Subsections of Homeworks
Week 1 HW: Principles and Practices
Process of Researching the project:
I looked at some iGem projects to gain inspiration and a sense of the types of topics that people have addressed. https://video.igem.org/c/2025_presentations/videos
First, describe a biological engineering application or tool you want to develop and why.
Vitiligo is an autoimmune condition characterized by the selective loss of melanocytes, leading to depigmented skin lesions. A growing body of evidence suggests that elevated oxidative stress in melanocytes precedes and exacerbates immune-mediated destruction by increasing cellular damage and inflammatory signaling.
My project proposes an oxidative-stress–responsive genetic circuit in melanocytes that activates cytoprotective pathways only under pathological redox conditions and automatically deactivates once oxidative stress resolves. By enhancing stress resilience rather than inducing pigmentation, this approach aims to reduce melanocyte vulnerability while minimizing unintended cosmetic or immune effects.
Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals
Goals:
Prevent Harm: Prevent the technology from causing biological, social, or psychological harm. • A1. Prevent biological harm from uncontrolled gene expression, persistence, or immune disruption. • A2 Prevent social harm, including stigmatization, cosmetic misuse, or reinforcement of colorism. • A3 Prevent misuse or repurposing beyond therapeutic, supportive contexts.
Promote Safe and Constructive Innovation: Enable beneficial research and translation without unnecessary restriction.
• B1. Encourage disease-aligned, upstream interventions (stress resilience vs cosmetic alteration). • B2. Avoid chilling effects on legitimate academic research. • B3. Support iterative learning and transparency around failures.
Respect Autonomy, Equity, and Trust • C1. Incorporate patient perspectives into design and deployment decisions. • C2. Avoid one-size-fits-all assumptions about desirability of treatment. • C3. Ensure accessibility and avoid disproportionate burdens on marginalized groups.
Next, describe at least three different potential governance “actions” by considering the four aspects below (Purpose, Design, Assumptions, Risks of Failure & “Success”).
Governance Action 1: Mandatory Design-Layer Safeguards for Stress-Responsive Therapeutics
Governance Action 2: Patient-Guided Scope Limitation via Advisory Committees
Governance Action 3: Transparency & Post-Research Monitoring Norms
3. Scoring Governance Actions Against Policy Goals
(1 = best, 3 = weakest, n/a = not applicable)
| Governance Action / Policy Goal | Option 1 Design Safeguards | Option 2 Patient Governance | Option 3 Transparency |
|---|---|---|---|
| A. Prevent Biological Harm | 1 | 2 | 2 |
| • A1 Biological safety | 1 | 3 | 2 |
| • A2 Social harm | 2 | 1 | 2 |
| • A3 Misuse prevention | 2 | 2 | 1 |
| B. Promote Constructive Use | 2 | 1 | 1 |
| • B1 Disease alignment | 1 | 1 | 2 |
| • B2 Not impede research | 2 | 1 | 1 |
| • B3 Learning from failure | 2 | 2 | 1 |
| C. Autonomy & Trust | 3 | 1 | 2 |
| • C1 Patient voice | n/a | 1 | 2 |
| • C2 Respect diversity | 2 | 1 | 2 |
| • C3 Equity | 2 | 2 | 1–2 |
Recommended approach
Prioritize a combined strategy centered on Option 1 (Design Safeguards) and Option 2 (Patient Governance), with Option 3 (Transparency) as a supporting layer. Option 1 scored strongest on preventing biological harm (A1) and performed well on misuse prevention. Because the MelanoGuard system directly manipulates cellular stress-response pathways, technical harm prevention must occur at the level of the biological system itself, not solely through external oversight. Option 2 scored best on social harm prevention (A2), autonomy and trust (C), and constructive use (B). Vitiligo is a visible, identity-linked condition, so ethical failure is more likely to occur at the social level than the molecular level. Option 3 scored highest for misuse detection (A3) and learning from failure (B3), but weaker on direct harm prevention and trust building.
Lecture 2 Slides 1 Q/A
Nature’s machinery for copying DNA is called polymerase.
1:10⁶ (error:base-pairs)
Human genomee is ~3.2 X 10⁹ base pairs. That would be 3200 errors. Biology has error correction and has a proof reading step and a post-replication mismatch repair.
Lecture 2 Slides 2 Q/A
- What is the most commonly used method for oligo synthesis today?
all commercial oligos today are made using phosphoramidite chemistry. - Why is it difficult to make oligos longer than ~200 nt by direct synthesis?
Error accumulation per synthesis cycle and errors compound exponentially. - Why can’t you make a 2000 bp gene via direct oligo synthesis?
Direct synthesis scales exponentially poorly, not linearly and errors would be everywhere
Lecture 2 Slides 3 Q/A
What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”? 1. Histidine 2. Isoleucine 3. Leucine 4. Lysine 5. Methionine 6. Phenylalanine 7. Threonine 8. Tryptophan 9. Valine 10. Arginine
So, #4 shows that all animals have in Lysine. Microbes and plants can synthesize all 20 canonical amino acids. This is why lysine biosynthesis pathways exist in bacteria but not animals. In animals (including humans) Lysine is always externally supplied, Cells already operate under lysine dependency. Therefore, Lysine contingency does NOT inherently restrict survival and is not a meaningful containment mechanism in animal hosts. Lysine contingency can function as containment in microbes, but only if: The environment lacks lysine, No cross-feeding occurs, No bypass pathways evolve. Lysine contingency should never be used alone and should be combined with the following so that animals can’t supply it, Microbes can’t evolve it easily and Environment doesn’t contain it:
• Genetic code recoding
• Non-standard amino acid dependence
• Codon reassignment
• Virus resistance
• Metabolic isolation
Knowing that lysine is one of the 10 essential amino acids in all animals makes it clear that lysine contingency alone is not a robust biocontainment strategy; it only becomes meaningful when combined with genetic code engineering or dependence on non-standard amino acids. Because lysine is essential to all animals and ubiquitously available through diet and environment, “lysine contingency” by itself is biologically weak and only functions as a safety mechanism when embedded in a larger framework of genetic code recoding and metabolic isolation.
[Given slides #2 & 4 (AA:NA and NA:NA codes)] What code would you suggest for AA:AA interactions?
Collapse AA pairs into a small set of symbols:
• HΦ: hydrophobic packing (Leu/Ile/Val/Phe…)
• HB: hydrogen-bond pairing (polar donors/acceptors)
• SB: salt bridge (Asp/Glu ↔ Lys/Arg/His; pH-dependent)
• π: aromatic stacking / cation–π (Phe/Tyr/Trp with Lys/Arg)
• SS: disulfide bond (Cys–Cys)
• M: metal coordination (His/Cys/Asp/Glu and specific metals)
• G/P: geometry breakers (Gly flexibility, Pro rigidity)
• X: steric/charge clash (strongly unfavorable)
Now an AA:AA “codeword” is something like:
• (Asp, Lys) → SB
• (Cys, Cys) → SS
• (Phe, Tyr) → π
• (Leu, Ile) → HΦ
ChatGPT prompts:
- Tell me about NRF2–KEAP1 pathway and the role it plays in Vitiligo
- Are there any scientific issues with my explanation: “Vitiligo is is an autoimmune disease where oxidative stress in Melanocytes triggers immune system to attack them causing pigmentation. The idea I’m considering is developing a oxidative stress sensing mechanism that regulates a genetic circuit which turns on the protective mechanism in melanocytes until oxidative stress is active, and turns off the the protection when no oxidative stress is detected.”
- Create a narrative based on the slides (for all the slides from lecture 2)
- What’s Lysine Contingency
- If Lysine is already produced endogenously, why is used as a biocontainment strategy?
Week 2 HW: DNA Read, Write, and Edit
Part 3: DNA Design Challenge

3.1 Protein of interest:
I’m interested in exploring JAK1 protein that is implicated in autoimmune diseases. I chose this by finding which are the priority protein targets for treatment of conditions like Vitiligo. Link to the protein from Uniprot: https://rest.uniprot.org/uniprotkb/P23458.fasta
Protein Sequence:
>sp|P23458|JAK1_HUMAN Tyrosine-protein kinase JAK1 OS=Homo sapiens OX=9606 GN=JAK1 PE=1 SV=2
MQYLNIKEDCNAMAFCAKMRSSKKTEVNLEAPEPGVEVIFYLSDREPLRLGSGEYTAEEL
CIRAAQACRISPLCHNLFALYDENTKLWYAPNRTITVDDKMSLRLHYRMRFYFTNWHGTN
DNEQSVWRHSPKKQKNGYEKKKIPDATPLLDASSLEYLFAQGQYDLVKCLAPIRDPKTEQ
DGHDIENECLGMAVLAISHYAMMKKMQLPELPKDISYKRYIPETLNKSIRQRNLLTRMRI
NNVFKDFLKEFNNKTICDSSVSTHDLKVKYLATLETLTKHYGAEIFETSMLLISSENEMN
WFHSNDGGNVLYYEVMVTGNLGIQWRHKPNVVSVEKEKNKLKRKKLENKHKKDEEKNKIR
EEWNNFSYFPEITHIVIKESVVSINKQDNKKMELKLSSHEEALSFVSLVDGYFRLTADAH
HYLCTDVAPPLIVHNIQNGCHGPICTEYAINKLRQEGSEEGMYVLRWSCTDFDNILMTVT
CFEKSEQVQGAQKQFKNFQIEVQKGRYSLHGSDRSFPSLGDLMSHLKKQILRTDNISFML
KRCCQPKPREISNLLVATKKAQEWQPVYPMSQLSFDRILKKDLVQGEHLGRGTRTHIYSG
TLMDYKDDEGTSEEKKIKVILKVLDPSHRDISLAFFEAASMMRQVSHKHIVYLYGVCVRD
VENIMVEEFVEGGPLDLFMHRKSDVLTTPWKFKVAKQLASALSYLEDKDLVHGNVCTKNL
LLAREGIDSECGPFIKLSDPGIPITVLSRQECIERIPWIAPECVEDSKNLSVAADKWSFG
TTLWEICYNGEIPLKDKTLIEKERFYESRCRPVTPSCKELADLMTRCMNYDPNQRPFFRA
IMRDINKLEEQNPDIVSEKKPATEVDPTHFEKRFLKRIRDLGEGHFGKVELCRYDPEGDN
TGEQVAVKSLKPESGGNHIADLKKEIEILRNLYHENIVKYKGICTEDGGNGIKLIMEFLP
SGSLKEYLPKNKNKINLKQQLKYAVQICKGMDYLGSRQYVHRDLAARNVLVESEHQVKIG
DFGLTKAIETDKEYYTVKDDRDSPVFWYAPECLMQSKFYIASDVWSFGVTLHELLTYCDS
DSSPMALFLKMIGPTHGQMTVTRLVNTLKEGKRLPCPPNCPDEVYQLMRKCWEFQPSNRT
SFQNLIEGFEALLK
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
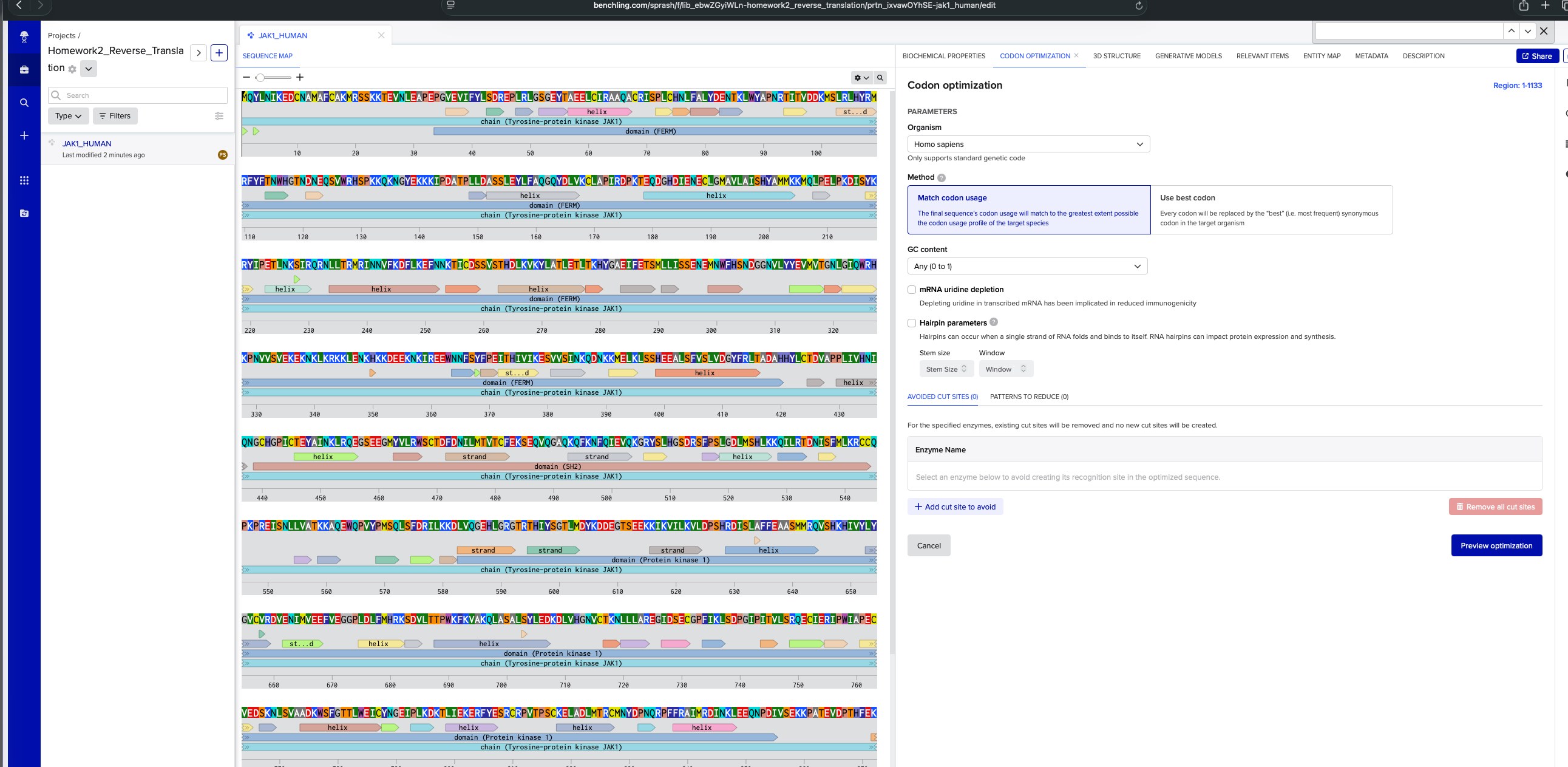

