Week 2 HW: DNA Read Write and Edit
✨Part 1: Benchling & In silico Gel Art
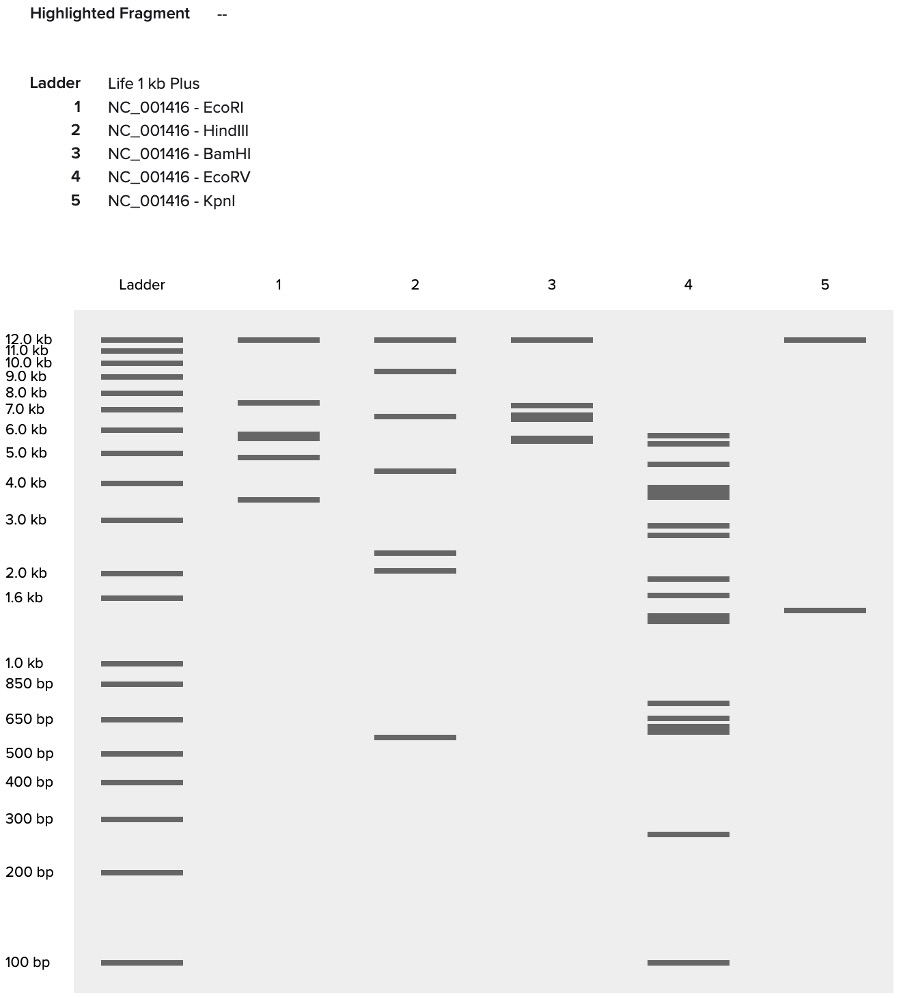
I simulated a restriction digest on λ DNA in Benchling using enzymes like EcoRI, HindIII, and BamHI, EcoRV, Kpnl. By comparing the band patterns, I could visualize how different enzymes cut the DNA into fragments of varying sizes. This simulation helped me understand how we verify DNA fingerprints before moving to synthesis.

✨ Part 3: DNA Design Challenge
3.1. Choose your protein
My Choice: For this assignment, I chose Lysostaphin, a glycylglycine endopeptidase enzyme. This protein is naturally produced by Staphylococcus simulans to kill rival bacteria.
Why I chose it:
My background is in dentistry and tissue engineering, where peri-implantitis (infection around dental implants) is a critical failure mode. These infections are often caused by antibiotic-resistant Staphylococcus aureus (MRSA) forming biofilms on the titanium surface. Lysostaphin is capable of slicing through the cell wall of S. aureus, destroying the biofilm effectively where traditional antibiotics fail. It represents a potential “biological scalpel” for saving failing implants.
