Week 3 HW: Lab Automation
Week 3 – Lab Automation
✨ Week 3 - Homework ✨
You can view my Automation Art design here: Opentrons Art Link
After creating this shell pattern using Opentrons Art, I duplicated the provided Colab notebook to develop a Python protocol. To program the Opentrons robot to physically recreate the artwork on a plate, I systematically entered the coordinate data from my design step-by-step into the script. Once the protocol was complete, it successfully generated the images shown below.
Digital Shell Design
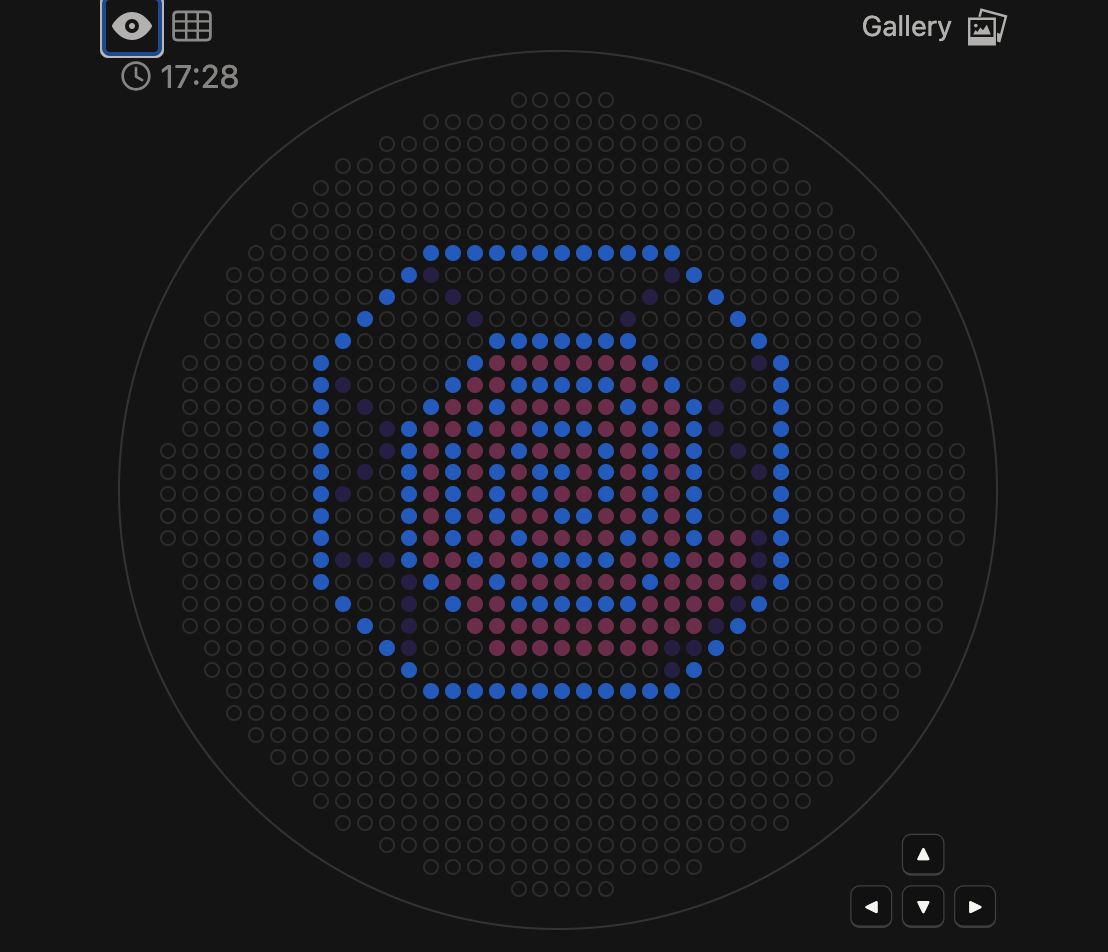

✨ Post-Lab Questions ✨
1) Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Article Title: An Automation Workflow for High-Throughput Manufacturing and Analysis of Scaffold-Supported 3D Tissue Arrays
Authors: Ruonan Cao, Nancy T. Li, Simon Latour, Jose L. Cadavid, Cassidy M. Tan, Ari Forman, Hartland W. Jackson, Alison P. McGuigan
Year: 2023
DOI: 10.1002/adhm.202202422
Article 1
This paper tackles a real bottleneck in advanced 3D culture: patient-derived organoids and complex co-cultures are powerful, but hard to scale and hard to analyze at single-cell resolution when manufacturing and handling are manual. The authors focus on the SPOT platform (Scaffold-supported Platform for Organoid-based Tissues), which generates flat, thin, dimensionally controlled microtissues in 96- and 384-well plate formats compatible with longitudinal imaging—yet historically limited by manual fabrication.