DNA Sequence for homosapien using codon optimization tool in benchling:
ATGCAGTACCTGAACATCAAGGAGGACTGCAACGCCATGGCCTTCTGCGCCAAGATGAGAAGCAGCAAGAAGACCGAGGTGAACCTGGAGGCCCCCGAGCCCGGCGTGGAGGTGATCTTCTACCTGAGCGACAGAGAGCCCCTGAGGCTGGGCAGCGGCGAGTACACCGCCGAGGAGCTGTGCATCAGAGCCGCCCAGGCCTGCAGGATCAGCCCCCTGTGCCACAACCTGTTCGCCCTGTACGACGAGAACACCAAGCTGTGGTACGCCCCCAACAGGACCATCACCGTGGACGACAAGATGAGCCTGAGGCTGCACTACAGGATGAGGTTCTACTTCACCAACTGGCACGGCACCAACGACAACGAGCAGAGCGTGTGGAGGCACAGCCCCAAGAAGCAGAAGAACGGCTACGAGAAGAAGAAGATCCCCGACGCCACCCCCCTGCTGGACGCCAGCAGCCTGGAGTACCTGTTCGCCCAGGGCCAGTACGACCTGGTGAAGTGCCTGGCCCCCATCAGGGACCCCAAGACCGAGCAGGACGGCCACGACATCGAGAACGAGTGCCTGGGCATGGCCGTGCTGGCCATCAGCCACTACGCCATGATGAAGAAGATGCAGCTGCCCGAGCTGCCCAAGGACATCAGCTACAAGAGGTACATCCCCGAGACCCTGAACAAGAGCATCAGGCAGAGGAACCTGCTGACCAGAATGAGGATCAACAACGTGTTCAAGGACTTCCTGAAGGAGTTCAACAACAAGACCATCTGCGACAGCAGCGTGAGCACCCACGACCTGAAGGTGAAGTACCTGGCCACCCTGGAGACCCTGACCAAGCACTACGGCGCCGAGATCTTCGAGACCAGCATGCTGCTGATCAGCAGCGAGAACGAGATGAACTGGTTCCACAGCAACGACGGCGGCAACGTGCTGTACTACGAGGTGATGGTGACCGGCAACCTGGGCATCCAGTGGAGGCACAAGCCCAACGTGGTGAGCGTGGAGAAGGAGAAGAACAAGCTGAAGAGGAAGAAGCTGGAGAACAAGCACAAGAAGGACGAGGAGAAGAACAAGATCAGGGAGGAGTGGAACAACTTCAGCTACTTCCCCGAGATCACCCACATCGTGATCAAGGAGAGCGTGGTGAGCATCAACAAGCAGGACAACAAGAAGATGGAGCTGAAGCTGAGCAGCCACGAGGAGGCCCTGAGCTTCGTGAGCCTGGTGGACGGCTACTTCAGGCTGACCGCCGACGCCCACCACTACCTGTGCACCGACGTGGCCCCCCCCCTGATCGTGCACAACATCCAGAACGGCTGCCACGGCCCCATCTGCACCGAGTACGCCATCAACAAGCTGAGACAGGAGGGCAGCGAGGAGGGCATGTACGTGCTGAGGTGGAGCTGCACCGACTTCGACAACATCCTGATGACCGTGACCTGCTTCGAGAAGAGCGAGCAGGTGCAGGGCGCCCAGAAGCAGTTCAAGAACTTCCAGATCGAGGTGCAGAAGGGCAGGTACAGCCTGCACGGCAGCGACAGGAGCTTCCCCAGCCTGGGCGACCTGATGAGCCACCTGAAGAAGCAGATCCTGAGGACCGACAACATCAGCTTCATGCTGAAGAGGTGCTGCCAGCCCAAGCCCAGAGAGATCAGCAACCTGCTGGTGGCCACCAAGAAGGCCCAGGAGTGGCAGCCCGTGTACCCCATGAGCCAGCTGAGCTTCGACAGGATCCTGAAGAAGGACCTGGTGCAGGGCGAGCACCTGGGCAGAGGCACCAGGACCCACATCTACAGCGGCACCCTGATGGACTACAAGGACGACGAGGGCACCAGCGAGGAGAAGAAGATCAAGGTGATCCTGAAGGTGCTGGACCCCAGCCACAGGGACATCAGCCTGGCCTTCTTCGAGGCCGCCAGCATGATGAGGCAGGTGAGCCACAAGCACATCGTGTACCTGTACGGCGTGTGCGTGAGGGACGTGGAGAACATCATGGTGGAGGAGTTCGTGGAGGGCGGCCCCCTGGACCTGTTCATGCACAGGAAGAGCGACGTGCTGACCACCCCCTGGAAGTTCAAGGTGGCCAAGCAGCTGGCCAGCGCCCTGAGCTACCTGGAGGACAAGGACCTGGTGCACGGCAACGTGTGCACCAAGAACCTGCTGCTGGCCAGGGAGGGCATCGACAGCGAGTGCGGCCCCTTCATCAAGCTGAGCGACCCCGGCATCCCCATCACCGTGCTGAGCAGGCAGGAGTGCATCGAGAGGATCCCCTGGATCGCCCCCGAGTGCGTGGAGGACAGCAAGAACCTGAGCGTGGCCGCCGACAAGTGGAGCTTCGGCACCACCCTGTGGGAGATCTGCTACAACGGCGAGATCCCCCTGAAGGACAAGACCCTGATCGAGAAGGAGAGGTTCTACGAGAGCAGATGCAGGCCCGTGACCCCCAGCTGCAAGGAGCTGGCCGACCTGATGACCAGGTGCATGAACTACGACCCCAACCAGAGGCCCTTCTTCAGAGCCATCATGAGGGACATCAACAAGCTGGAGGAGCAGAACCCCGACATCGTGAGCGAGAAGAAGCCCGCCACCGAGGTGGACCCCACCCACTTCGAGAAGAGGTTCCTGAAGAGGATCAGGGACCTGGGCGAGGGCCACTTCGGCAAGGTGGAGCTGTGCAGATACGACCCCGAGGGCGACAACACCGGCGAGCAGGTGGCCGTGAAGAGCCTGAAGCCCGAGAGCGGCGGCAACCACATCGCCGACCTGAAGAAGGAGATCGAGATCCTGAGGAACCTGTACCACGAGAACATCGTGAAGTACAAGGGCATCTGCACCGAGGACGGCGGCAACGGCATCAAGCTGATCATGGAGTTCCTGCCCAGCGGCAGCCTGAAGGAGTACCTGCCCAAGAACAAGAACAAGATCAACCTGAAGCAGCAGCTGAAGTACGCCGTGCAGATCTGCAAGGGCATGGACTACCTGGGCAGCAGGCAGTACGTGCACAGGGACCTGGCCGCCAGGAACGTGCTGGTGGAGAGCGAGCACCAGGTGAAGATCGGCGACTTCGGCCTGACCAAGGCCATCGAGACCGACAAGGAGTACTACACCGTGAAGGACGACAGGGACAGCCCCGTGTTCTGGTACGCCCCCGAGTGCCTGATGCAGAGCAAGTTCTACATCGCCAGCGACGTGTGGAGCTTCGGCGTGACCCTGCACGAGCTGCTGACCTACTGCGACAGCGACAGCAGCCCCATGGCCCTGTTCCTGAAGATGATCGGCCCCACCCACGGCCAGATGACCGTGACCAGACTGGTGAACACCCTGAAGGAGGGCAAGAGGCTGCCCTGCCCCCCCAACTGCCCCGACGAGGTGTACCAGCTGATGAGGAAGTGCTGGGAG
3.4. You have a sequence! Now what? What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
At first, Transcribe the DNA sequence to mRNA by inserting the DNA into living cells. The DNA template strand is read 3’ to 5’ and mRNA is synthesized 5’ to 3’. The resulting mRNA contains 5’ UTR, start codon (AUG), coding sequence, stop codon, and 3’ UTR.
Next is Translation. The ribosome binds the mRNA, it recognizes the start codon (AUG), reads the codons sequentially, tRNA delivers corresponding amino acids. Peptide bonds are formed and it stops at stop codon (UAA, UAG, UGA). The polypeptide then folds, may undergo post-translational modification and could get either secreted or remain intracellular.
3.5. [Optional] How does it work in nature/biological systems? Describe how a single gene codes for multiple proteins at the transcriptional level.
Single genes code for multiple proteins through the process of alternate splicing where different combination of exons are expressed.
Superimposed DNA -> AA
Part 4: Prepare a Twist DNA Synthesis Order
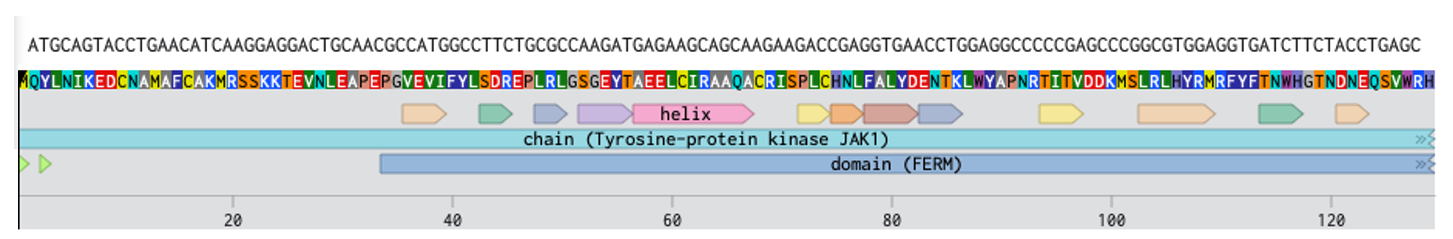