Sequence:
Using UniProt, I obtained the amino acid sequence for Lysostaphin:
sp|P10547|LSTP_STASI Lysostaphin OS=Staphylococcus simulans OX=1286 GN=lss PE=1 SV=2 MKKTKNNYYTRPLAIGLSTFALASIVYGGIQNETHASEKSNMDVSKKVAEVETSKAPVEN TAEVETSKAPVENTAEVETSKAPVENTAEVETSKAPVENTAEVETSKAPVENTAEVETSK APVENTAEVETSKAPVENTAEVETSKAPVENTAEVETSKAPVENTAEVETSKAPVENTAE VETSKAPVENTAEVETSKAPVENTAEVETSKAPVENTAEVETSKAPVENTAEVETSKALV QNRTALRAATHEHSAQWLNNYKKGYGYGPYPLGINGGMHYGVDFFMNIGTPVKAISSGKI VEAGWSNYGGGNQIGLIENDGVHRQWYMHLSKYNVKVGDYVKAGQIIGWSGSTGYSTAPH LHFQRMVNSFSNSTAQDPMPFLKSAGYGKAGGTVTPTPNTGWKTNKYGTLYKSESASFTP NTDIITRTTGPFRSMPQSGVLKAGQTIHYDEVMKQDGHVWVGYTGNSGQRIYLPVRTWNK STNTLGVLWGTIK
3.2. Reverse Translate
Using the online resource at https://www.bioinformatics.org/, I converted the amino acid sequence(taken from https://www.uniprot.org) of the Lysostaphin protein back into its potential DNA sequence. This technique follows the Central Dogma of Molecular Biology, which outlines the flow of genetic information from DNA to RNA and finally to protein. By reversing this sequence, the tool creates a logical nucleotide chain capable of producing that specific protein.

Converted Sequence:
reverse translation of Untitled to a 1479 base sequence of most likely codons. atgaaaaaaaccaaaaacaactattatacccgcccgctggcgattggcctgagcaccttt gcgctggcgagcattgtgtatggcggcattcagaacgaaacccatgcgagcgaaaaaagc aacatggatgtgagcaaaaaagtggcggaagtggaaaccagcaaagcgccggtggaaaac accgcggaagtggaaaccagcaaagcgccggtggaaaacaccgcggaagtggaaaccagc aaagcgccggtggaaaacaccgcggaagtggaaaccagcaaagcgccggtggaaaacacc gcggaagtggaaaccagcaaagcgccggtggaaaacaccgcggaagtggaaaccagcaaa gcgccggtggaaaacaccgcggaagtggaaaccagcaaagcgccggtggaaaacaccgcg gaagtggaaaccagcaaagcgccggtggaaaacaccgcggaagtggaaaccagcaaagcg ccggtggaaaacaccgcggaagtggaaaccagcaaagcgccggtggaaaacaccgcggaa gtggaaaccagcaaagcgccggtggaaaacaccgcggaagtggaaaccagcaaagcgccg gtggaaaacaccgcggaagtggaaaccagcaaagcgccggtggaaaacaccgcggaagtg gaaaccagcaaagcgccggtggaaaacaccgcggaagtggaaaccagcaaagcgctggtg cagaaccgcaccgcgctgcgcgcggcgacccatgaacatagcgcgcagtggctgaacaac tataaaaaaggctatggctatggcccgtatccgctgggcattaacggcggcatgcattat ggcgtggatttttttatgaacattggcaccccggtgaaagcgattagcagcggcaaaatt gtggaagcgggctggagcaactatggcggcggcaaccagattggcctgattgaaaacgat ggcgtgcatcgccagtggtatatgcatctgagcaaatataacgtgaaagtgggcgattat gtgaaagcgggccagattattggctggagcggcagcaccggctatagcaccgcgccgcat ctgcattttcagcgcatggtgaacagctttagcaacagcaccgcgcaggatccgatgccg tttctgaaaagcgcgggctatggcaaagcgggcggcaccgtgaccccgaccccgaacacc ggctggaaaaccaacaaatatggcaccctgtataaaagcgaaagcgcgagctttaccccg aacaccgatattattacccgcaccaccggcccgtttcgcagcatgccgcagagcggcgtg ctgaaagcgggccagaccattcattatgatgaagtgatgaaacaggatggccatgtgtgg gtgggctataccggcaacagcggccagcgcatttatctgccggtgcgcacctggaacaaa agcaccaacaccctgggcgtgctgtggggcaccattaaa
3.3. Codon optimization
- Why do we optimize codons?
