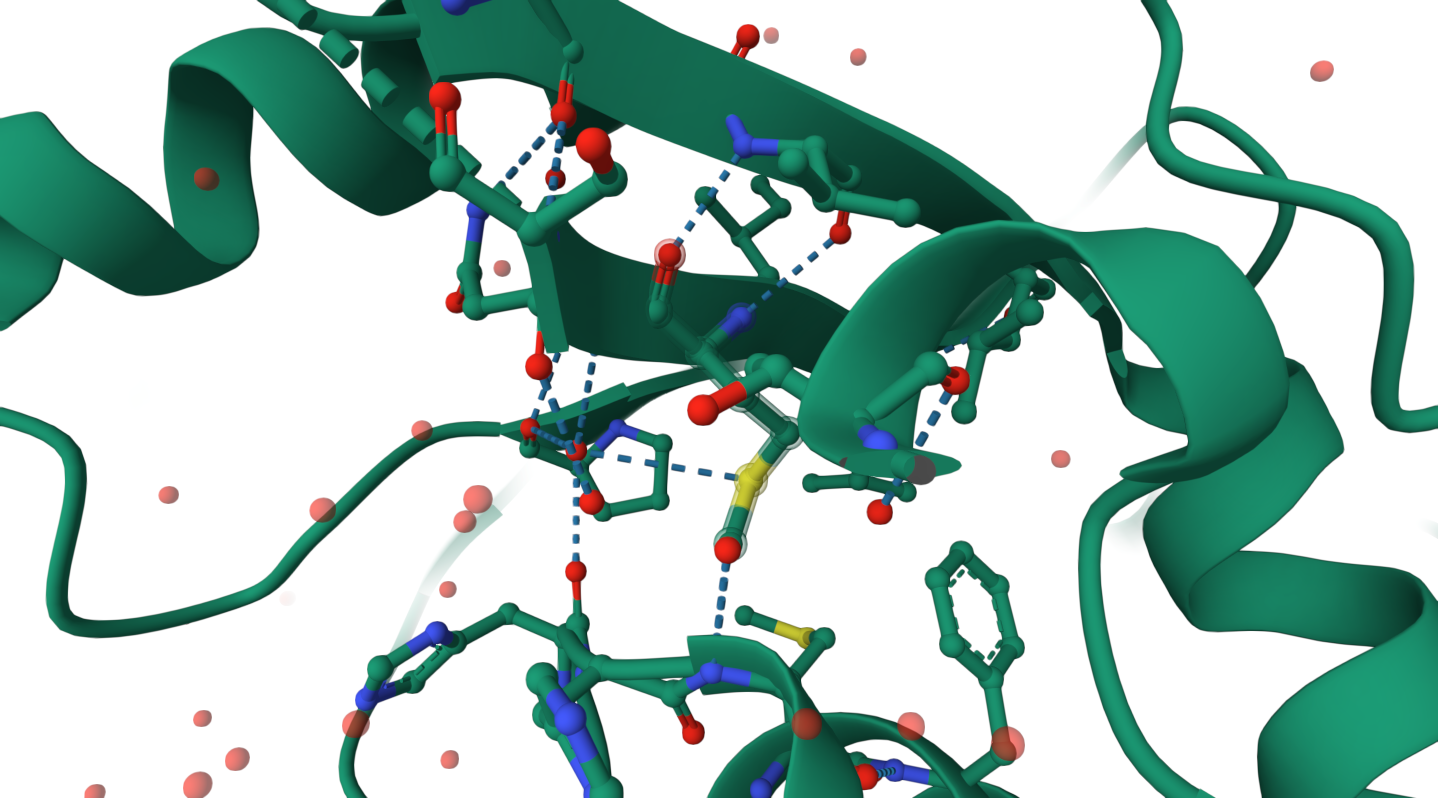
Does your protein belong to any structure classification family?
It belongs to the ATP-dependent AMP-binding enzyme family. This family includes enzymes that activate substrates through adenylation using ATP, forming an AMP-bound intermediate.
Open the structure of your protein in any 3D molecule visualization software: - PyMol Tutorial Here (hint: ChatGPT is good at PyMol commands) - Visualize the protein as “cartoon”, “ribbon” and “ball and stick”.