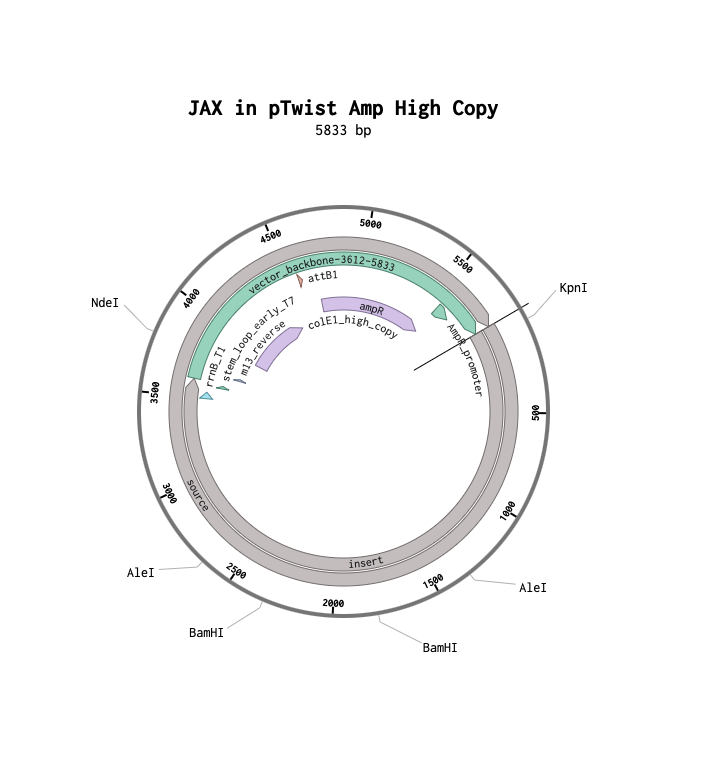
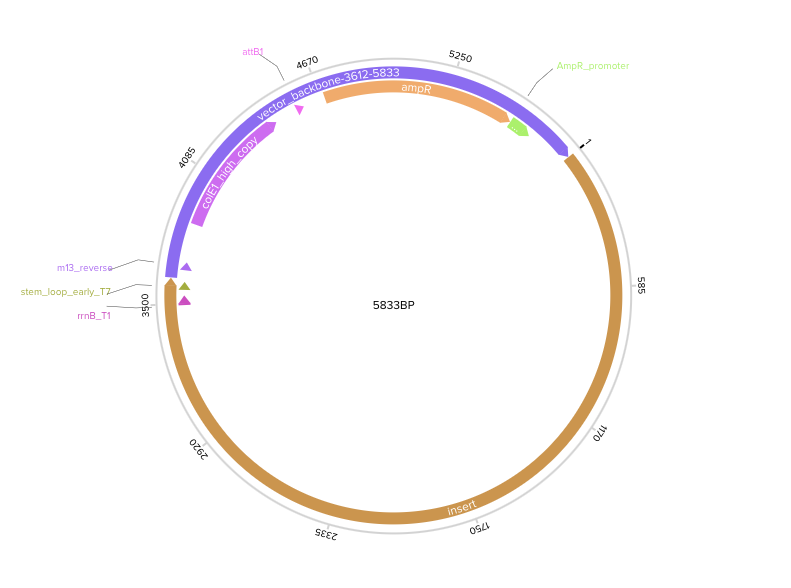
My annotated expression cassette
(JAX1 protein)[https://benchling.com/s/seq-kczgpR5k4eKn4BIEZYrC?m=slm-XqgAw2klZciZdJUaxluG]
Final Plasmid
Part 5: DNA Read/Write/Edit
Given my project for Vitiligo to develop a genetic circuit that detects reactive oxidative stress (ROS) and turns on genetic mechanisms to protect melanocytes, I would build the following bioengineering pipeline.
- DNA READ → Understand endogenous ROS regulatory DNA
- DNA WRITE → Build synthetic ROS-responsive circuit
- DNA EDIT → Insert circuit into melanocytes safely
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I would sequence the regulatory DNA that controls the oxidative stress response in melanocytes. My goal is to build a system that detects oxidative stress (ROS) in melanocytes and turn on protective genes. I would want to sequence:
The endogenous antioxidant response pathway genes in melanocytes
• Especially NRF2 (NFE2L2)
• KEAP1
• Antioxidant genes like HMOX1 (HO-1), NQO1, GCLC
Out of these, I would choose the NRF2, which is the master regulator of oxidative stress response. It activates antioxidant genes when ROS levels increase. If I understand the exact DNA sequence in melanocytes, I can:
• Identify natural promoter elements
• Find antioxidant response elements (AREs)
• Detect polymorphisms that affect stress response
• Design a synthetic circuit that mimics or enhances this pathway
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Also answer the following questions:
Is your method first-, second- or third-generation or other? How so?
What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
Since this is targeted sequencing of specific loci, I would use 2nd-gen, short-read sequencing (Illumina sequencing). Illumina sequencing has high accuracy, is suitable for variant detection. If I were studying structural variants or large rearrangements, I might use PacBio or Oxford Nanopore (third-gen). For my application, short-read high accuracy is ideal.
What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
Input: Genomic DNA extracted from melanocytes.
- DNA Extraction
- Lysis: Break open cells to release genomic DNA — ensures access to genetic material.
- Protein digestion: Remove histones and other proteins — prevents contamination and improves downstream efficiency.
- DNA purification: Isolate clean DNA — required for accurate fragmentation and library prep.
- Fragmentation: Mechanical (sonication) or enzymatic shearing: Break DNA into smaller pieces because sequencing platforms cannot read very long DNA directly. Target size ~300–500 bp: Optimal length for Illumina read chemistry and efficient cluster formation.
- End Repair: Generate blunt ends: Fix overhangs or damaged ends after fragmentation ensures uniform ends for adapter ligation.
- 4. A-tailing: Add single 3′ A overhang: Creates compatible ends for T-overhang adapters which increases ligation specificity and efficiency.
- Adapter Ligation
- Ligate Illumina adapters with index barcodes:
- Adapters allow DNA fragments to bind to the flow cell.
- Barcodes enable multiplexing of multiple samples in one sequencing run.
- PCR Amplification: Enrich adapter-ligated fragments: Selectively amplifies properly ligated DNA — increases library concentration and removes unligated fragments.
- Cluster Generation
- Bind to flow cell: Adapter sequences hybridize to complementary oligos on the flow cell surface which immobilizes fragments.
- Bridge amplification: Creates clonal clusters of identical DNA fragments — amplifies signal strength for accurate base detection during sequencing.
Illumina uses Sequencing by Synthesis (SBS):
1. Add fluorescently labeled reversible terminator nucleotides.
2. DNA polymerase incorporates ONE base.
3. Lasers excite fluorophores.
4. Camera records color → identifies base.
5. Terminator removed.
6. Cycle repeat, where each cycle = one base.
What is the output of your chosen sequencing technology?
The output is FASTQ files which contain Sequence reads, Quality scores (Phred scores). This can be used to align to reference genome and perform variant calling
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
I would synthesize a ROS-sensing genetic circuit that:
- Detects oxidative stress
- Activates antioxidant genes
- Protects melanocytes from damage
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
Also answer the following questions:
I would use Phosphoramidite DNA synthesis + Gene Assembly (Commercial gene synthesis, e.g., Twist Bioscience)
What are the essential steps of your chosen sequencing methods?
What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
I would use chemical DNA synthesis.
DNA Synthesis Workflow
- Solid-phase phosphoramidite synthesis: Chemically builds short DNA oligos base by base on a solid support to generate designed sequences.
- Oligo cleavage: Releases synthesized oligos from the solid support and removes protecting groups to obtain usable DNA.
- Oligo assembly (Gibson or enzymatic assembly): Joins overlapping oligos into a full-length gene construct.
- Cloning into plasmid backbone: Inserts the assembled gene into a plasmid for propagation and expression in cells.
- Sequence verification: Confirms the final DNA sequence is correct and free of synthesis errors.
Limitations
- Oligos longer than 200 nt are error-prone: Chemical synthesis efficiency decreases with length, increasing mutation rates.
- Repetitive elements reduce fidelity: Repeats promote misalignment during synthesis and assembly.
- GC-rich sequences are difficult: High GC content can impair synthesis and cloning efficiency.
- Cost increases with length: Longer constructs require more synthesis and validation steps.
- Cloning validation required: Errors introduced during synthesis must be identified by sequencing.
5.3 DNA Edit
(i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
- Edit melanocyte genomic DNA at a safe harbor locus (e.g., AAVS1).
- This allows stable insertion of a therapeutic construct without disrupting essential endogenous genes.
- Insert a ROS-responsive antioxidant genetic circuit. This enables melanocytes to activate protective genes only during oxidative stress, reducing unnecessary expression.
- Optionally enhance endogenous NRF2 pathway activity. Strengthening the native antioxidant response may improve resilience to oxidative damage implicated in vitiligo.
(ii) What technology or technologies would you use to perform these DNA edits and why?
I’ll use CRISPR-Cas9 genome editing. This system enables targeted DNA modification at precise genomic locations.
- Design a single guide RNA targeting the chosen safe harbor locus. The guide RNA determines the exact genomic cut site and ensures specificity.
- Provide a donor DNA template containing the ROS-sensing circuit with homology arms. Homology arms direct precise insertion via homology-directed repair.
- Deliver Cas9, guide RNA, and donor DNA into melanocytes ex vivo. Editing cells outside the body improves safety and allows selection of correctly modified cells.
- Screen and sequence edited clones. Verification ensures accurate insertion and absence of unintended mutations.
Also answer the following questions:
How does your technology of choice edit DNA? What are the essential steps?
- Cas9 introduces a double-strand break at the target site. The break initiates cellular DNA repair mechanisms.
- Cells repair the break using homology-directed repair when a donor template is present. This enables precise integration of the therapeutic circuit.
- The edited locus now contains the ROS-responsive construct. The inserted sequence becomes a permanent part of the melanocyte genome.
What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
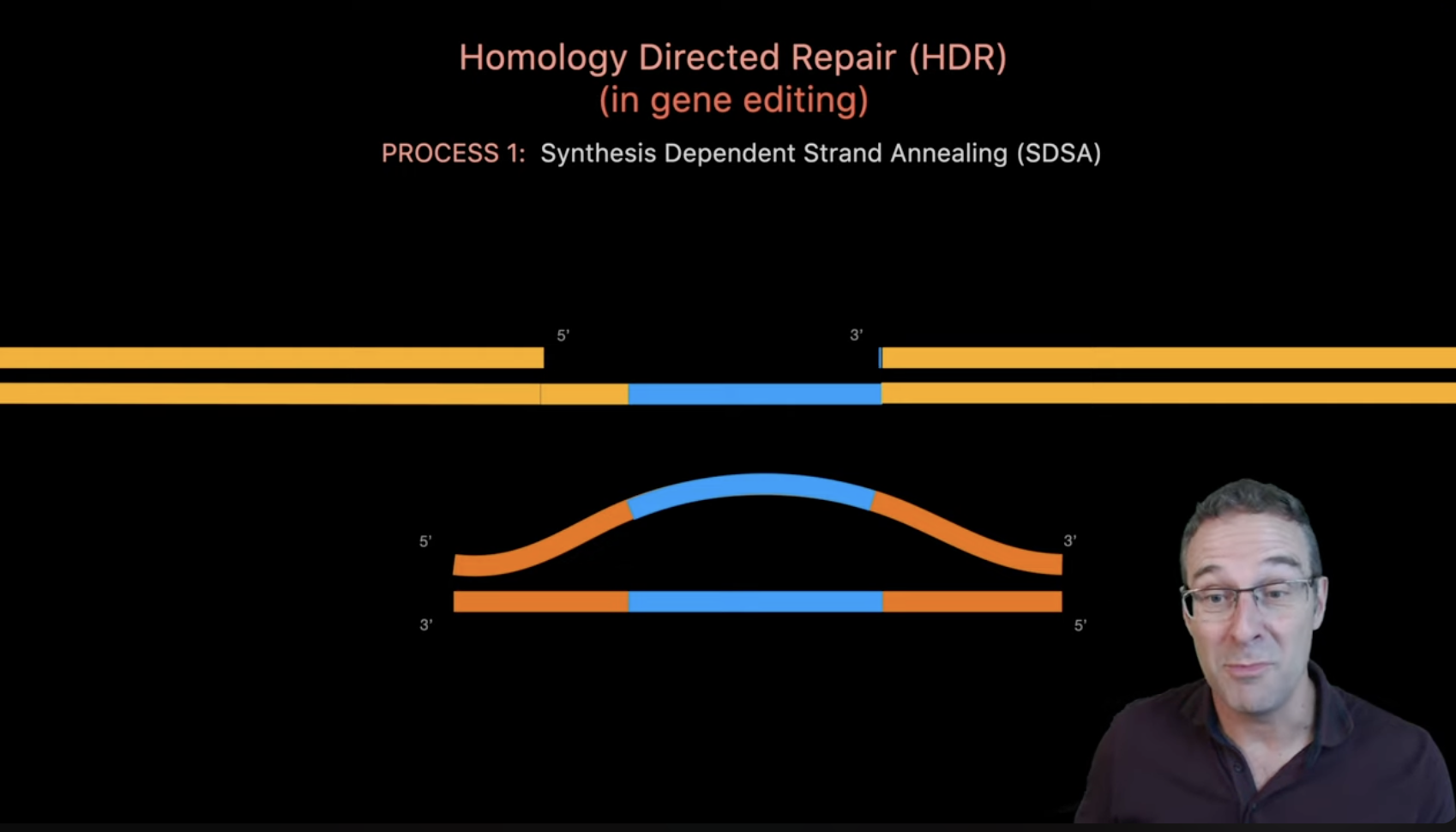
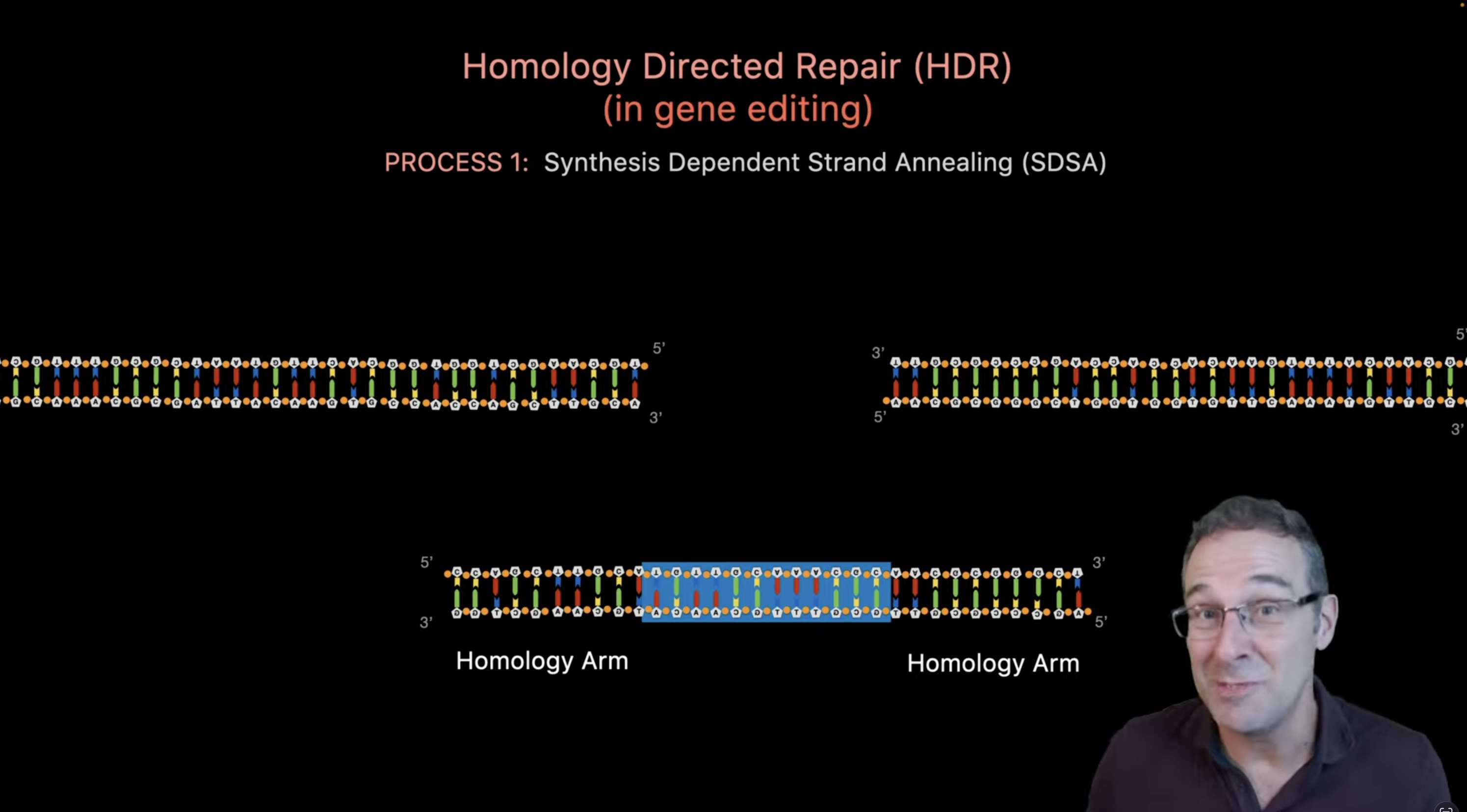




