Week 2: DNA Read, Write, and Edit
Homework #2 🧬
Part 1: Benchling & In-silico Gel Art
Overview:
Import the Lambda DNA.
Simulate Restriction Enzyme Digestion with the following Enzymes:
- EcoRI
- HindIII
- BamHI
- KpnI
- EcoRV
- SacI
- SalI
Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks.
**Attempt **
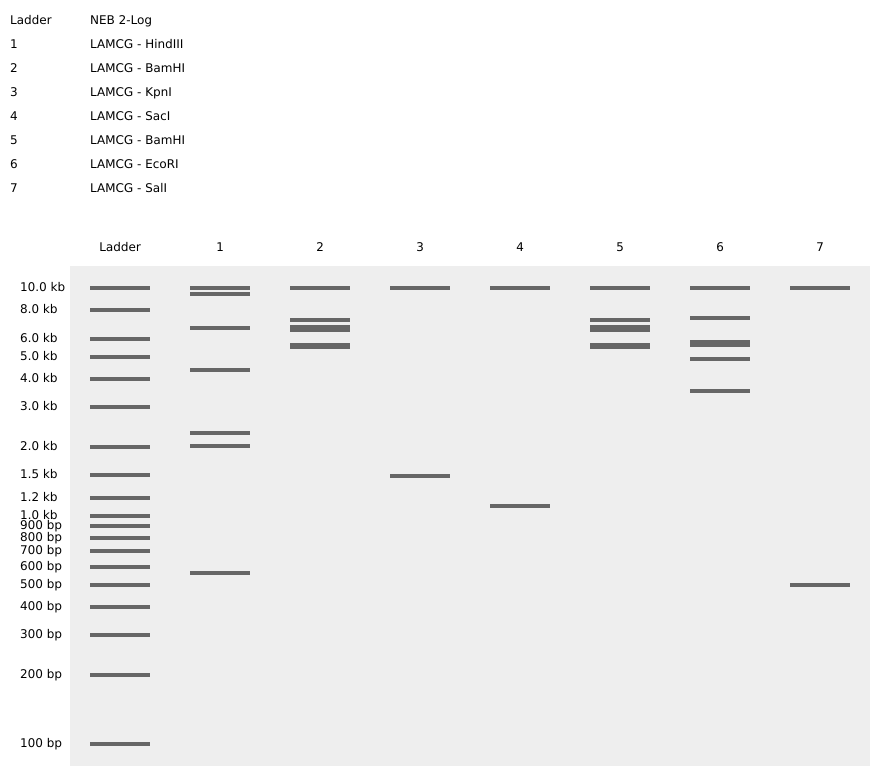

Part 3: DNA Design Challenge
3.1. Choose your protein.
In recitation, we discussed that you will pick a protein for your homework that you find interesting. Which protein have you chosen and why? Using one of the tools described in recitation (NCBI, UniProt, google), obtain the protein sequence for the protein you chose
Protein: Escherichia coli strain 29 beta-lactamase (TEM-1)
The protein I have chosen for the homework is PETase (poly(ethylene terephthalate) hydrolase) from the bacterium Piscinibacter sakaiensis (previously known as Ideonella sakaiensis). I find this protein particularly interesting because it represents a breakthrough in addressing one of the world’s major environmental challenges: plastic pollution. PETase is an enzyme that can break down polyethylene terephthalate (PET), a common plastic used in bottles, packaging, and textiles. Discovered in a bacterium isolated from plastic waste, PETase enables the microbe to use PET as a carbon and energy source by hydrolyzing its ester bonds. This natural biological degradation process offers hope for sustainable recycling and bioremediation of plastics, unlike traditional mechanical or chemical methods that are energy-intensive or produce pollutants. The enzyme’s specificity for PET and its activity at relatively mild temperatures also make it exciting for potential biotechnological applications, such as engineered variants for industrial plastic breakdown.
Using UniProt (one of the tools mentioned in recitation for protein information), I retrieved the protein sequence for PETase from Piscinibacter sakaiensis. The UniProt accession is A0A0K8P6T7, and here is the full amino acid sequence (290 residues):
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
The Central Dogma discussed in class and recitation describes the process in which DNA sequence becomes transcribed and translated into protein. The Central Dogma gives us the framework to work backward from a given protein sequence and infer the DNA sequence that the protein is derived from. Using one of the tools discussed in class, NCBI or online tools (google “reverse translation tools”), determine the nucleotide sequence that corresponds to the protein sequence you chose above.