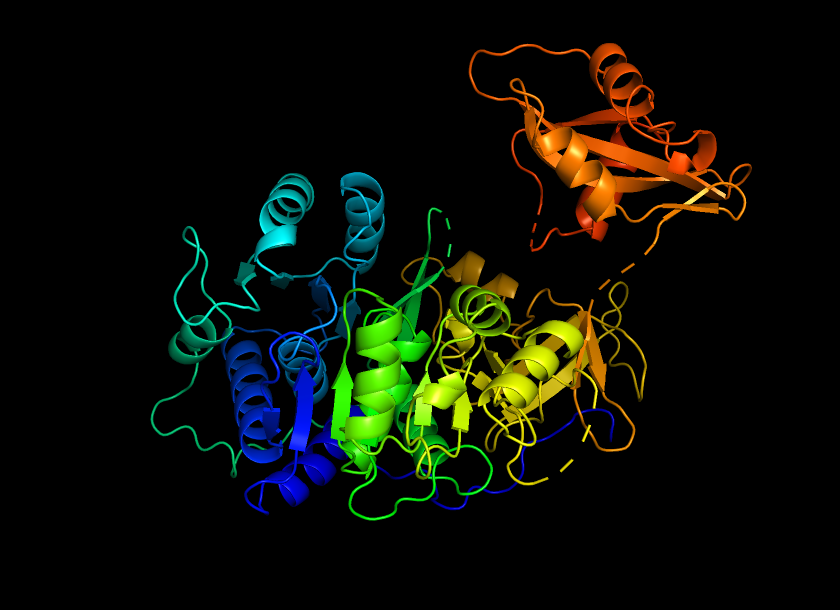
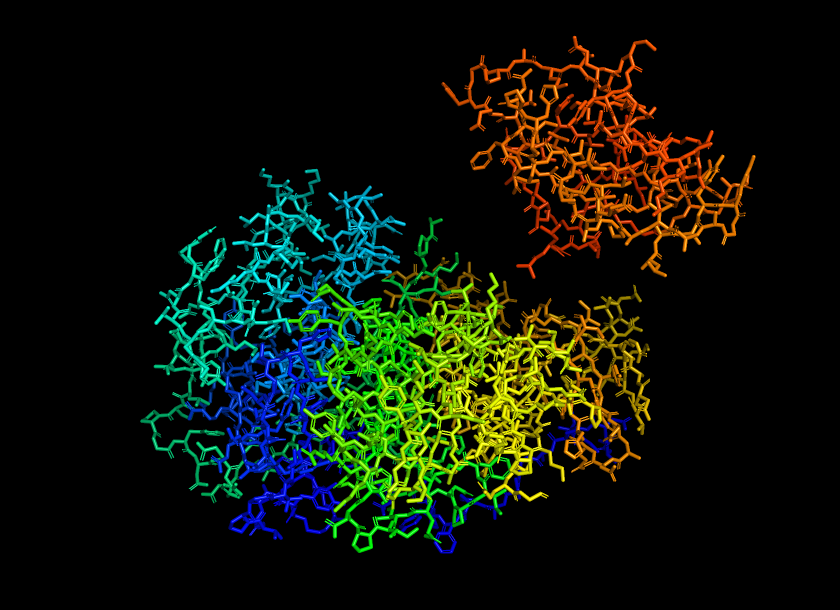
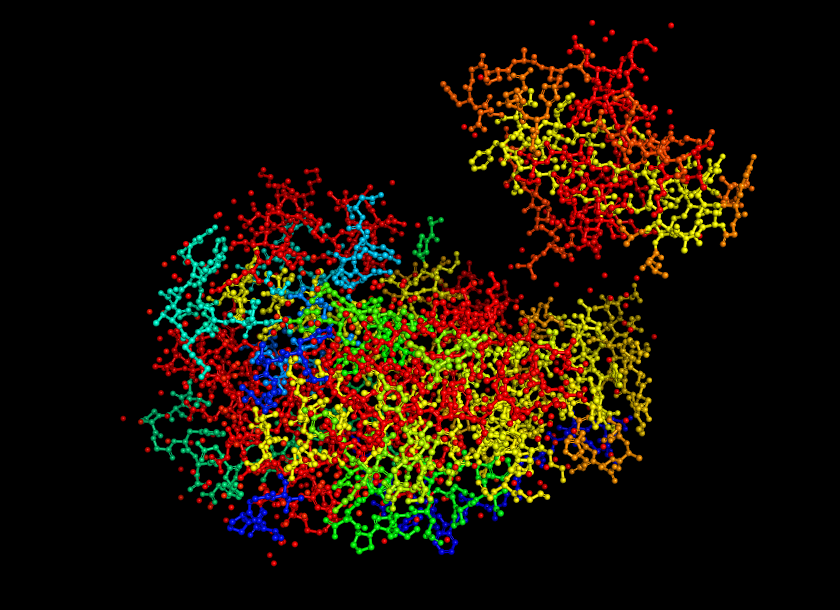
Color the protein by secondary structure. Does it have more helices or sheets?