 (Source)[https://www.youtube.com/watch?v=cLMo6DYdJRE]
(Source)[https://www.youtube.com/watch?v=cLMo6DYdJRE]
- Cas9 protein or expression plasmid. Provides the nuclease that cuts DNA
- Single guide RNA (sgRNA). Directs Cas9 to the target locus
- Donor DNA template with homology arms. Supplies the sequence to be inserted.
- Cultured melanocytes: Serve as the editable cellular substrate.
What are the limitations of your editing methods (if any) in terms of efficiency or precision?
- Homology-directed repair efficiency can be low. Precise insertion may occur in only a fraction of treated cells.
- Off-target edits are possible. Imperfect guide specificity may lead to unintended genomic changes.
- Delivery into primary melanocytes can be challenging. Some cell types are difficult to transfect efficiently.
- Permanent genomic edits raise regulatory and safety concerns. Long-term monitoring would be required before clinical use.
Week 3 HW: Opentrons and automation tools
Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Semiautomated Production of Cell-Free Biosensors
https://www.sciencedirect.com/org/science/article/abs/pii/S2161506325000725
Cell-free biosensors combine the gene expression machinery of cells with engineered genetic circuits, enabling detection of molecules through measurable outputs like fluorescence (GFP) or color change (LacZ). These systems have unique advantages - they can be freeze-dried, distributed cheaply, and used at the point of need without living cells, making them promising platforms for environmental and health diagnostics. However, traditional workflows rely on manual pipetting to assemble reactions, which is slow and introduces variability in performance.
To overcome this bottleneck, the authors developed and evaluated a semiautomated manufacturing protocol using the Opentrons OT-2 liquid handling robot. They compared traditional manual assembly of reactions with an automated protocol and demonstrated the approach by constructing a full 384-well plate of fluoride-sensing cell-free biosensors. The reactions assembled by Opentrons performed comparably to expectations from manual assembly, supporting the idea that liquid-handling automation can improve scalability, reproducibility, and quality control in biosensor production.
Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
One of my ideas is to build a Programmable Regenerative Biofilter for Selective VOC Destruction
I would use the cloud lab to answer:
- Which detector design is most sensitive? Test 30-100 versions the promoter (the “on switch” strength), the regulator amount, the reporter (GFP glow vs luciferase glow)
- Which enzyme destroys formaldehyde fastest?
- Which setup lasts the longest before it wears out?
Find the best formaldehyde detector (it glows only when formaldehyde is present). Then find the best enzyme that destroys formaldehyde. Then test how to glue that enzyme onto a filter material so it works in a cartridge. Finally, combine detection and cleanup into a system that turns itself on only when formaldehyde is in the air.
Automated Build/Test Pipeline (Formaldehyde Focused):
1. Design & DNA Assembly (Build)
Upload formaldehyde sensor variants and enzyme gene sequences to the cloud lab → automated DNA synthesis, Gibson assembly, transformation, and sequence verification.
2. High-Throughput Sensor Screening (Test)
Express sensor constructs in 96-well cell-free reactions → expose to a formaldehyde concentration gradient → measure fluorescence to identify lowest detection threshold and highest signal-to-noise ratio.
3. Enzyme Activity Optimization (Test)
Express candidate formaldehyde-degrading enzymes in cell-free → add formaldehyde substrate → quantify degradation rate via colorimetric or absorbance assay → rank by turnover and stability.
4. Immobilization & Stability Assay (Build/Test)
Attach top enzyme candidates to bead or hydrogel matrices → wash and stress-test (heat, humidity simulation) → measure retained catalytic activity over time.
5. Closed-Loop Integration Test (Build/Test)
Combine best sensor + enzyme modules in automated plate workflow → confirm: formaldehyde detection triggers enzyme expression → formaldehyde concentration decreases over time → quantify response dynamics and regeneration capacity.
Week 4 HW: Protein Design Part 1
Part A. Conceptual Questions
How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
Hey, so I’m a vegetarian, so I don’t take a single gram of meat if I can avoid it.
However, 500 grams of meat is approximately 3.011e+26 Daltons. If an average amino acid is 100 Daltons, we’ll end up with 3.011e+24 amino acid molecules. That’s assuming that all of the meat is protein, which it isn’t. Meat is approximately 20% protein, so 20% of 500 is 100g protein, or 3.01e+24Why do humans eat beef but do not become cows, eat fish but do not become fish?
Because we’re not modifying the human DNA by eating fish or beef. Our DNA makes us who we are. What we eat is digested as macromolecules that become generic molecular building blocksWhy are there only 20 natural amino acids?
Evolutionarily, 20 amino acids may be a near-optimal solution balancing chemical diversity, folding reliability, metabolic cost, and coding efficiency. There is enough redundancy and error minimization that similar codons encode chemically similar amino acids. These amino acids cover the functional space of protein physiochemical dimensions (hydrophobicity, etc)Can you make other non-natural amino acids? Design some new amino acids.
Yes. Non-natural amino acids (nnAAs) can be designed and incorporated into proteins. For example, a photo-switchable amino acid can have light-driven switching which can help Light-controlled enzyme activationWhere did amino acids come from before enzymes that make them, and before life started?
Amino acids likely formed through:Lightning-driven atmospheric chemistry
Strecker-type reactions in aqueous environments
Hydrothermal vent redox chemistry
UV photochemistry
Delivery from meteorites
They are a natural product of carbon, nitrogen, hydrogen, and oxygen chemistry under energetic planetary conditions.
If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
Can you discover additional helices in proteins?
Yes we can both identify and design new helices using structural, computational, and synthetic approaches. We can discovery by mining structural databases, through computational protein design and through non-natural amino acids.Why are most molecular helices right-handed?
Most molecular helices in biology are right-handed because life is built from chiral building blocks of one dominant handedness, and that stereochemistry biases the lowest-energy helical geometry.Why do β-sheets tend to aggregate?
β-sheets tend to aggregate because their hydrogen-bonding pattern and geometry make intermolecular extension energetically favorable and difficult to cap. In other words, the structure is inherently “open-ended.”What is the driving force for β-sheet aggregation?
The driving force for β-sheet aggregation is free-energy minimization dominated by backbone hydrogen bonding and hydrophobic collapse, with cooperativity that makes extension energetically downhill once nucleated.Why do many amyloid diseases form β-sheets?
Many amyloid diseases involve β-sheets because the β-sheet architecture is the lowest-energy, self-templating structure accessible to partially unfolded polypeptides. When proteins misfold, β-structure is the most thermodynamically and kinetically available aggregation solution.Can you use amyloid β-sheets as materials?
Yes. Amyloid β-sheets are exceptionally robust, nanoscale materials. The cross-β architecture is one of the strongest naturally occurring protein material motifs. Amyloid fibrils are mechanically stiff, chemically stable, and self-assembling, which are all desirable properties for materials engineering.Design a β-sheet motif that forms a well-ordered structure.
Part B: Protein Analysis and Visualization
Briefly describe the protein you selected and why you selected it.
I picked histone-based chromatin https://pdb101.rcsb.org/motm/314, which is the molecule of the month on PDB and arrived at archeal histone https://www.uniprot.org/uniprotkb/A0A060HNU7/entry#sequences. I thought their biological assembly looked cool - it has nice symmetry because they act as spools for DNA. I also found it interesting that Histones were long thought to be unique to the eukaryotic lineage. But recent studies have shown that histones exist in most archaeal and some bacterial cells, shedding light on DNA packaging mechanisms while also raising new questions about the evolution of histones.Identify the amino acid sequence of your protein.
MSSGPEFGLAAMYRIMKKSGAERVSDDAADELRKVLEEVAERIAKQAAELSMHAGRKTIKPEDIRLASKNVIRLHow long is it? What is the most frequent amino acid? You can use this Colab notebook to count the frequency of amino acids.
It’s 74 amino acids long. The most common amino acid is: A, which appears 11 times.How many protein sequence homologs are there for your protein? Hint: Use Uniprot’s BLAST tool to search for homologs.
The BLAST tool found 250 results in UniProtKBDoes your protein belong to any protein family?
The sequence similarity shows it belongs to the archaeal histone HMF familyIdentify the structure page of your protein in RCSB
There isn’t an exact match. The closest similarity search results in CRYSTAL STRUCTURE OF THE HISTONE HMFA FROM METHANOTHERMUS FERVIDUS https://www.rcsb.org/3d-view/1B67When was the structure solved? Is it a good quality structure? Good quality structure is the one with good resolution. Smaller the better (Resolution: 2.70 Å)
Yes, it’s very well resolved for the similar protein in RCSB
Are there any other molecules in the solved structure apart from protein?
NoDoes your protein belong to any structure classification family?
No it doesn’t.Open the structure of your protein in any 3D molecule visualization software:
PyMol Tutorial Here (hint: ChatGPT is good at PyMol commands)
Visualize the protein as “cartoon”, “ribbon” and “ball and stick”.
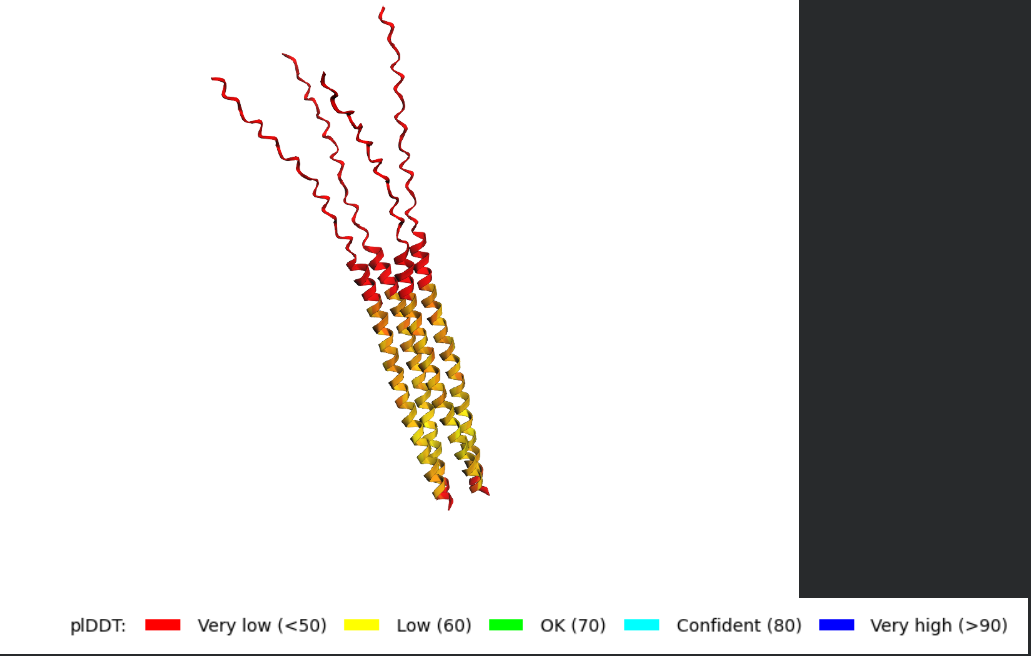
Cartoon: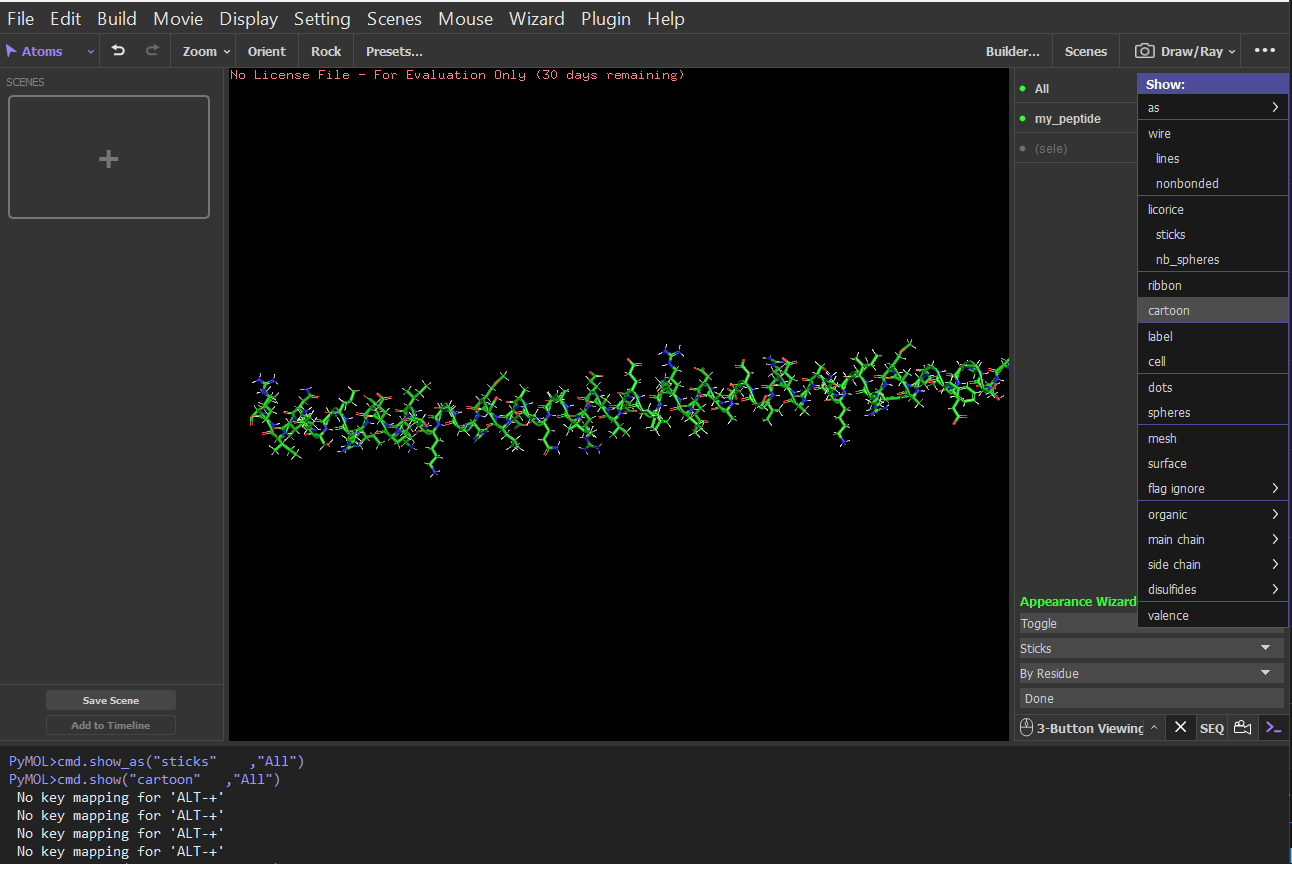
 Ribbon:
Ribbon: 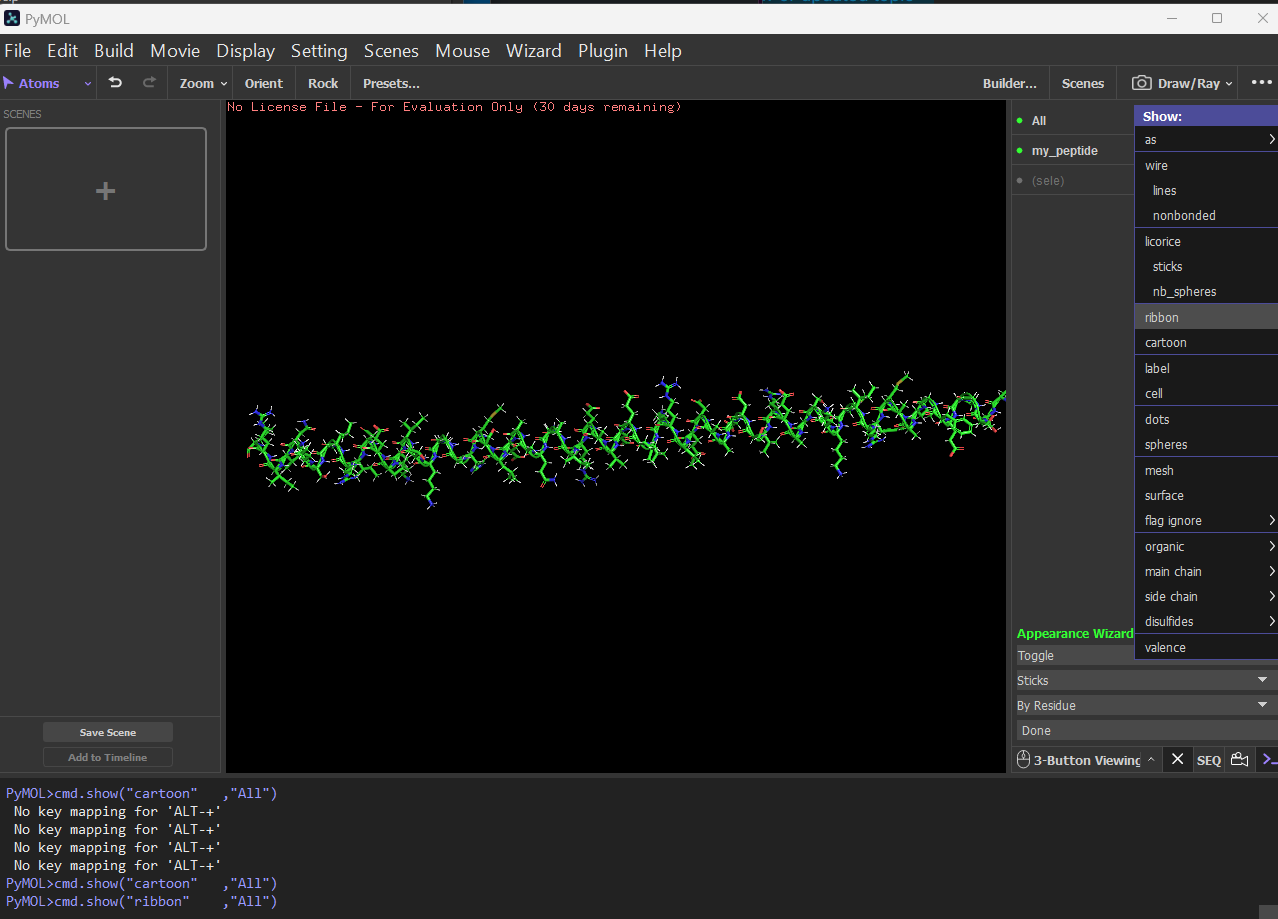
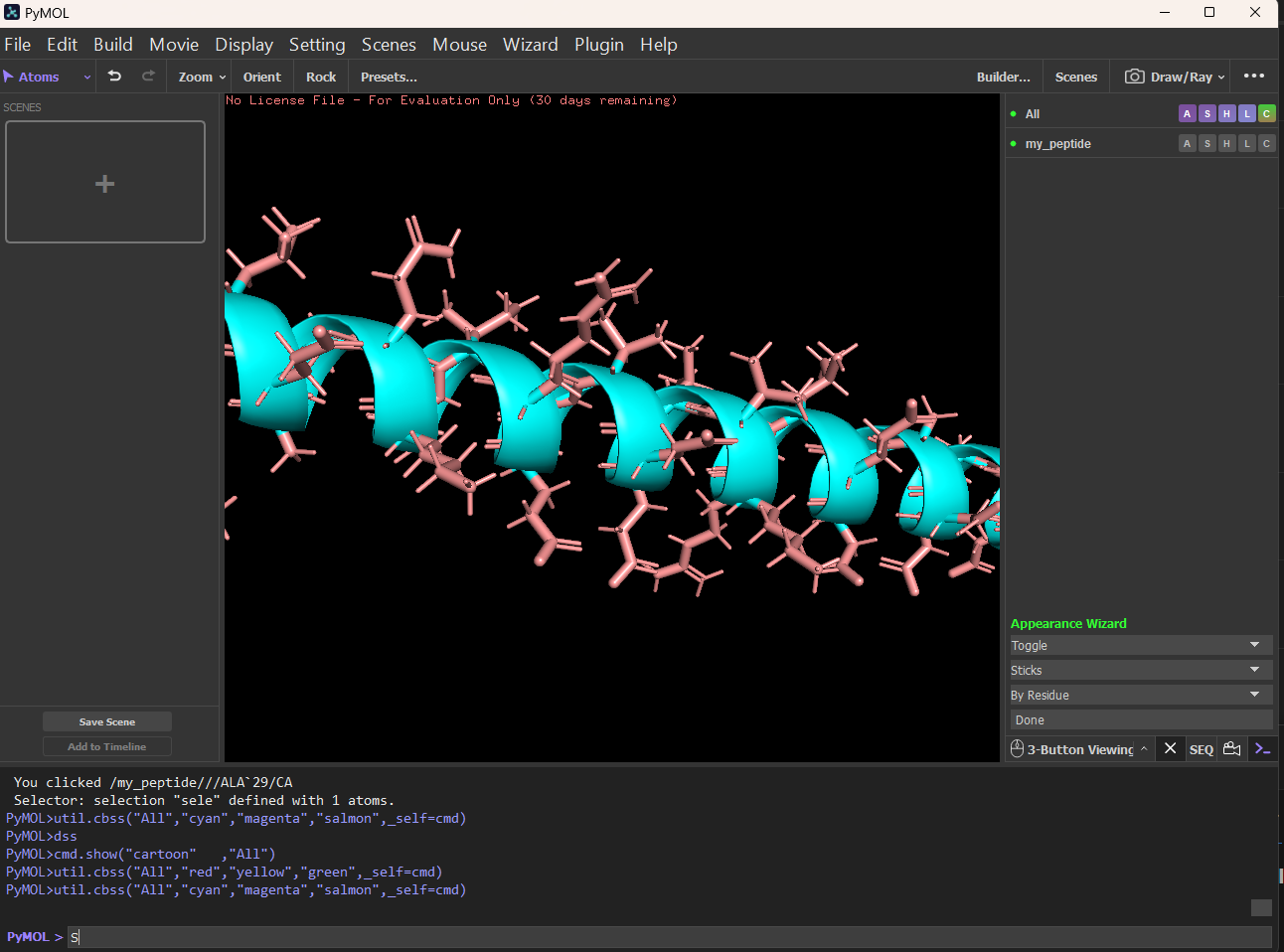
 Sticks:
Sticks: 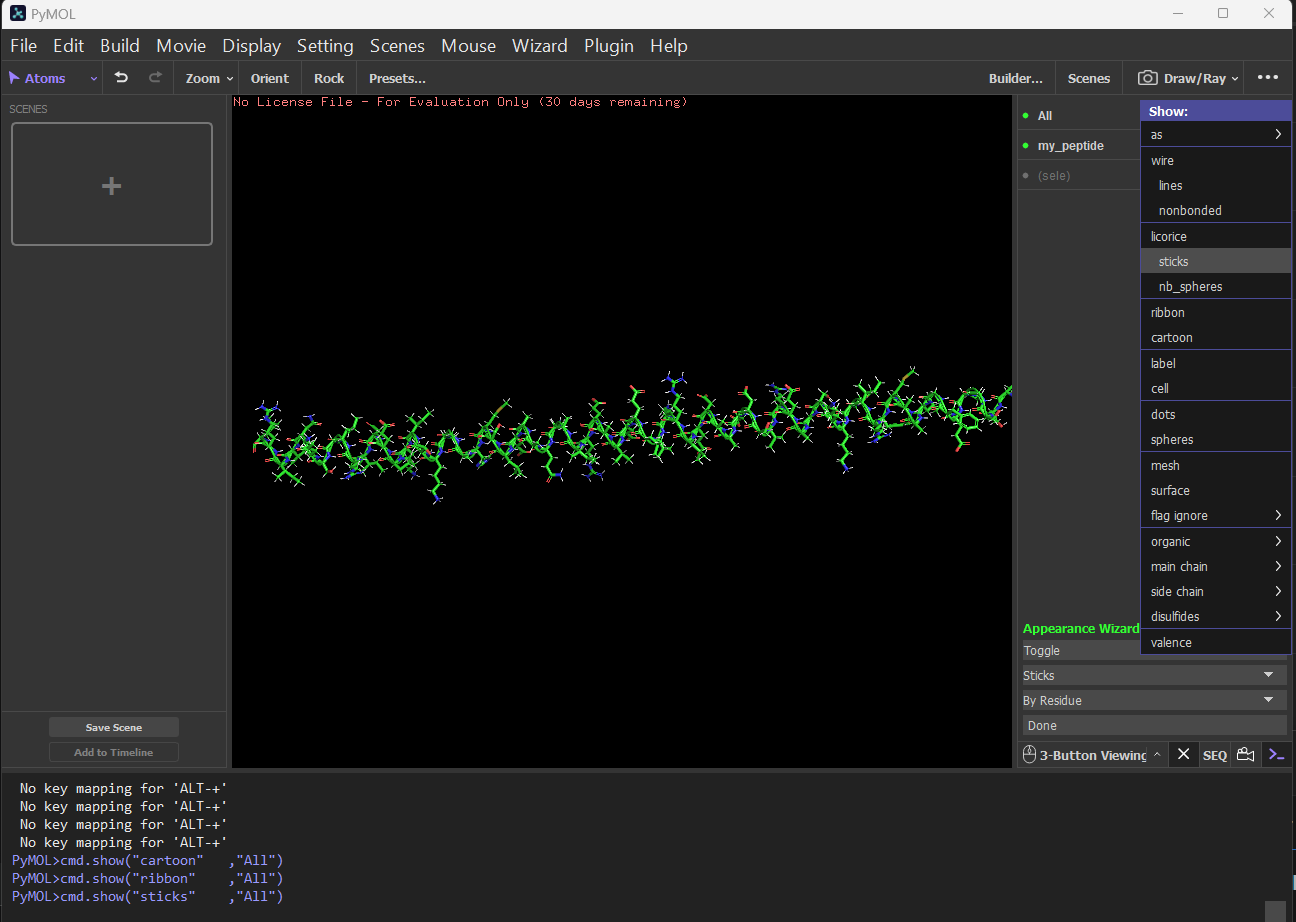