What’s automated with Opentrons OT-2 (and what makes it novel):
Automated 3D microtissue manufacturing (seeding): They use the Opentrons OT-2 to dispense a cell–gel mixture into 96/384-SPOT plates and optimize the process so automated manufacturing is comparable to manual consistency.
Temperature control + custom hardware for reliability: Two temperature modules set to 4 °C keep the SPOT plate and cell-gel cold during seeding, and a custom aluminum plate improves support and heat conduction for the thin plate—showing how automation often needs small mechanical/thermal design choices to work well.
]Automation beyond seeding (screening + single-cell endpoints): The OT-2 also supports drug/reagent addition and culture maintenance, and it automates gel digestion to recover single cells for high-throughput flow cytometry.
Multiplexed CyTOF enablement: A particularly strong “novel application” angle is that OT-2 is used to generate a barcode master plate and automate parts of the barcoding/washing/pooling workflow to reduce manual errors—enabling scalable CyTOF proteomic readouts.
Proof-of-value biology: They generate 3D complex tissues with different tumor/stromal ratios and show the workflow can incorporate primary patient-derived organoids, supporting scalable, patient-relevant screening and analysis.
2) Write a description about what you intend to do with automation tools for your final project.
Across all three ideas, the Opentrons OT-2 is my core “execution engine” for repeatable, programmable liquid handling—reducing variability, scaling to multi-sample workflows, and producing clean run logs/plate maps. Where formats aren’t standard labware, I’d use custom 3D-printed holders.
Idea 1 — Automated Seeding of Patient-Specific Bone Scaffolds
Goal: Improve cell distribution and viability deep inside porous bone scaffolds by replacing static pipetting with automated, repeatable dynamic “drip-seeding.”
What I would automate on the OT-2
- A custom 3D-printed scaffold holder mounting multiple scaffolds on the OT-2 deck.
- A timed protocol dispensing cell suspension (e.g., MSCs) + osteogenic media cues across scaffolds in multi-pass patterns.
- Optional scheduled media refresh + standardized sampling for assays.