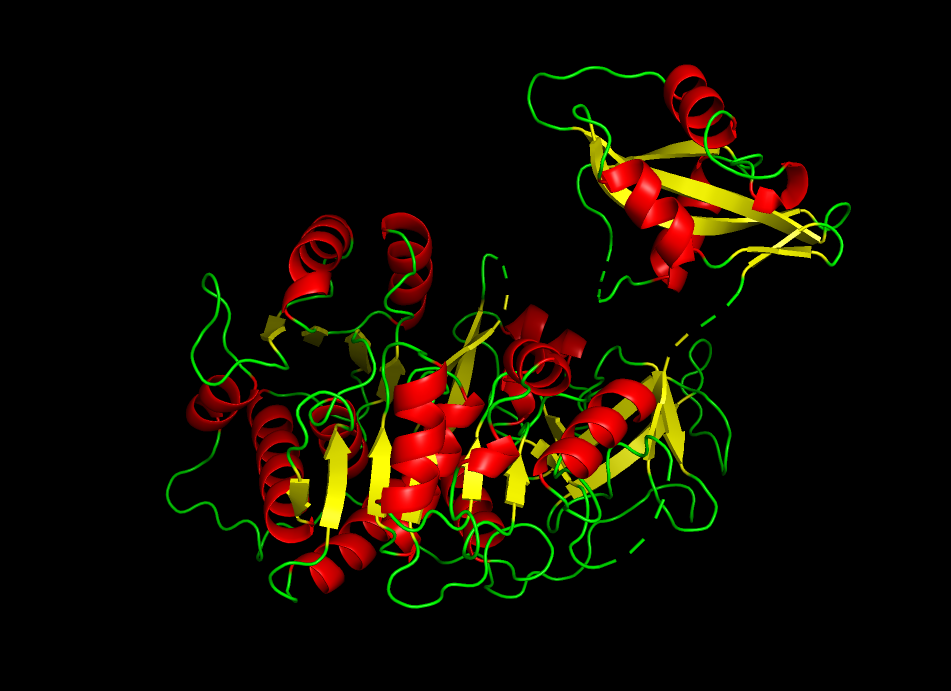
The protein shows a predominance in alpha helix (red) compared to beta helix (green). This indicates that firefly luciferase is mainly an alpha-helical protein with a smaller portion of beta-sheet structures
Color the protein by residue type. What can you tell about the distribution of hydrophobic vs hydrophilic residues?
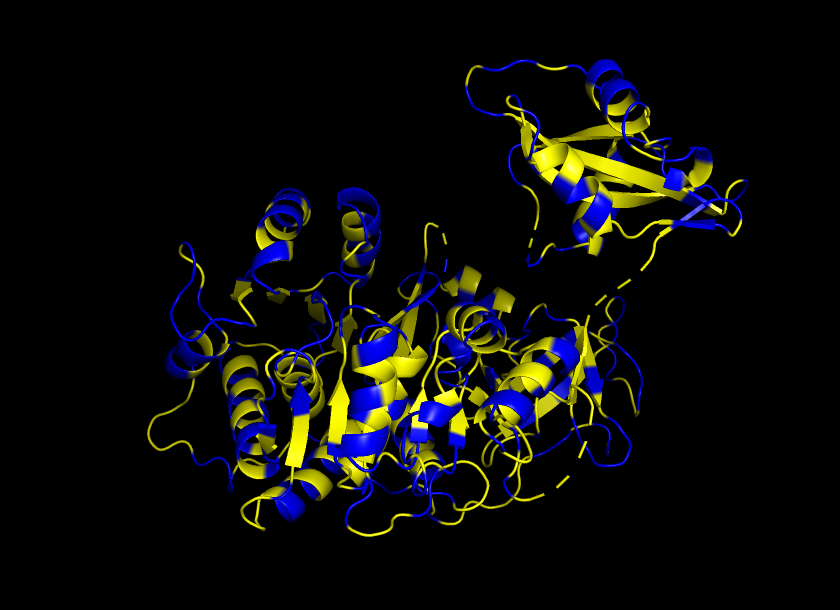
It is shown that this particular protein/enzyme, which operates in an aqueous environment, has an exterior with hydrophilic residues as protagonists and its core with hydrophobic residues.
Visualize the surface of the protein. Does it have any “holes” (aka binding pockets)?