Color the protein by secondary structure. Does it have more helices or sheets?
It has more helices and no sheets. It has loops. \Color the protein by residue type. What can you tell about the distribution of hydrophobic vs hydrophilic residues?
| Property | Hydrophobic (Orange) | Hydrophilic (Marine) |
|---|---|---|
| Common Residues | Leu, Ile, Val, Phe | Ser, Thr, Asp, Glu, Lys |
| Typical Location | Buried in the core | Exposed on the surface |
| Role | Stability & Folding (Hydrophobic effect) | Solubilization & Ligand Binding |
- Visualize the surface of the protein. Does it have any “holes” (aka binding pockets)?
There are no true internal binding pockets in this structure. structure is still a single, extended α-helix (likely a coiled or amphipathic helix), not a folded globular domain. A binding pocket requires:
- Tertiary packing
- Enclosed or partially enclosed cavity volume
- Multiple secondary structure elements forming a concave region
A single helix is essentially a cylinder with protruding side chains. Cylinders do not form pockets unless:
- Multiple helices pack together (helix bundle)
- The helix bends sharply
- Oligomerization creates an interface groove
Part C:
- Deep Mutational Scans
a. Use ESM2 to generate an unsupervised deep mutational scan of your protein based on language model likelihoods.
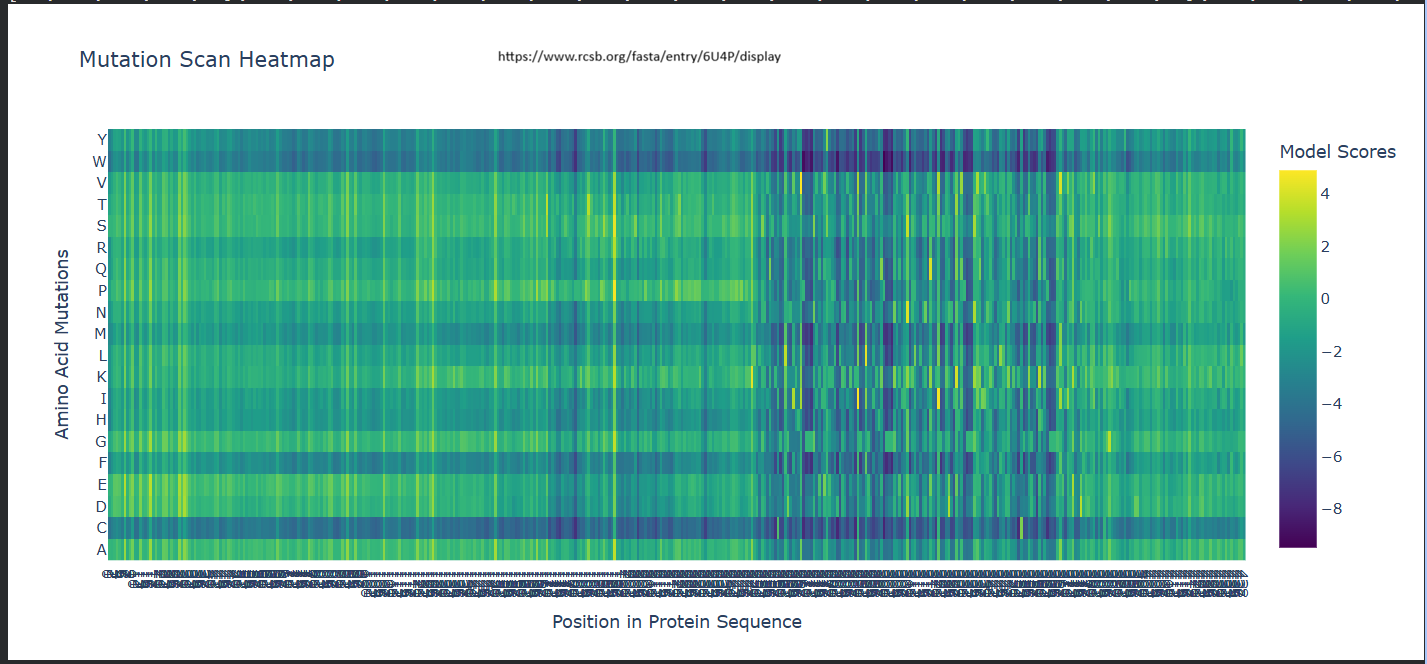