Tool: Reverse translate Gene Corner https://www.genecorner.ugent.be/rev_trans.html
Piscinibacter sakaiensis PETase protein gene
3.3. Codon optimization.
Once the nucleotide sequence of your protein is determined, you need to codon optimize your sequence. You may, once again, utilize Google for a “codon optimization tool”.
Tool: Codon Optimization IDT https://www.idtdna.com/CodonOpt
In your own words, describe why do you need to optimize codon usage. Which organism have you chose to optimize the codon sequence for and why?
Optimization is vital to achieve improvements in protein synthesis efficiency, either in terms of stability, structure, and speed of the processes. This is achieved by employing specific codons that are preferred by the organism of interest. This translates into increased protein expression.
In this case, I selected Escherichia coli , one of the model organisms in protein production in biotechnology. The preference is associated with the ease of manipulation of its genes and rapid proliferation/growth as it is an organism that is not very demanding in terms of conditions. This makes it an ideal organism for this type of experiments.
3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
In this case, it is possible to use both methods:
Cell-free methods: based on the use of cell extracts or synthetic compounds with the ability to perform translation and transcription by having the respective machinery (ribosomes, RNA polymerase, etc.), without the need for living cells. These are usually encapsulated in cell-free protein synthesis systems (CFPs), capable of producing proteins that are collected directly. An example of this is through the use of a system that incorporates the preparation of a bacterial lysate and encapsulation in vesicles. There are also commercial CFPs kits that could be used to produce a protein of interest.
Cell-dependent methods: based on the use of live cells, in this case it is possible to work with plasmids for the production of recombinant proteins in E. coli . One of the most widely used series in recent years is the pET line, allowing efficient protein translation. In these systems, the incorporated machinery of the cells is what allows these processes to be executed, and it is also necessary to have: a DNA sequence, a terminator, a regulatory sequence, ARN polymerase, enhancers, and start and termination codons, among others. In addition to the insertion of the gene or genes, it is also necessary to carry out bacterial transformation processes, induce expression, and finally extract the purified protein.
Part 4: My first Benchling plasmid 🧬
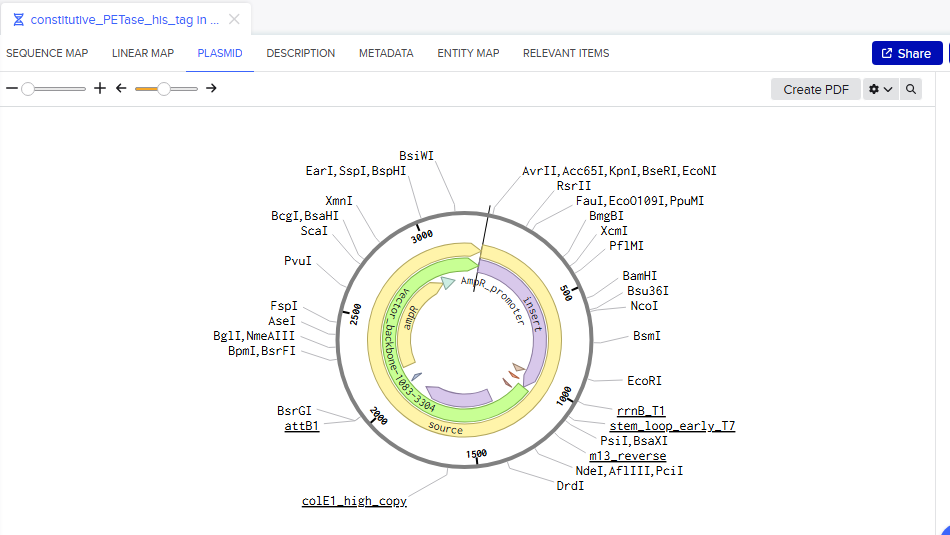

Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I consider that it could be of interest to work with the eae gene of the enteropathogenic pathotype of E. coli (EPEC), responsible for encoding the intimin protein, necessary for adherence to the intestinal epithelium and which causes diarrheal affections as a consequence worldwide. This could be very useful for environmental monitoring and the study of epidemiological patterns in developing countries such as Ecuador. Since it is one of the main pathogens of public health risk, sequencing is proposed as an alternative for the study in complex environments such as river waters or important sources of high contamination.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why? Also answer the following questions:
a. Is your method first-, second- or third-generation or other? How so?
The first-generation Sanger method is proposed for this case. It is positioned in this category as one of the first methods used in DNA sequencing in 1977. It is based on the addition of deoxynucleotides that facilitate DNA chain elongation. It is also useful in this case because of its accuracy, ease, cost, and, above all, because the size of the strand of interest is manageable for the technology (881bp).
b. What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
- Extraction of DNA from study samples (e.g. contaminated water). The use of an extraction kit is suggested to ensure higher purity of the sample and avoid other contaminants.