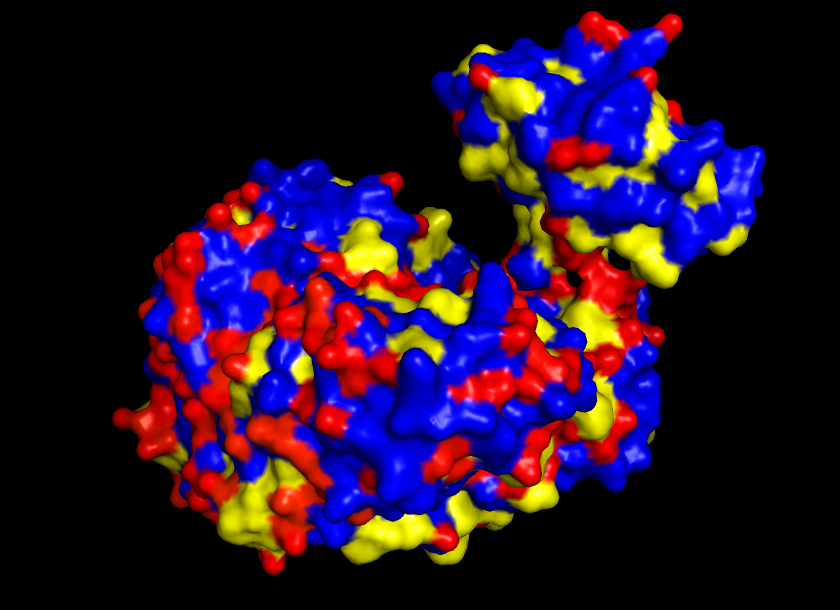
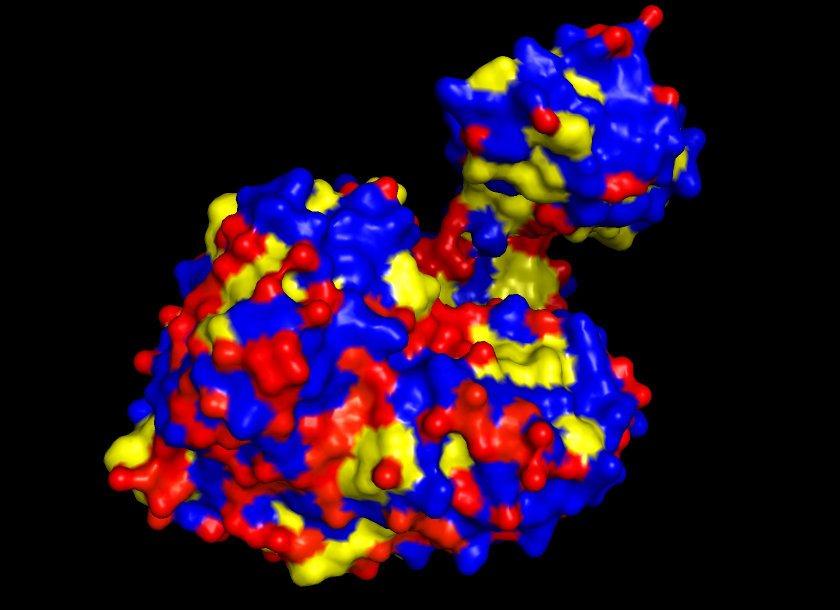
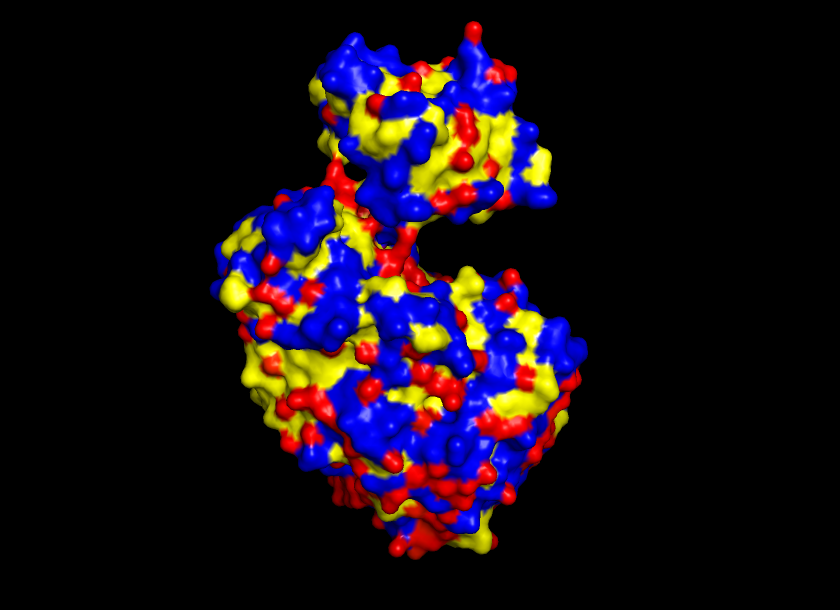
Yes, the protein surface shows a variety of pockets. One with a predominant size, and others that are small. It is precisely in the big pocket that the ATP binds with the D-luciferine to form Luciferil-AMP and then binds together with oxygen molecules that finally form oxyluciferine and light.