I need to ensure my DNA “reads” fluently in the host organism. If the codons are rare in the host, protein production will stall. Optimization replaces these rare codons with the host’s preferred ones without changing the final protein structure.
- Which organism did you choose and why?
I chose Escherichia coli (E. coli) for codon optimization. While my final application is for dental patients, E. coli is the industrial standard for manufacturing proteins. By optimizing for E. coli, I can grow large vats of bacteria, induce them to produce Lysostaphin, and then purify the enzyme to be applied as a dental gel or coating for implants.
Optimization Result:
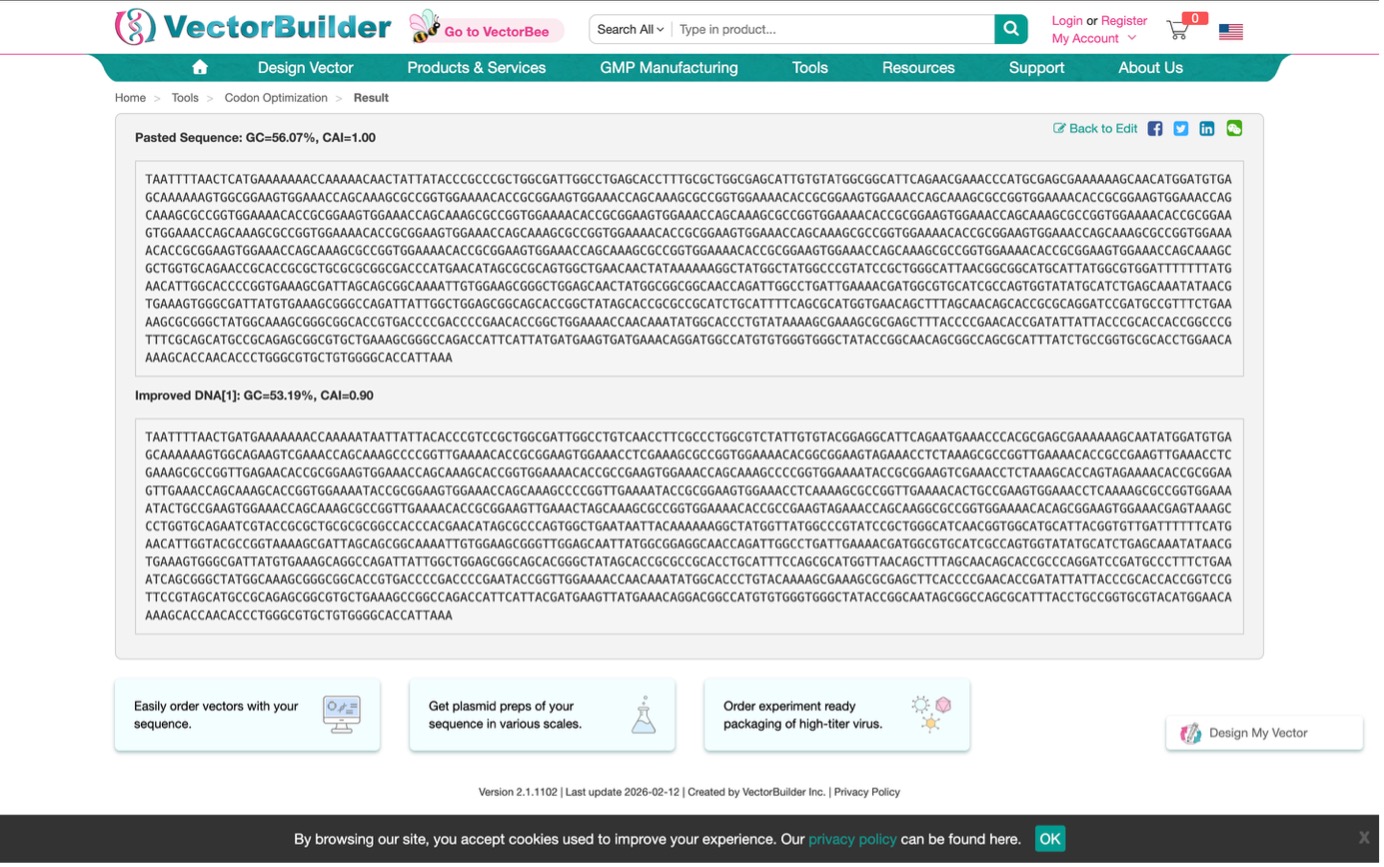

3.4. You have a sequence! Now what?
Now that I have the optimized DNA sequence, the goal is recombinant protein production to create a therapeutic solution for peri-implantitis.
- Cloning
I will insert the optimized Lysostaphin gene into an expression vector (plasmid). This plasmid acts as the delivery vehicle, containing a strong promoter that signals the host cell to begin producing the protein.
- Transformation
The recombinant plasmid is put into $E. coli$ bacteria. This is achieved through a process called transformation (such as heat-shock), which allows the bacterial cells to take up the foreign DNA and host it within their own systems.
- Expression
The bacteria act as biological factories, following the Central Dogma of Molecular Biology. The $E. coli$ cells read the optimized DNA instructions to produce mRNA via transcription, which is then translated into the Lysostaphin Protein. Because the codons were optimized for $E. coli$ (K12), the translation process is highly efficient with a high protein yield.