b. Can you explain any particular pattern? (choose a residue and a mutation that stands out)
It seems like for position 333, Wildtype residue G, It is highly likey for the amino acid to be
-– Scores for Position 331 (Wild-type Residue: P) -–
Amino Acid (Y) | Score (Z)
-————————
A | 0.2499
C | -4.0639
D | -2.3773
E | -1.1166
F | -2.0289
G | -1.5281
H | -1.6049
I | 0.9267
K | 0.0000
L | 0.5773
M | -0.5974
N | -1.6772
P | -0.3847
Q | 1.5133
R | -1.1004
S | -0.9229
T | -0.0714
V | 2.6539
W | -3.7854
Y | -2.9304
My first question was why isn’t the Wild-Type (P) exactly 0?
I understood that when the code calculates these scores, it masks (hides) the residue at position 331 and asks the model: ‘Given the rest of the protein, what should be here?’
The Model’s Preference: The model thinks Lysine (K) fits that spot slightly better than the original Proline (P). Since the ‘Relative’ score is log(P_mutation) - log(P_wildtype), and the model happened to predict Lysine with a probability identical to what it expected for a wild-type residue in that context, it came out to 0.
The Penalty for P: The -0.3847 for Proline means that when the model looks at the surrounding sequence, it’s actually a bit ‘surprised’ to see a Proline there compared to its top choices (like Valine or Lysine).
The Baseline: The true ‘baseline’ is always the Wild-Type residue’s original probability. Any amino acid with a positive score is something the model likes more than the original, and anything negative is something it likes less.
Positive Scores (e.g., V: 2.6539): These are favored mutations. A positive score means the model thinks Valine is actually more likely to be found at this position than the original Proline. This could suggest the mutation might increase stability or is common in similar protein sequences evolutionarily.
Negative Scores (e.g., C: -4.0639): These are disfavored mutations. A negative score indicates the model predicts this change is unlikely or potentially destabilizing. The more negative the number, the more ‘surprised’ the model would be to see that amino acid there.
-–
Latent Space Analysis
a. Use the provided sequence dataset to embed proteins in reduced dimensionality.
Done in code.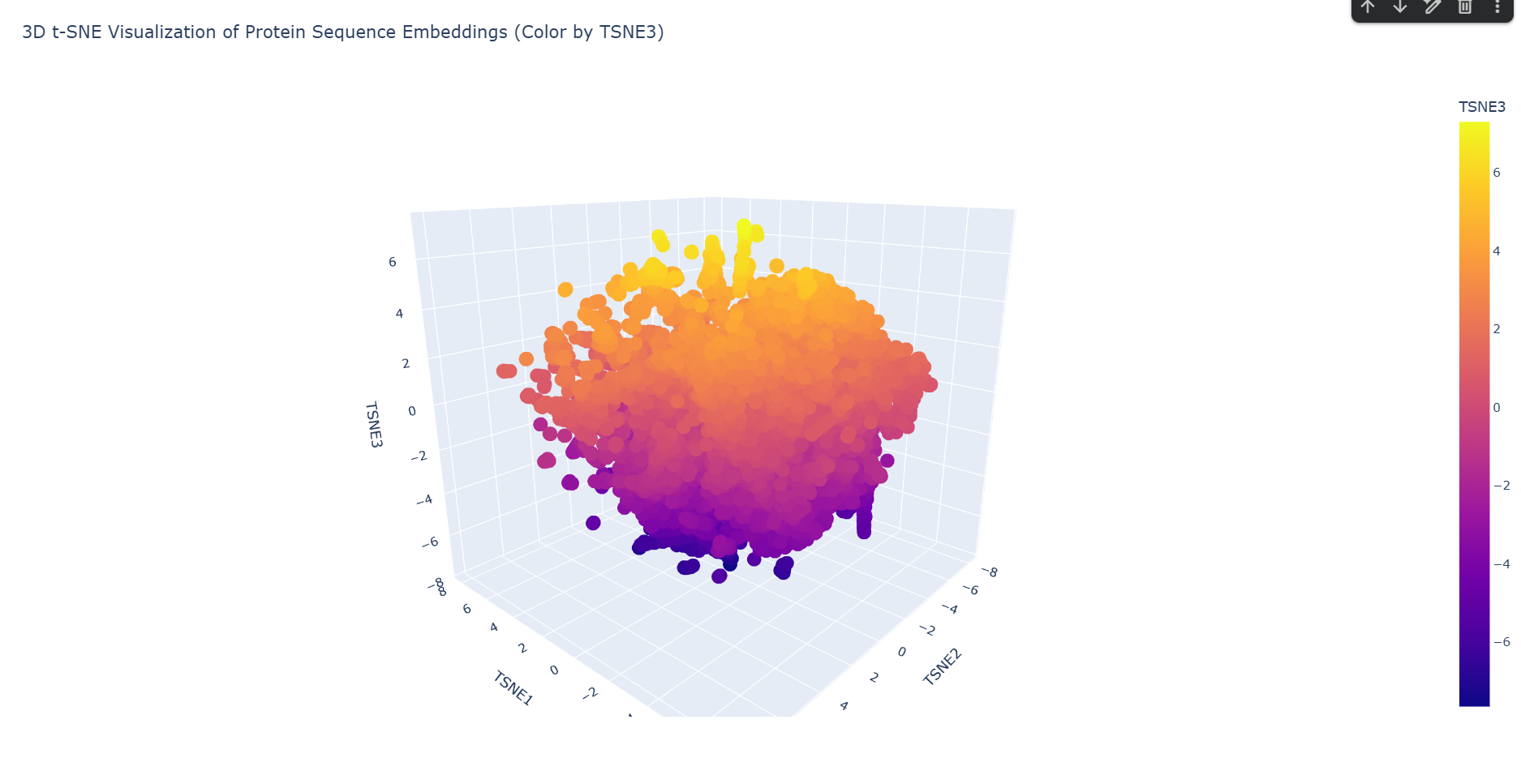
 b. Analyze the different formed neighborhoods: do they approximate similar proteins?
b. Analyze the different formed neighborhoods: do they approximate similar proteins?
Yes. Based on the structural neighborhood report, Cluster 6 is the most ‘pure’ (defined as the highest percentage of its dominant fold after ‘automated matches’ are set aside).Here are the top 5 most common structural descriptions in that neighborhood:
- T-cell antigen receptor (Dominant Motif)
- Transglutaminase, two C-terminal domains
- Fibronectin type-III domai ncontaining protein 3a
- Neogenin
- Chondroitinase AC
This neighborhood primarily clusters proteins with Immunoglobulin-like folds and large multi-domain enzymatic proteins, showing that the latent space successfully groups these complex beta-sheet rich structures together.
c. Place your protein in the resulting map and explain its position and similarity to its neighbors.
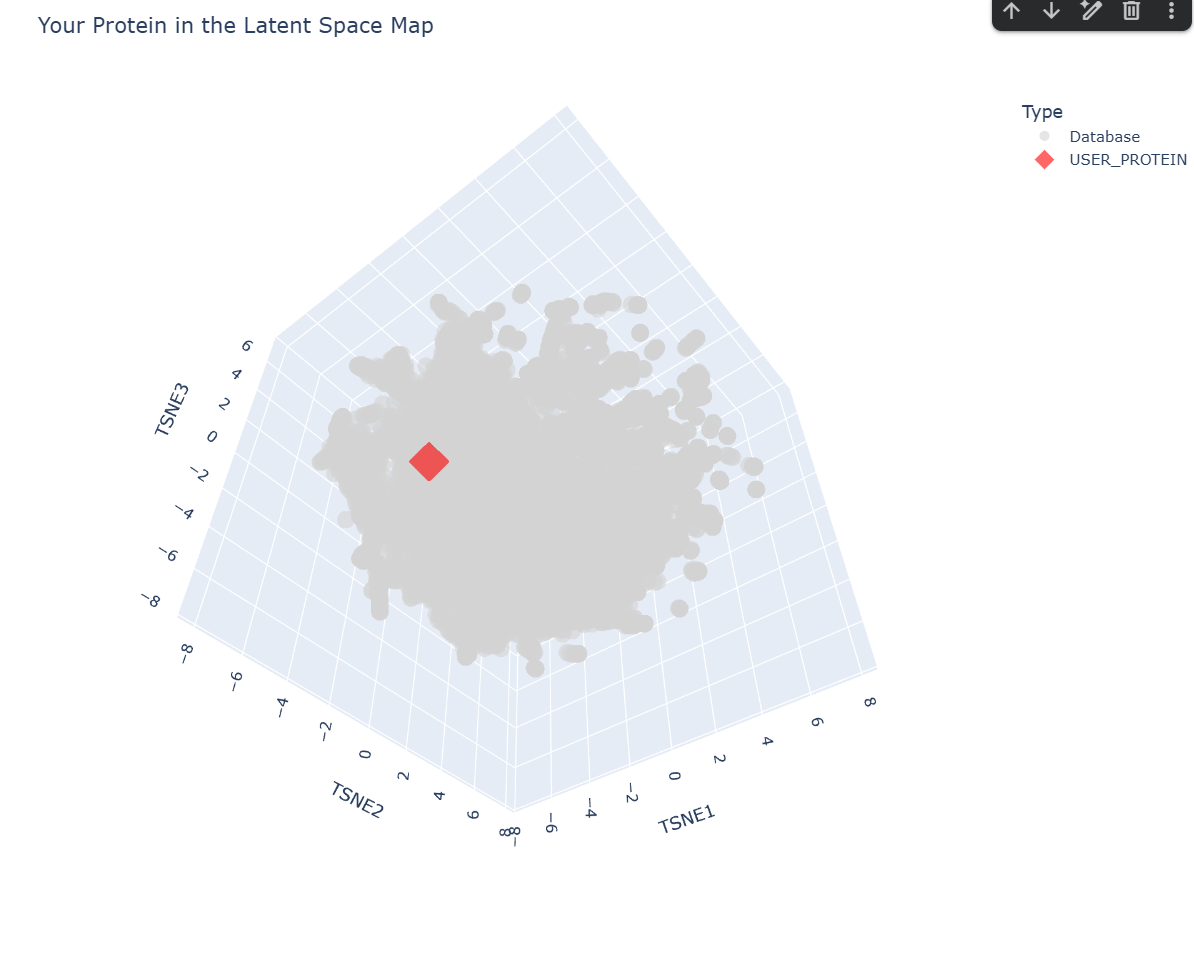