Part C. Using ML-Based Protein Design Tools
C1. Protein Language Modeling
Deep Mutational Scans
a. Use ESM2 to generate an unsupervised deep mutational scan of your protein based on language model likelihoods. b. Can you explain any particular pattern? (choose a residue and a mutation that stands out). c. (Bonus) Find sequences for which we have experimental scans, and compare the prediction of the language model to experiment.
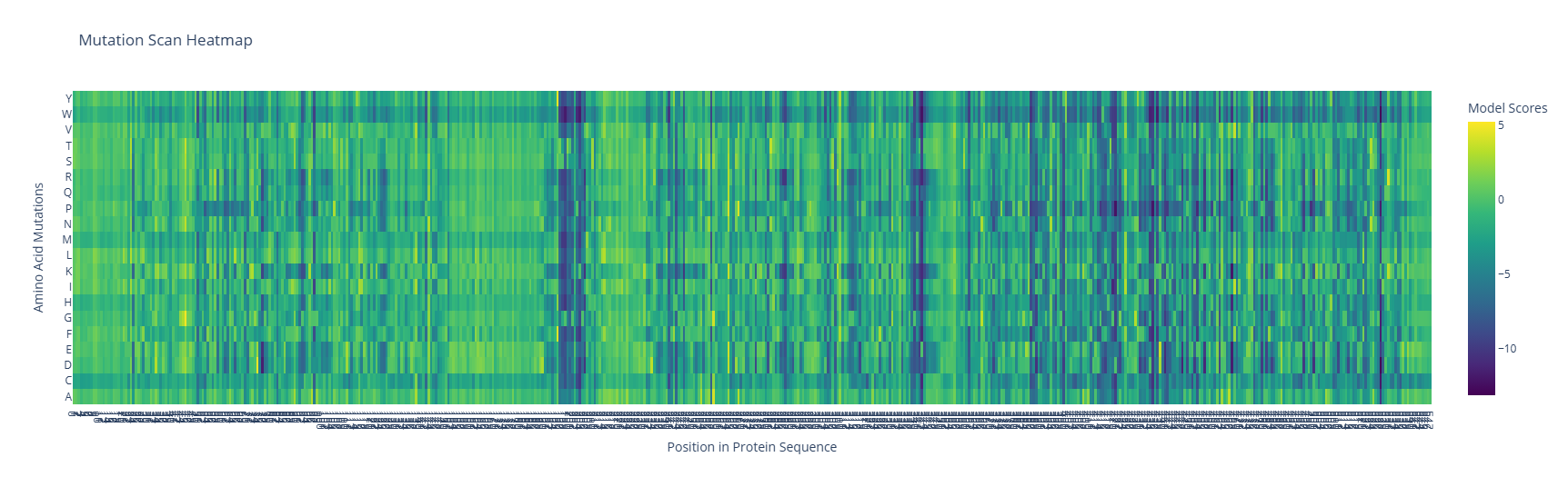
The map shows a large number of possible mutations, although two main regions should not be changed because the protein could collapse; those regions are shown as two columns of dark blue. Also, three subtle rows show color consistency corresponding to W, M, and C.
Latent Space Analysis