- Purification Finally, I will extract the protein from the bacterial culture. Through a series of filtration and chromatography steps, the Lysostaphin is isolated from other bacterial proteins. The result is a pure protein that can be formulated into a bioactive gel designed to target and eliminate $Staphylococcus$ biofilms in patients with peri-implantitis.
3.5. [Optional] How does it work in nature/biological systems?
- Describe how a single gene codes for multiple proteins at the transcriptional level.
A gene is first transcribed into a long RNA molecule called pre-mRNA. This pre-mRNA contains both coding regions (exons) and non-coding regions (introns). Through a process called Alternative Splicing, the cell can cut out the introns and stitch the exons together in different combinations. Just like editing a movie scene in different ways, different combinations of exons create different final mRNA molecules. $$Different \ mRNA \ variants \rightarrow Different \ Proteins$$ Because the mRNA sequence changes, the resulting amino acid sequence changes too. This allows a single gene to code for multiple different protein isoforms, maximizing the efficiency of the genome.
- Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein!
In nature, the enzyme RNA Polymerase reads the DNA template strand and synthesizes a single-stranded RNA molecule based on base complementarity.
• A pairs with U (Uracil replaces Thymine in RNA).
• T pairs with A.
• G pairs with C.
• C pairs with G.
After transcription, the Ribosome reads the mRNA in groups of three nucleotides called codons. Each codon corresponds to one specific amino acid.
Alignment for Lysostaphin (Start of Sequence):
Here is the flow of information for the first 6 amino acids of my Lysostaphin protein (MTTTPD…).
• DNA (Coding Strand): ATG ACC ACC ACC CCG GAT
• mRNA (Transcription): AUG ACC ACC ACC CCG GAU
• Protein (Translation): M T T T P D
Key:
• M (Methionine): The “Start” signal.