Example pseudocode (conceptual)
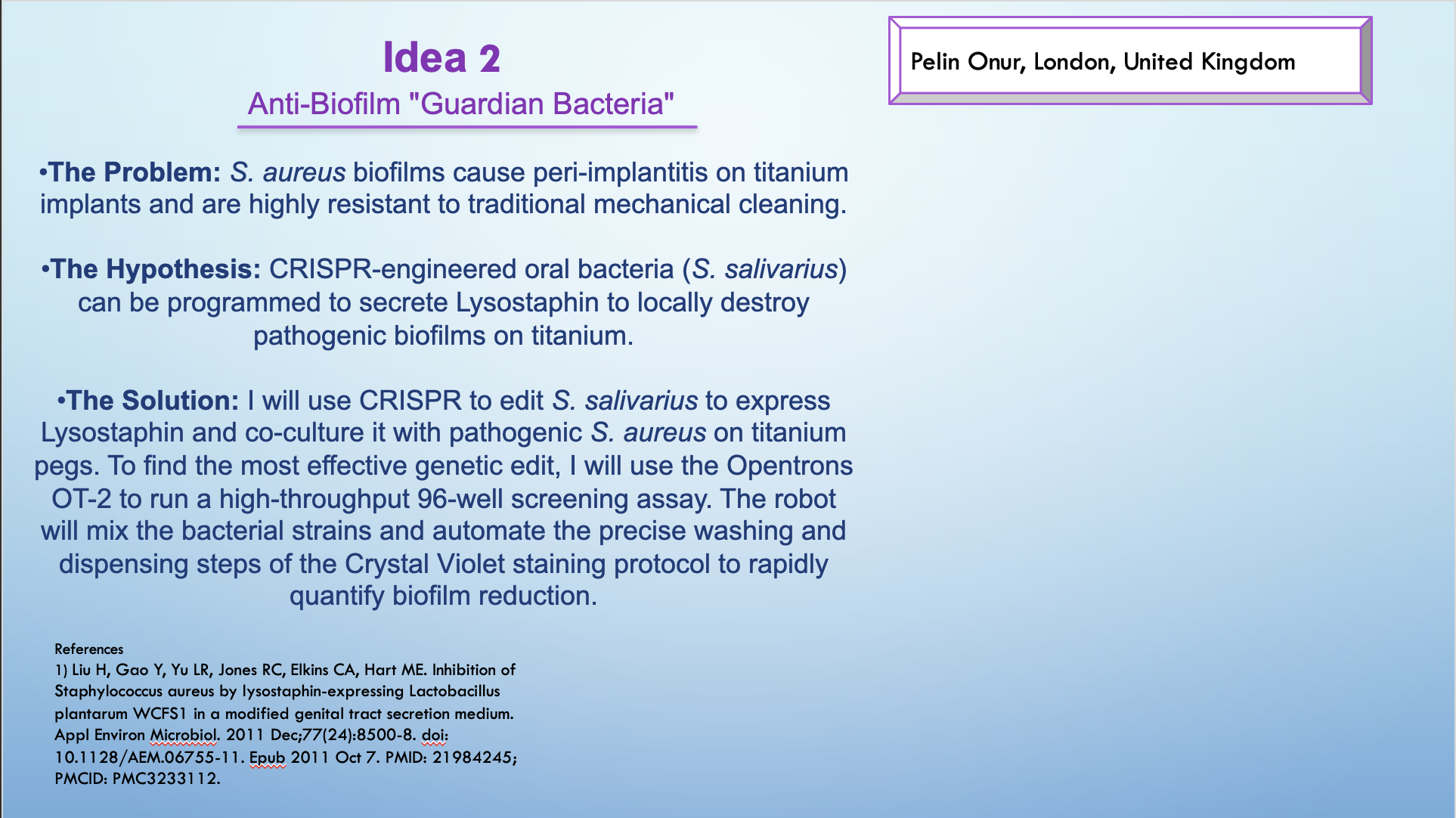
Idea 2 — Anti-Biofilm “Guardian Bacteria” (high-throughput screening)
Goal: Run a high-throughput anti-biofilm screen on titanium-relevant surfaces using a plate-based assay format.
What I would automate on the OT-2
- A 96-well screening layout (controls + variants + replicates).
- Automated mixing, dispensing, wash steps, and readout reagent handling.
- Standardized timing + plate map + run log for comparability.