-– Top 10 Nearest Structural Neighbors to myProtein -–
1. Distance: 0.3048 | Description: d2ffta1 g.88.1.1 (A:2-84) Thylakoid soluble phosphoprotein TSP9 {Spinach (Spinacia oleracea) [TaxId: 3562]}
2. Distance: 0.5750 | Description: d2b5lc_ d.384.1.1 (C:) SV5-V core {Simian virus 5 [TaxId: 11207]}
3. Distance: 0.6194 | Description: d4lmsa_ d.184.1.0 (A:) automated matches {Chroomonas sp. [TaxId: 3029]}
4. Distance: 0.6318 | Description: d1xg0a_ d.184.1.1 (A:) Phycoerythrin 545 alpha-subunits {Cryptophyte (Rhodomonas sp. CS24) [TaxId: 79257]}
5. Distance: 0.8230 | Description: d1pbyc_ a.137.9.1 (C:) Quinohemoprotein amine dehydrogenase C chain {Paracoccus denitrificans [TaxId: 266]}
6. Distance: 0.8820 | Description: d4it7a_ d.17.1.0 (A:) automated matches {Ascaris lumbricoides [TaxId: 6252]}
7. Distance: 0.9125 | Description: d5ez2a_ b.60.1.0 (A:) automated matches {Sander vitreus [TaxId: 283036]}
8. Distance: 0.9148 | Description: d4m0wa2 d.3.1.23 (A:63-319) automated matches {SARS coronavirus [TaxId: 227859]}
9. Distance: 0.9433 | Description: d3nwaa_ e.76.1.1 (A:) Glycoprotein B {Human herpesvirus 1 [TaxId: 10298]}
10. Distance: 0.9935 | Description: d2a73a6 g.94.1.1 (A:578-643) Complement C3 linker domain {Human (Homo sapiens) [TaxId: 9606]}
Based on the ESM-2 latent space analysis, my protein’s closest structural neighbor is TSP9 (Thylakoid soluble phosphoprotein) from Spinach.
Analysis of Neighbors:
- TSP9 (Distance: 0.3048): This is a very close match in the embedding space. TSP9 is often associated with photosynthesis and has highly flexible regions.
- Viral and Algal Proteins: The presence of SV5-V core and Phycoerythrin in the top neighbors suggests your protein might share motifs with proteins involved in large complex assemblies or pigment binding.
- Structural Diversity: The neighbors include a mix of alpha-helical and linker domains (like the Complement C3 linker). This suggests your protein might have a modular or relatively disordered/flexible structure, which is common for proteins that show up near ’linker’ or ‘phosphoprotein’ domains in the latent space.
C2. Protein Folding
Folding a protein
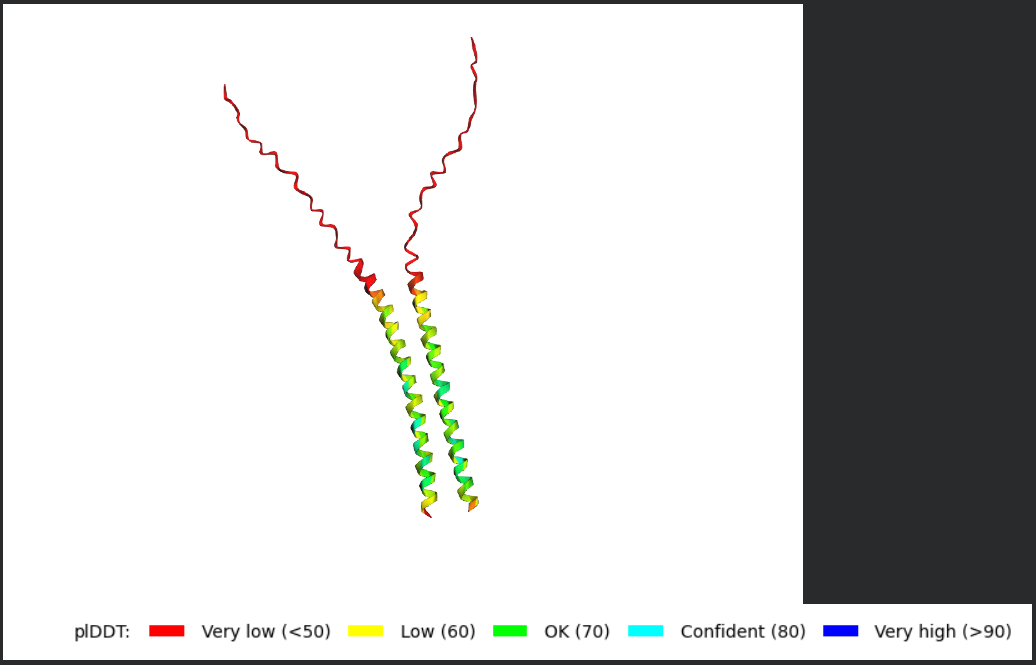
- Fold your protein with ESMFold. Do the predicted coordinates match your original structure?
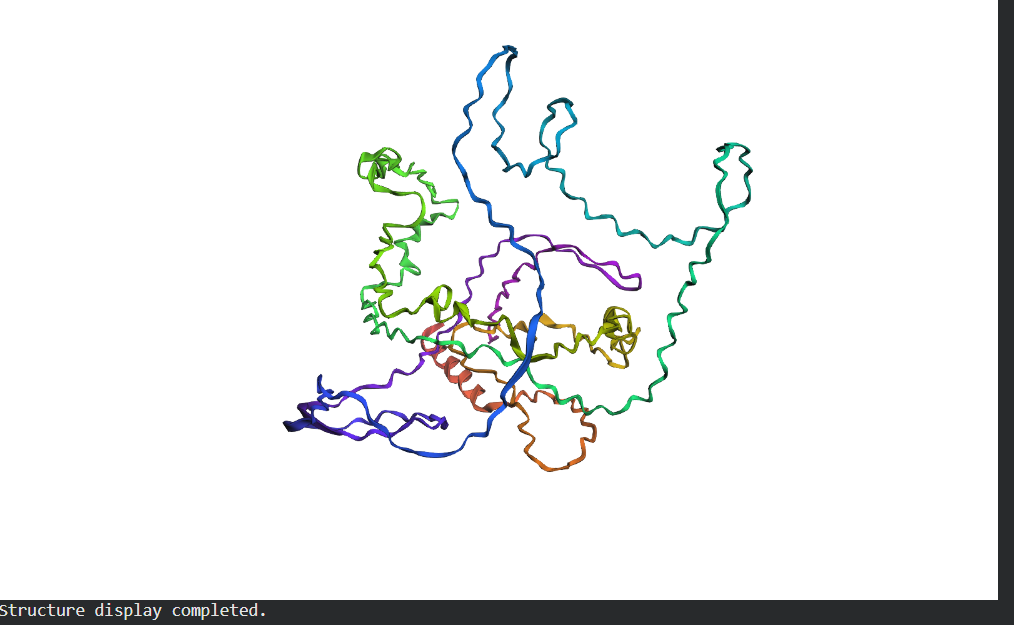