a. Use the provided sequence dataset to embed proteins in reduced dimensionality. b. Analyze the different formed neighborhoods: do they approximate similar proteins? c. Place your protein in the resulting map and explain its position and similarity to its neighbors.
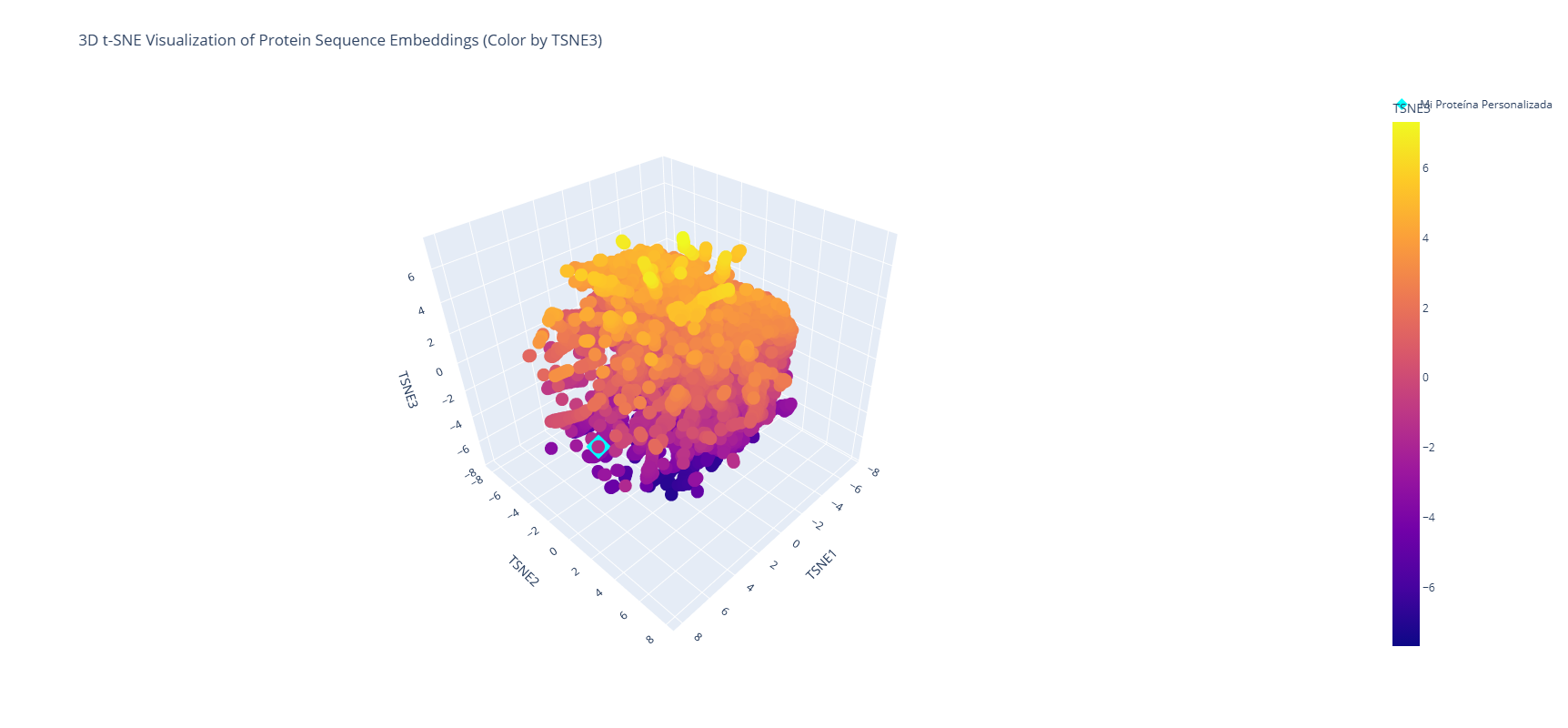
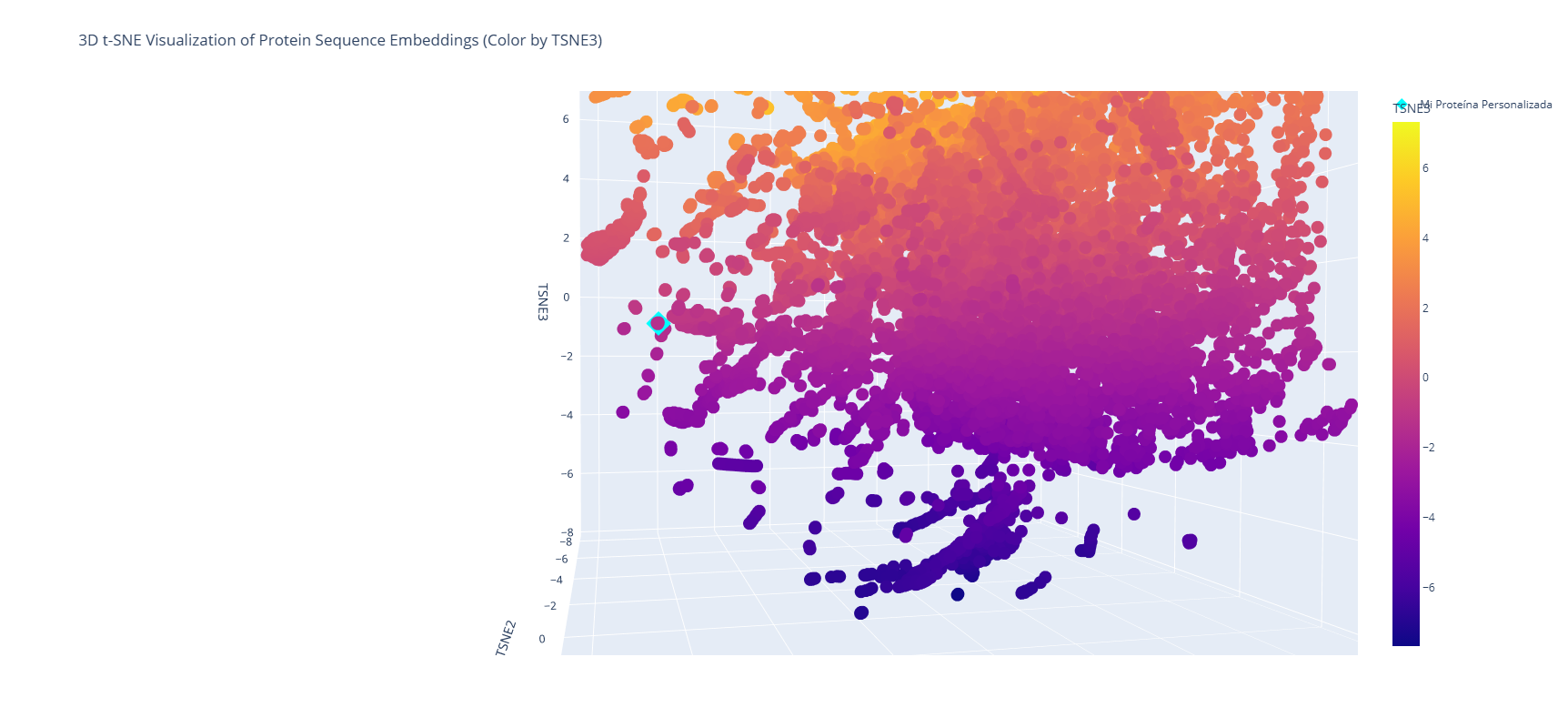
It is shown that near the analyzed protein (Firefly Luciferase - Photinus pyralis) is located the Luciferase Luciola Cruciata, a protein produced by another type of firefly. The first one, PP, is from North America, while the second one, LC, is from Japan. The main difference is the geographical location and its molecular composition, which is expressed in a slightly different type of color, and the stability of the enzyme. Although both proteins use D-Luciferin and ATP to produce light, PP Luciferase is widely used in biotech as a reporter gene. In contrast, LC Luciferase is used to understand how active-site residues interact with the substrate.
C2. Protein Folding
