Week 3: Lab Automation
Homework #3 🤖
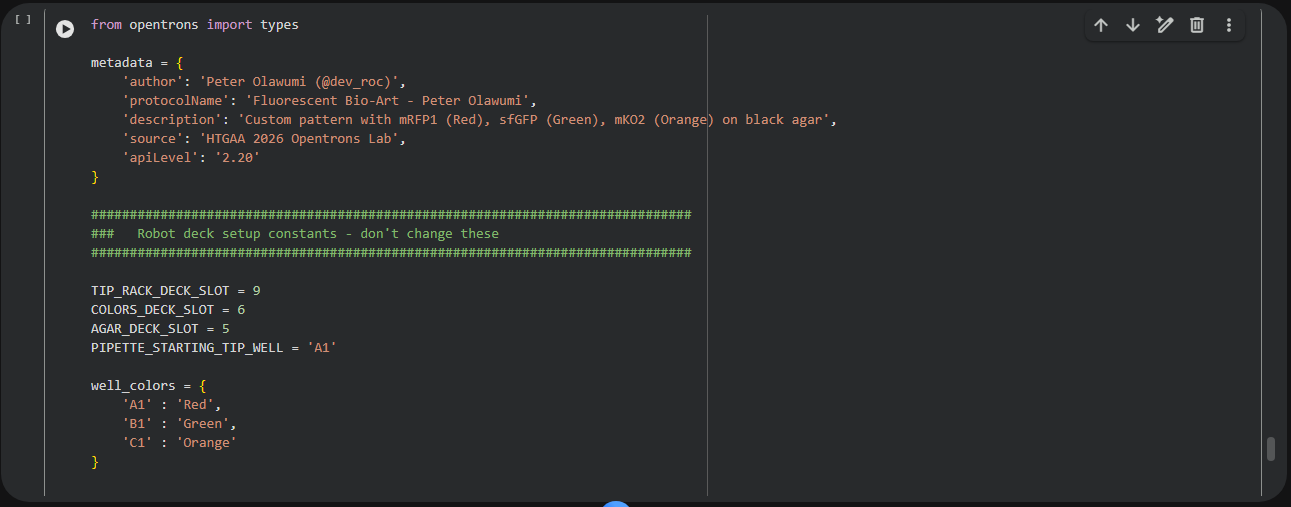
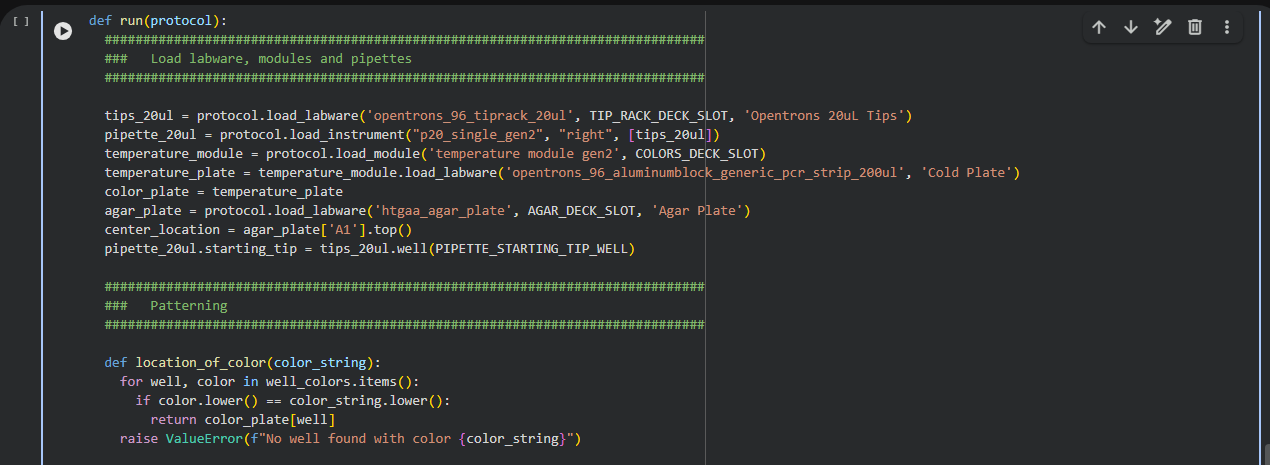
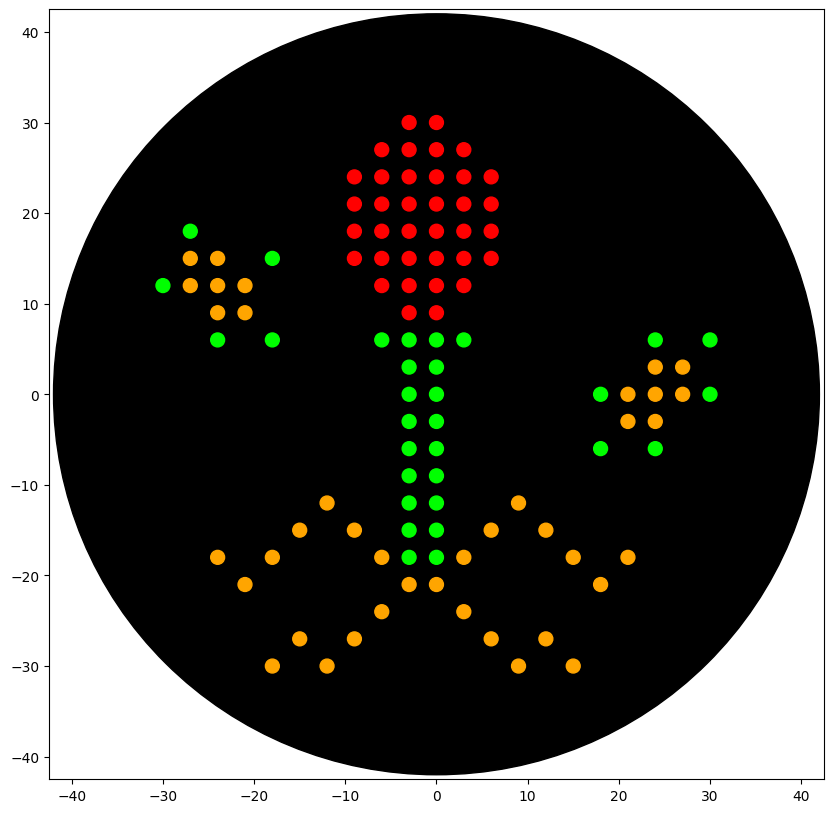
Part 2: Python code for the petri dish design
Link of colab: (https://colab.research.google.com/drive/1mW0Mx0VoMFat2BOsuizyHktBI3Vd4LSO?usp=sharing)



Part 3: Post Lab Questions
1. Published Paper Using Opentrons for Novel Biological Applications
One compelling example is the paper “Semi-automated Production of Cell-Free Biosensors” by Dylan M. Brown, Daniel A. Phillips, and colleagues (bioRxiv preprint October 13, 2024; formally published in ACS Synthetic Biology, 2025).
The team used the affordable Opentrons OT-2 liquid-handling robot to scale up manufacturing of cell-free synthetic biology biosensors for point-of-need diagnostics (e.g., detecting fluoride in drinking water). They developed a semi-automated protocol that precisely assembles viscous cell-free reaction mixes (DNA template + PANOx extract + buffers) into full 384-well plates in ~30 minutes—something that was previously done manually with high operator-to-operator variability.