• T (Threonine): A polar amino acid.
• P (Proline): Adds structural rigidity.
• D (Aspartic Acid): Negatively charged.
Part 4: Prepare a Twist DNA Synthesis Order
I created a new sequence in Benchling named Lysostaphin_e.coli. I combined my optimized gene with the standard parts required for E. coli expression:
Promoter (BBa_J23106):
TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGC
RBS (BBa_B0034):
CATTAAAGAGGAGAAAGGTACC
Start Codon:
ATG
Coding Sequence:
ATGAAAAAAACGAAAAACAATTACTATACCCGCCCGCTGGCCATTGGCCTGAGCACTTTTGCGCTGGCGAGCATCGTGTACGGCGGCATTCAGAACGAAACCCATGCGAGCGAAAAAAGCAATATGGATGTAAGCAAAAAAGTGGCGGAAGTTGAAACCAGCAAAGCGCCGGTCGAAAACACCGCGGAAGTGGAAACTAGCAAAGCGCCGGTCGAAAACACCGCCGAAGTGGAAACCAGCAAAGCGCCGGTTGAAAACACCGCCGAAGTGGAGACCAGCAAAGCGCCGGTGGAAAATACCGCCGAAGTAGAAACCAGCAAAGCCCCGGTGGAAAATACCGCGGAAGTGGAGACCTCAAAAGCGCCGGTTGAAAACACCGCGGAAGTGGAAACGAGCAAAGCACCGGTGGAGAATACCGCGGAAGTGGAAACCAGCAAAGCGCCGGTGGAAAATACCGCGGAAGTGGAAACGAGCAAAGCCCCAGTTGAAAATACGGCCGAGGTGGAAACCAGCAAAGCGCCGGTGGAAAACACCGCCGAAGTTGAAACCTCCAAAGCCCCGGTTGAAAATACCGCGGAAGTAGAAACCTCGAAAGCACCGGTGGAAAACACCGCCGAAGTGGAAACCTCAAAAGCCCCGGTGGAAAACACCGCGGAAGTTGAAACCTCTAAAGCGCCGGTGGAAAATACGGCGGAAGTGGAAACCAGCAAAGCCCTGGTCCAGAACCGCACCGCGCTGCGCGCGGCAACCCATGAACATAGCGCGCAGTGGCTGAATAACTACAAAAAAGGCTATGGCTATGGCCCGTATCCGCTGGGCATTAATGGCGGCATGCATTATGGTGTCGACTTTTTCATGAACATCGGCACCCCGGTTAAAGCGATTTCGAGCGGTAAAATCGTGGAAGCCGGCTGGAGCAACTACGGCGGCGGCAACCAGATTGGTCTGATTGAAAATGATGGCGTGCATCGCCAGTGGTACATGCATCTGAGCAAATACAACGTCAAAGTGGGTGATTATGTGAAAGCAGGTCAGATTATTGGCTGGAGCGGCAGCACCGGCTACAGCACCGCACCGCACCTGCATTTCCAGCGTATGGTGAATAGCTTCAGCAATAGCACCGCGCAGGATCCGATGCCGTTTCTGAAATCAGCGGGCTATGGCAAAGCGGGCGGCACCGTGACCCCGACCCCGAATACCGGCTGGAAAACCAACAAATATGGCACCCTGTATAAAAGCGAAAGCGCGAGCTTTACCCCGAACACCGATATCATTACCCGCACCACCGGCCCGTTCCGCAGCATGCCGCAGTCAGGCGTGCTGAAAGCGGGCCAGACCATTCATTATGATGAAGTGATGAAACAGGATGGCCATGTGTGGGTGGGTTATACCGGCAACTCGGGCCAGCGCATCTACCTGCCGGTGCGCACCTGGAACAAAAGCACCAACACCCTGGGTGTACTGTGGGGTACCATTAAA
7x His Tag:
CATCACCATCACCATCATCAC
Stop Codon:
TAA
Terminator (BBa_B0015):
CCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTG TCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA
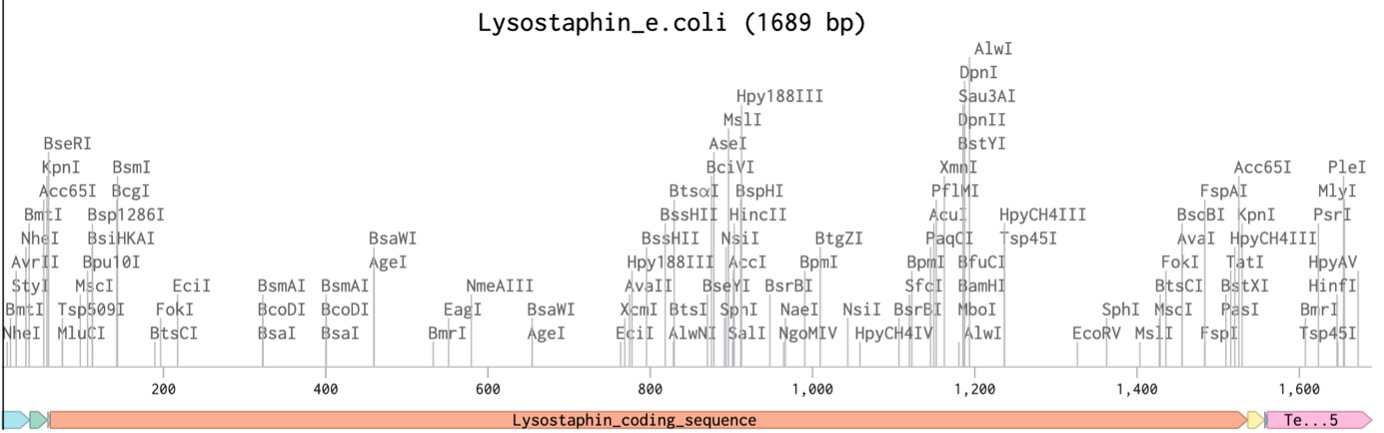

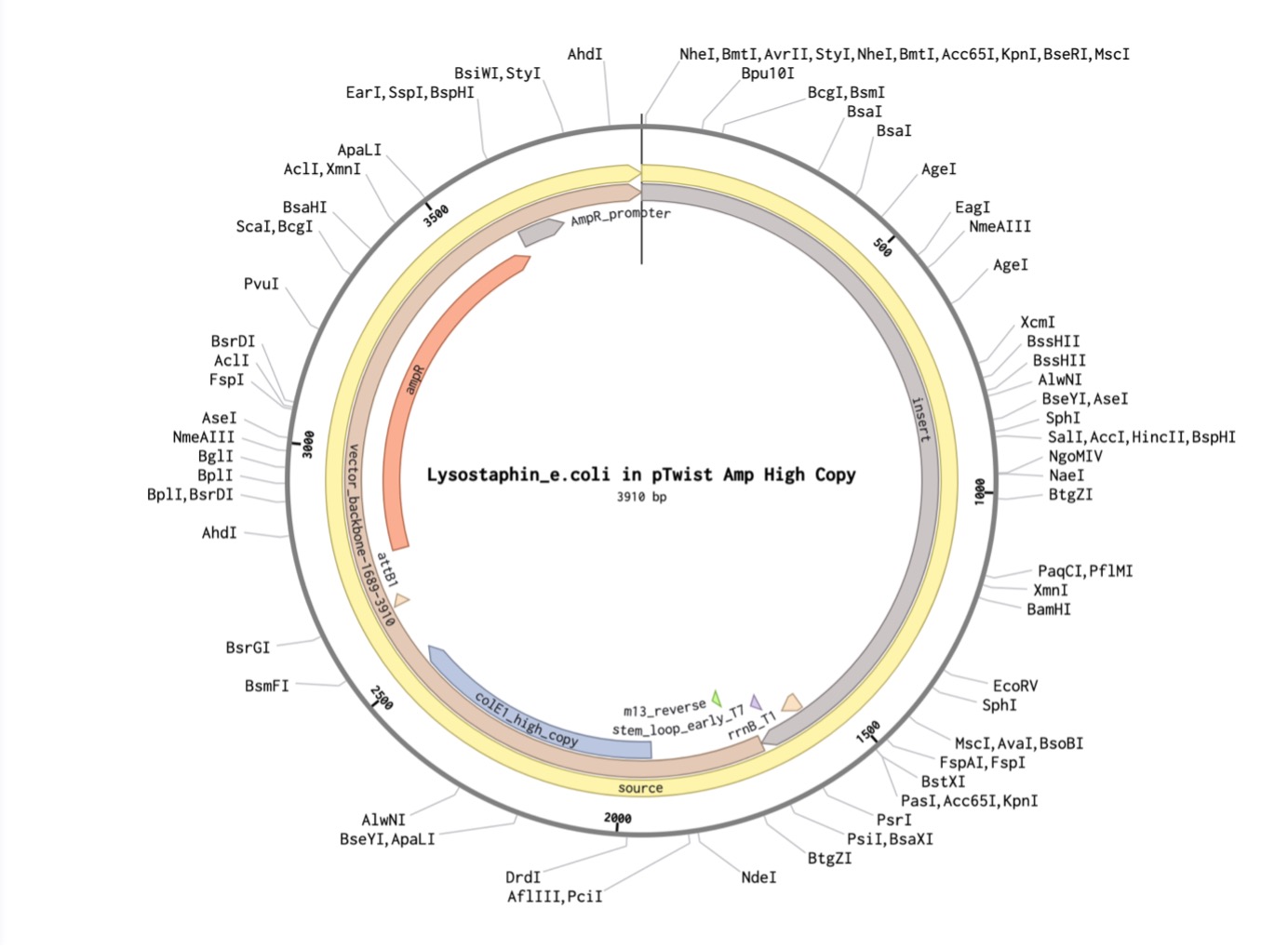

Part 5: DNA Read/Write/Edit
5.1 DNA Read
What to Read: I would sequence the biofilm microbiome found in the pockets of failing dental implants.
Why:
Current treatment for peri-implantitis is often “blind” mechanical cleaning. By sequencing the DNA of the infection site, we can identify exactly which pathogens are present (e.g., P. gingivalis vs. S. aureus) and detect if they carry Antibiotic Resistance Genes (AMR). This allows for precision dentistry—choosing the right treatment rather than guessing.
Technology:
I would use the Oxford Nanopore MinION. • Reason: It is portable and rapid. I could theoretically bring it into a dental clinic, swab an implant, and get sequencing data in real-time to guide surgery. • Process: Extract DNA from plaque $\rightarrow$ Load into MinION $\rightarrow$ Nanopore reads electrical signals of DNA strands $\rightarrow$ Output is the pathogen profile.
5.2 DNA Write
What to Write:
I want to synthesize the gene for Lysostaphin (as designed in Part 3).
Why:
Nature provided S. simulans with this weapon, but we need to mass-produce it to use it as a medicine. By writing (synthesizing) this DNA, we can create a pure, high-concentration anti-biofilm agent that dissolves the cell walls of MRSA, saving titanium implants that would otherwise need to be removed.
Technology:
I would use Twist Bioscience silicon-based synthesis. • Reason: It allows me to order the exact “Expression Cassette” I designed, ensuring the sequence is perfect for my E. coli factories.
5.3 DNA Edit
What to Edit:
I would use CRISPR to edit commensal oral bacteria (like Streptococcus salivarius) to naturally secrete Lysostaphin.
Why:
Instead of applying a gel, we could introduce a “guardian bacteria” into the patient’s mouth. This edited bacteria would live on the gums and constantly produce small amounts of Lysostaphin, preventing the dangerous S. aureus from ever forming a biofilm on the implant in the first place.