- Performing a conventional PCR to obtain an adequate amount of the fragment, ensuring that it is in a pure form. Only PCR conventional components are required as normal nucleotides (dNTPs) and a thermostable DNA polymerase (Taq polymerase).
c. What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
For Sanger sequencing the DNA obtained from PCR is mixed with other reagents: nucleotides (dNTPs) and other special nucleotides that are fluorescently labeled (ddNTPs).
The polymerase then synthesizes a new strand and when a ddNTP is added, the process is stopped, resulting in fragments of different lengths.
These fragments are separated in a capillary electrophoresis process where the shorter fragments migrate faster and in turn, the fragments are excited by a laser which emits a specific signal for each fragment.
These signals can then be recorded by a detector and translated into a nucleotide sequence.
d. What is the output of your chosen sequencing technology?
The method generates an electropherogram, which is a graph showing the fluorescence peaks corresponding to each nucleotide in the DNA sequence. Where each color represents a specific base (A, T, C, G).
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
For this section, I would be interested in synthesizing DNA associated with Shiga toxin as the Stx2 responsible for multiple outbreaks at the global level and the cause of hemolytic uremic syndrome. This toxin is usually produced by serotypes of pathogenic E. coli ( STEC), so its synthesis could be of interest in the development of recombinant vaccines, by obtaining attenuated antigens.
(ii) What technology or technologies would you use to perform this DNA synthesis and why? Also, answer the following questions:
I would make use of the Gibson Assembly technology because it is highly accurate and efficient compared to others such as Golden Gate, and I consider this to be essential in vaccine development. In addition, it is sufficiently suitable for the assembly of a plasmid with an attenuated version of the toxin and is flexible in case modifications are necessary to improve the immune response.
What are the essential steps of your chosen sequencing methods?
In the first instance, it is necessary to synthesize or amplify an attenuated version of the protein (toxin) of interest. This means removing the domains or parts associated with toxicity but retaining the elements that activate the immune response in patient’s body. This gene can be obtained by PCR and must have overlapping ends that match the plasmid where the insertion will be made. The plasmid to be used is also pre-designed and linearized to facilitate insertion.
The next step is the assembly, which consists of mixing these components in a tube with Gibson’s mix containing: exonuclease responsible for generating the overlapping ends, polymerase that fills these spaces, and ligase that joins these fragments.
Finally, the next step is the transformation of the organism chosen, in this case, E. coli, by the addition of this recombinant plasmid.
b. What are the limitations of your sequencing method (if any) in terms of speed, accuracy, and scalability?
Among the limitations of this method are the possible formation of secondary structures and the need for long overlapping sequences which could lead to complications in the design and synthesis. The cost could also be relatively high compared to the other alternatives.
5.3 DNA Edit.
(i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
For this part of the paper, I would again bring up the idea of modifying the genes of plants that are subject to desiccation problems such as bananas. I believe that the agricultural sector in countries like Ecuador has great potential to test these technologies and improve yield and productivity levels.
(ii) What technology or technologies would you use to perform these DNA edits and why? Also answer the following questions:
How does your technology of choice edit DNA? What are the essential steps?
It starts with the design of the construct of interest, in this case consisting of the DREB1A gene, which is inserted into an expression vector together with its promoter.
This vector is then introduced into A. tumefaciens and the plants of interest are infected in an in vitro culture, which will allow the integration of the gene of interest. The principle of this technology is based on the ability of this bacterium to transfer DNA to other cells, using its Ti plasmid in which the region associated with the tumors is replaced by the region of interest. Thus, when this bacterium infects plant tissue, this genetic alteration is also transferred.
Subsequently, the plants that have been transformed correctly are selected, this can be through a fluorescent marker such as GFP.
Additionally, expression tests can be performed by RT-qPCR, and lastly, the regeneration and re-planting of the culture of interest is performed.
b. What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
This process requires the selected gene of interest, a suitable vector compatible with A. tumefaciens including a promoter, terminator, and selection marker. Also, designed primers, restriction enzymes, ligases, culture media, and growth hormones.
c. What are the limitations of your editing methods (if any) in terms of efficiency or precision?
The main limitations revolve around the efficacy of the transformation because it is subject to a process of transgenesis, which could compromise the specificity and accuracy of the editing. In addition to possible unwanted adverse effects due to random insertions.