Key novel application: They created and lyophilized hundreds of identical fluoride-riboswitch biosensors that can be rehydrated in the field and give a clear colorimetric or fluorescent readout. By optimizing robot parameters (dispense height, mix volume, aspiration rate), they achieved reproducibility that matched or exceeded manual assembly while drastically reducing hands-on time and batch-to-batch variation. This opens the door to cheap, deployable diagnostics in low-resource settings (they reference prior field tests in Kenya and Costa Rica). The work is especially elegant because it shows how open-source automation turns cell-free systems from lab curiosities into manufacturable products—exactly the kind of scalability we need in synthetic biology.
2. What I Intend to Do with Automation Tools for My Final Project
Project Title: Microbial “Plastic Eaters” – Engineering On-Site Industrial Recycling Pods with Recombinant PETase/MHETase in a Cell-Free + Bacterial Pipeline
My final project builds a portable “recycling pod” that uses engineered bacteria (or their secreted enzymes) to break down PET plastic waste directly on factory floors. The bottleneck is rapid optimization of PETase and MHETase variants for faster degradation, higher temperature tolerance, and better secretion. Automation will let me screen dozens-to-hundreds of variants in parallel, run degradation assays remotely, and iterate in days instead of weeks.
Here is exactly what I plan to automate:
A. High-Throughput Variant Library Assembly & Cell-Free Expression Screening (Primary automation goal – inspired by the cell-free biosensor paper above)
- Opentrons OT-2 (or cloud lab equivalent) will perform Golden Gate assembly of PETase mutant libraries (active-site saturation + secretion-signal variants).
- Echo transfer or Opentrons p20 multi-channel will dispense 50–100 ng of each linearized plasmid + cofactors into 96-well or 384-well plates.
- Bravo / Opentrons stamps in the cell-free protein synthesis (CFPS) master mix (E. coli lysate + energy components).
- Multiflo dispenses the full reaction volume to start expression.
- PlateLoc seals the plate.
- Inheco or Opentrons temperature module incubates at 30 °C / 37 °C for 4–16 h.
- XPeel removes seal.
- PHERAstar or plate reader measures either (a) fluorescence (GFP-fused PETase) or (b) enzymatic activity via p-nitrophenyl ester surrogate substrate at 405 nm.
Pseudocode / Opentrons Python sketch:
B. 3D-Printed Custom Holders (from Opentrons 3D Printing Directory style)
I will design and print (using the class Prusa or lab printer) a PET-flake assay tray: a 96-well-compatible holder that securely positions 5 mm × 5 mm shredded PET flakes or thin PET film strips at the bottom of each well. The holder has sloped walls and a mesh bottom so supernatant can be easily aspirated for downstream HPLC or weight-loss measurements without losing plastic particles. This turns a messy manual assay into a clean, robot-friendly 96-well format.
C. Cloud-Lab Integration (Ginkgo Nebula / similar remote biofoundry)
Once top variants are identified on the Opentrons, I will upload the best 10–20 constructs to Ginkgo Nebula (or equivalent cloud laboratory) for larger-scale bacterial expression and real PET degradation in 1 L bioreactors. The cloud lab will:
- Run parallel fermentations with automated sampling.
- Perform continuous OD600, pH, and TPA/EG monomer quantification via inline HPLC.
- Return lyophilized enzyme powders ready for pod prototyping.
D. Full Degradation Validation Loop
After cell-free hits, Opentrons will set up 24–48 replicate mini-reactions with purified enzyme + real factory PET scraps, incubate with shaking, and automatically sample at 0/24/48/72 h for mass-loss and LC-MS readout. This closed loop (design → assemble → express → assay → analyze) will run with minimal intervention, letting me test 50+ variants per week.
By combining the Opentrons for precision liquid handling, 3D-printed custom labware for PET-specific assays, and cloud-lab scale-up, I will move from gene sequence to validated high-performance enzyme cocktail in a matter of weeks—exactly what an industrial recycling pod needs. This automation plan directly mirrors the cell-free biosensor paper’s success in scaling reproducible reactions and will make my project robust, repeatable, and genuinely ready for Lagos factory floors.