Week 9 HW: Cell Free Systems
HTGAA Week 9: Cell-Free Systems
Part A: General and Lecturer-Specific Questions
1. Explain the main advantages of cell-free protein synthesis (CFPS) over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
The biggest advantage I see is that CFPS turns “biology” into “chemistry.” In traditional in vivo systems, the cell membrane is a wall that prevents us from easily tweaking the internal environment. In a cell-free setup, I have an “open” system. I can add non natural amino acids, adjust salt concentrations (Mg²⁺ and K⁺) in real time, or even add detergents to help fold tricky proteins.
Two cases where this is a game changer:
- Toxic Protein Production: If I’m trying to express an antimicrobial peptide designed to kill Streptococcus mutans, a live E. coli host would likely die before it can produce a high yield. In CFPS, the “host” is already an extract, so toxicity isn’t an issue.
- Screening Biofilm Disruptors: I can rapidly test dozens of enzyme variants that degrade biofilm matrices (like glucanases) without the long turnaround time of bacterial transformation and culture.
2. Describe the main components of a cell-free expression system and explain the role of each component.
- Cell Extract: The “engine” containing ribosomes, tRNAs, and initiation factors harvested from a host cell.
- Energy Buffer: A mix of NTPs (ATP, GTP, etc.) that power the translation process.
- Amino Acids: The building blocks used to assemble the protein.
- DNA/mRNA Template: The genetic instructions (blueprint) for the specific protein I want to make.
- Salts/Cofactors: Essential for stabilizing the ribosome and metabolic enzymes in the extract.
- Energy Regeneration System: Reagents like Phosphoenolpyruvate (PEP) that recycle spent ADP back into ATP to keep the reaction running for hours.
3. Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Energy is the literal fuel for the ribosome. Without a regeneration system, the “pool” of ATP would be depleted in minutes due to the high metabolic demand of protein synthesis and background phosphatase activity. To keep the reaction going, I’d use the PANOX system (which utilizes PEP and the enzyme Pyruvate Kinase). This allows the system to constantly “recharge” the ATP so the reaction can last for 10+ hours.
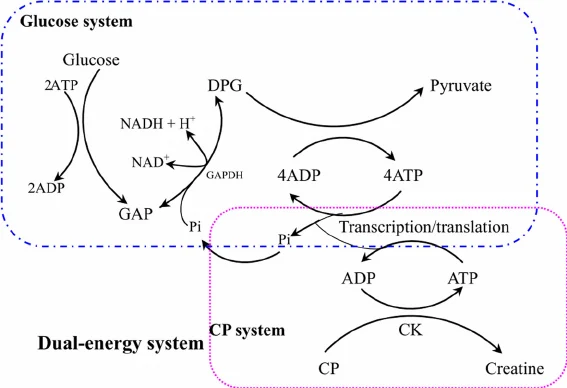

Figure: ATP regeneration pathways for cell-free protein synthesis showing the glucose system (top) and creatine phosphate (CP) system (bottom). In the dual-energy system, inorganic phosphate released by the CP system is recycled into glycolysis. Source: Kim et al., Biotechnology and Bioengineering, 2007.
4. Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
- Prokaryotic (E. coli): Best for high speed and high yield. I’d use this for Mutanase, an enzyme that breaks down the (1→3)-α-glucan in dental biofilms. It’s a bacterial enzyme, so it folds well in a bacterial extract.
- Eukaryotic (CHO or Wheat Germ): Better for complex human proteins. I’d use this for Histatin-5, a salivary protein with antifungal properties. Eukaryotic extracts are better at handling the specific folding and potential modifications this protein might need to be fully active.
5. How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
Membrane proteins are hydrophobic and tend to aggregate if they aren’t stuck in a lipid bilayer. To optimize this, I’d use Nanodiscs; tiny, discoidal pieces of lipid bilayer held together by a scaffold protein. I would add these nanodiscs directly to the cell-free reaction so the membrane protein can “sit” in the lipid environment as it’s being made, keeping it stable and functional.
6. Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
- RNA Degradation: RNases in the extract might be eating my mRNA. Fix: Add a potent RNase inhibitor like RNasin.
- Magnesium Titration: The Mg²⁺ concentration might be off. Ribosomes are very picky. Fix: Run a “magnesium sweep” (testing a range of 5–15 mM) to find the peak yield.
- Energy Exhaustion: The reaction might be running out of steam. Fix: Switch to a “fed batch” approach where I add fresh energy mix every few hours.
Homework Question from Kate Adamala: Synthetic Minimal Cell (SMC)
Pick a function and describe it. What would your synthetic cell do? What is the input and what is the output?
I want to design a “Biofilm Sentinel.” Its job is to sense when the dental biofilm is becoming acidogenic and release a neutralizing enzyme.
- Input: High acidity (low pH) or Quorum Sensing signals (like CSP from S. mutans).