Fold your protein with ESMFold. Do the predicted coordinates match your original structure? Try changing the sequence, first try some mutations, then large segments. Is your protein structure resilient to mutations?
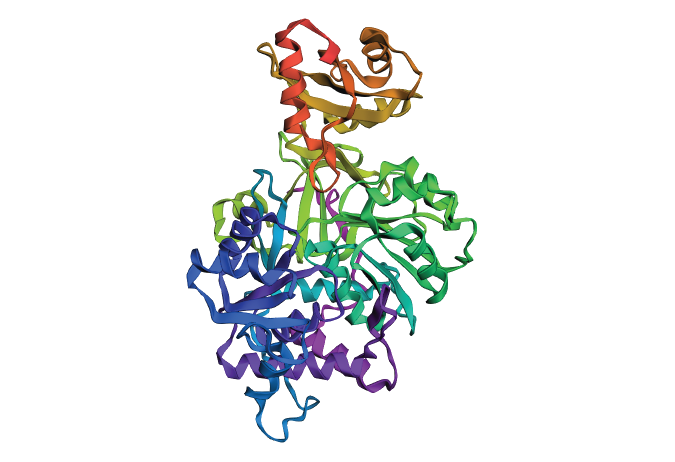
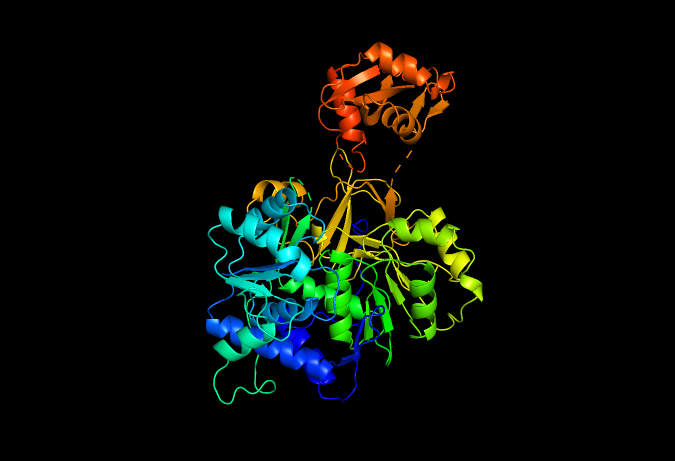
When folded with ESMFold, the protein shows an almost identical structure to the original one, but when given some mutations, it presents a few changes, not very radical ones, but a few anomalies, meaning that the protein is resilient in a high percentage.
C3. Protein Generation
Inverse-Folding a protein Let’s now use the backbone of your chosen PDB to propose sequence candidates via ProteinMPNN Analyze the predicted sequence probabilities and compare the predicted sequence vs the original one. Input this sequence into ESMFold and compare the predicted structure to your original.