Example pseudocode (conceptual)
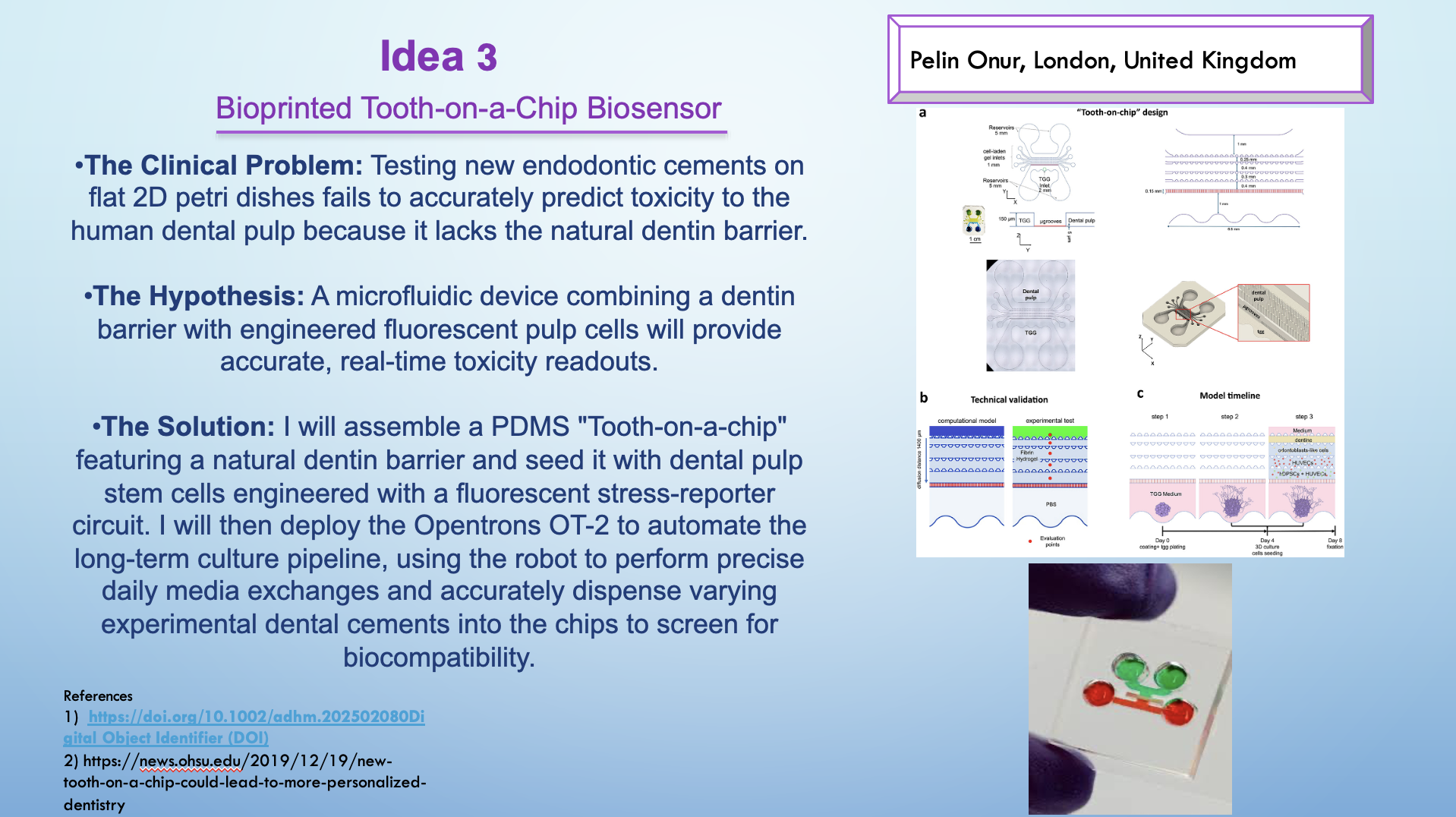
Idea 3 — Bioprinted Tooth-on-a-Chip Biosensor (automated long-term culture + exposure)
Goal: Improve dental material testing realism using a chip that includes a dentin barrier + engineered reporter pulp cells for real-time toxicity/biocompatibility readouts.
What I would automate on the OT-2
- Daily/recurring media exchange across multiple chips.
- Controlled dosing/exposure scheduling for different materials.
- Optional sampling workflow into plates for downstream measurements.
Example pseudocode (conceptual)