- Output: Urease (which produces ammonia to raise the pH) or a bacteriocin to kill the pathogens.
Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
No. If it’s not encapsulated, the sensors and enzymes would just wash away with saliva. Encapsulation allows the SMC to stay localized within the “niche” of the biofilm and concentrate the response where it’s needed.
Could this function be realized by genetically modified natural cell?
Possibly, but natural cells are hard to control and might be outcompeted by the existing oral microbiome. An SMC is more predictable and won’t “evolve” into something else.
Describe the desired outcome of your synthetic cell operation.
The outcome is a more balanced oral microbiome where the “bad” bacteria are kept in check, preventing the pH from dropping low enough to cause demineralization.
Design all components that would need to be part of your synthetic cell.
- Membrane: DOPC/Cholesterol for a robust vesicle.
- Internal Machinery: E. coli S30 extract.
- Communication: I need a pH sensitive promoter or a receptor for CSP (Competence Stimulating Peptide). I’ll use the α-hemolysin (aHL) pore to allow the output (Urease) to exit the cell.
Experimental details (Lipids and Genes):
- Lipids: POPC, Cholesterol.
- Genes:
- comX/comE: The sensing system for S. mutans quorum signals.
- ureA/ureB: Genes for Urease to neutralize acid.
- hlyA: To create the α-hemolysin pores.
- Measurement: I’d measure the pH of the surrounding medium over time using a pH sensitive dye like Bromocresol Purple.
Homework Question from Peter Nguyen: Cell-Free Materials
Choose one application field—Architecture, Textiles/Fashion, or Robotics—and propose an application using cell-free systems that are functionally integrated into the material.
Field: Architecture (Healthcare Surface Design)
Summary Pitch: I propose “Active Antimicrobial Coatings” for dental clinic surfaces that only activate when they detect dental pathogens.
Detailed Mechanism: I envision a transparent polymer coating embedded with freeze-dried, cell-free reaction pellets. These pellets contain a genetic circuit that is triggered by AI-2 (Autoinducer-2), a common signaling molecule in oral biofilms. When a pathogen like Porphyromonas gingivalis settles on the surface, its signals rehydrate and activate the CFPS, which then expresses and secretes a localized dose of Lysostaphin to kill the bacteria and prevent biofilm formation.
Societal Challenge: This reduces the reliance on harsh chemical disinfectants and helps prevent the spread of cross contaminants in clinical settings.
Addressing Limitations: To handle “one time use,” the coating would be applied as a “smart film” that can be peeled and replaced once the color indicator (built into the circuit) shows the “bioink” has been spent.
Homework Question from Ally Huang: Genes in Space Proposal
1. Provide background information that describes the space biology question or challenge you propose to address.
Astronauts often suffer from “Space Gingivitis.” Microgravity and radiation seem to shift the oral microbiome toward a more pathogenic state, and the lack of traditional hygiene tools makes it worse. This isn’t just about gum health; oral pathogens are linked to systemic issues like heart disease, which is a major risk on a 3 year mission to Mars.
2. Name the molecular or genetic target that you propose to study.
The target is the gtfB gene (Glucosyltransferase B) of Streptococcus mutans, which is the primary enzyme responsible for the “glue” that allows biofilms to stick to teeth.
3. Describe how your molecular or genetic target relates to the space biology question or challenge your proposal addresses.
Biofilms behave differently in microgravity. They often grow thicker and more resistant to antibiotics. By studying how gtfB is expressed and how its resulting protein (GTF-B) folds and functions in space using BioBits, we can understand if the “stickiness” of dental plaque increases in microgravity.
4. Clearly state your hypothesis or research goal and explain the reasoning behind it.
Hypothesis: I hypothesize that the BioBits® system can be used to produce a functional GTF-B inhibitor (a specific nanobody or peptide) in microgravity, and that its binding affinity to the GTF-B protein will be altered by the lack of convection in space.
The reasoning is that if we can’t brush effectively in space, we need “on demand” biological therapeutics. Using BioBits to produce biofilm disrupting proteins allows astronauts to create fresh, personalized oral health treatments without needing a cold chain supply from Earth.
5. Outline your experimental plan.
I will use BioBits® to express the gtfB gene and a fluorescently labeled version of a known inhibitor.
- Test Sample: BioBits + gtfB DNA + Inhibitor DNA.
- Control: BioBits + gtfB DNA (no inhibitor).
- Measurements: I will use the P51 viewer to check for fluorescence. If the inhibitor is being produced and binding correctly, we should see a specific “quench” or shift in the signal. I’ll also use the miniPCR to check if the S. mutans DNA found in astronaut saliva samples shows an “upregulation” of the gtfB gene during flight compared to ground controls.
Homework Part B: Individual Final Project
I’ve put my slide to the slide deck and submit my final project form.