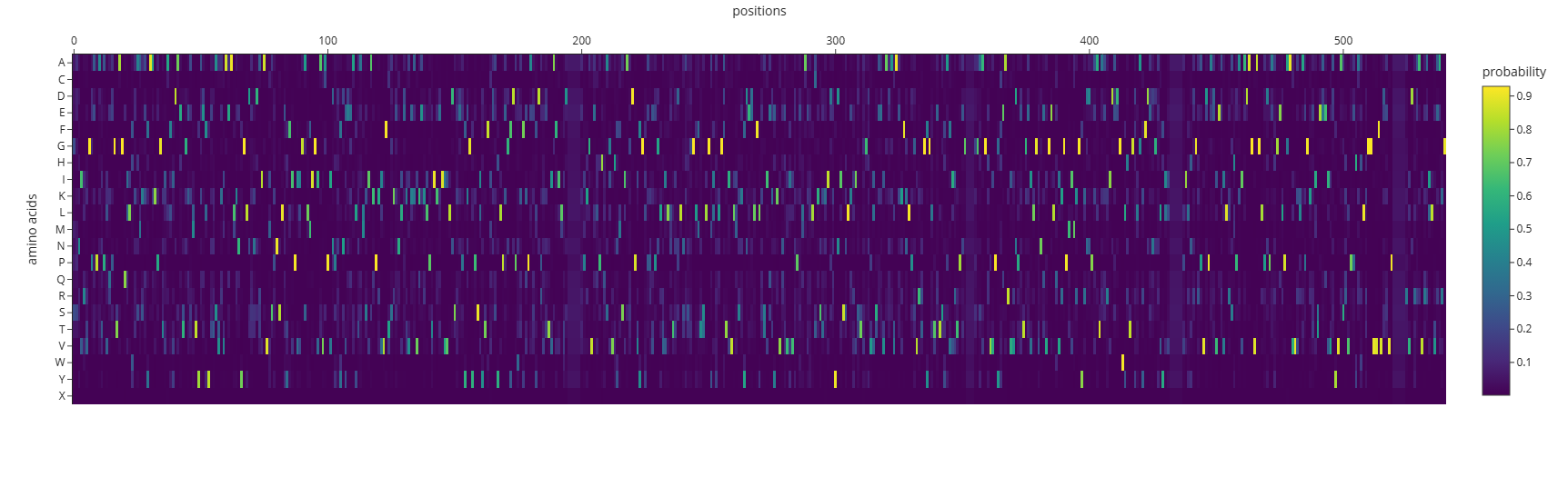
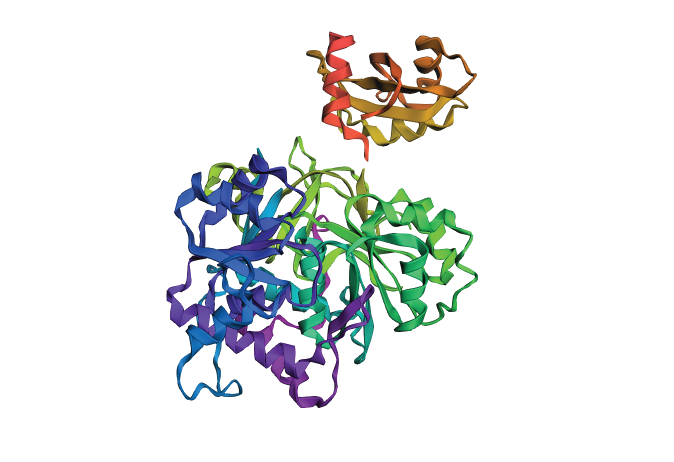
Even though the predicted structure has a completely different type of amino acid distribution, the structure remains the same. This is why the protein shown in 3D is very similar to the original in its alpha- and beta-structures. The backbone is not altered, nor is the logic of the distribution of certain types of amino acids, either.
Part D. Group Brainstorm on Bacteriophage Engineering
- Find a group of ~3–4 students
- Read through the Phage Reading material listed under “Reading & Resources” below.
- Review the Bacteriophage Final Project Goals for engineering the L Protein:
- Increased stability (easiest)
- Higher titers (medium)
- Higher toxicity of lysis protein (hard)
- Brainstorm Session
- Choose one or two main goals from the list that you think you can address computationally (e.g., “We’ll try to stabilize the lysis protein,” or “We’ll attempt to disrupt its interaction with E. coli DnaJ.”).
- Write a 1-page proposal (bullet points or short paragraphs) describing:
- Which tools/approaches from recitation you propose using (e.g., “Use Protein Language Models to do in silico mutagenesis, then AlphaFold-Multimer to check complexes.”).
- Why do you think those tools might help solve your chosen sub-problem?
- Name one or two potential pitfalls (e.g., “We lack enough training data on phage–bacteria interactions.”).
- Include a schematic of your pipeline.
- This resource may be useful: HTGAA Protein Engineering Tools
- Each individually put your plan on your HTGAA website
- Include your group’s short plan for engineering a bacteriophage