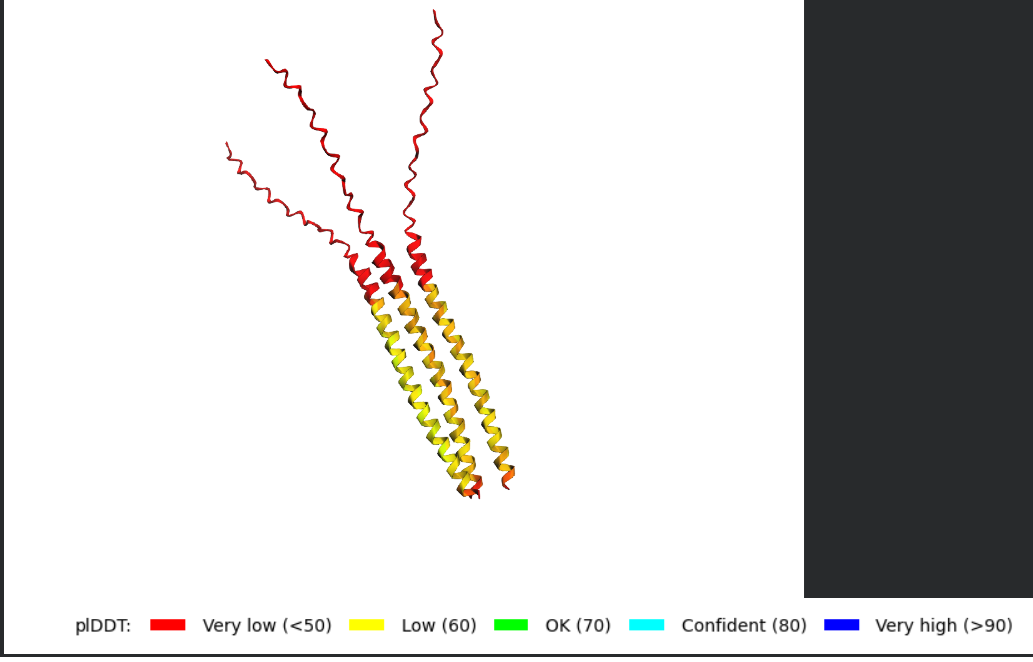
- Try changing the sequence, first try some mutations, then large segments. Is your protein structure resilient to mutations?
Folded Protein Small Change change:

Folded Protein Big Change change:


## C3. Protein Generation
Inverse-Folding a protein: Let’s now use the backbone of your chosen PDB to propose sequence candidates via ProteinMPNN
Generating sequences...
>10KR, score=2.4363, fixed_chains=[], designed_chains=[‘A’], model_name=v_48_020
QIVYKPVDLSKVTSKCGSLGNIHHKPGGGQMEVKSEKLDFKDRVQSKIGSLDNITHVPGGGNKKIETHKLT
>T=0.1, sample=0, score=1.3713, seq_recovery=0.2535
TVVVTPVVTVVTTVTTGAVGTVVVDPGGGEEVVTVEGVVVTGPVVVEVGVHGVTVTTPAGGTVVTVVVTET
New Sequence:TVVVTPVVTVVTTVTTGAVGTVVVDPGGGEEVVTVEGVVVTGPVVVEVGVHGVTVTTPAGGTVVTVVVTET
Analyze the predicted sequence probabilities and compare the predicted sequence vs the original one.
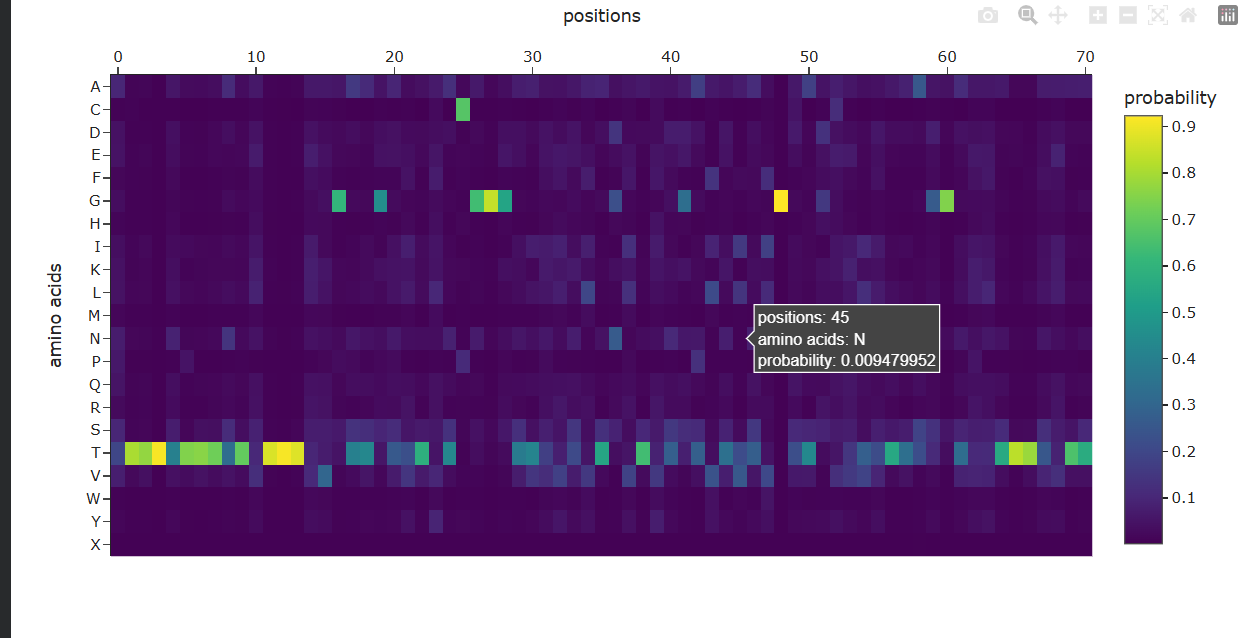

Input this sequence into ESMFold and compare the predicted structure to your original.

Week 5 HW: Protein Design Part 2
Part A: SOD1 Binder Peptide Design (From Pranam)
Part 1: Generate Binders with PepMLM
Begin by retrieving the human SOD1 sequence from UniProt (P00441) and introducing the A4V mutation. Unprot Link for SOD1 sequence
MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQMutant SOD1 sequence A4V:
MATVAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQUsing the PepMLM Colab linked from the HuggingFace PepMLM-650M model card
My notebookGenerate four peptides of length 12 amino acids conditioned on the mutant SOD1 sequence.
To your generated list, add the known SOD1-binding peptide FLYRWLPSRRGG for comparison.
Record the perplexity scores that indicate PepMLM’s confidence in the binders.
Binder Pseudo Perplexity WRYPAAALALAE 6.78294 WSYGVVALALGK 10.1846 WRYPVAGLGWAE 20.4686 WRSPAAAAGHGK 8.71141 FLYRWLPSRRGG 20.5338
Part 2 Evaluate Binders with Alphafold3
- Navigate to the AlphaFold Server: alphafoldserver.com
- For each peptide, submit the mutant SOD1 sequence followed by the peptide sequence as separate chains to model the protein-peptide complex.
- Peptide 1: WRYPAAALALAE
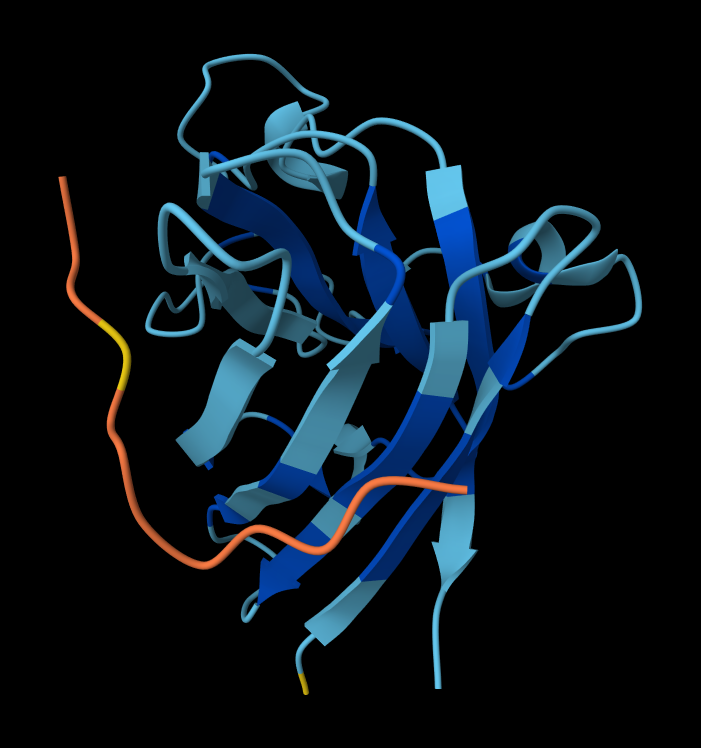

- Peptide 2: WSYGVVALALGK
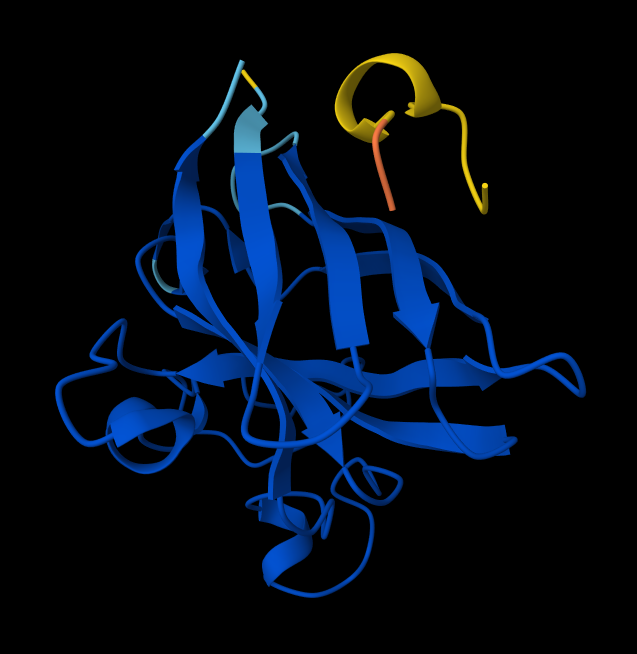

- Peptide 3: WRYPVAGLGWAE
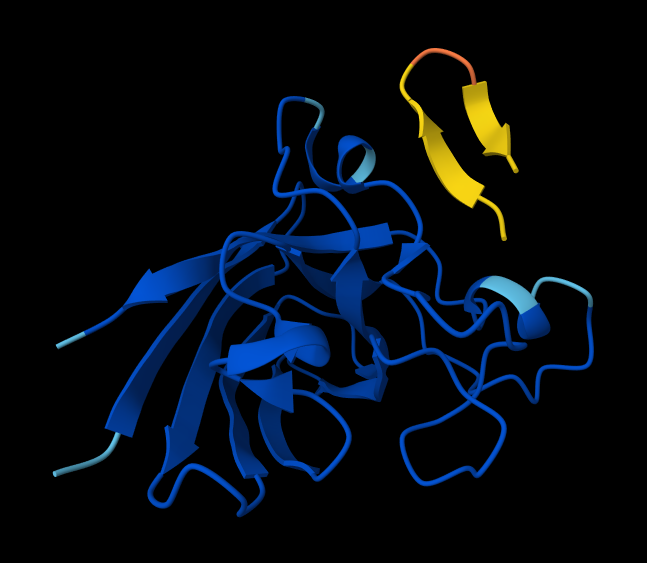

- Peptide 4: WRYPVAGLGWAE
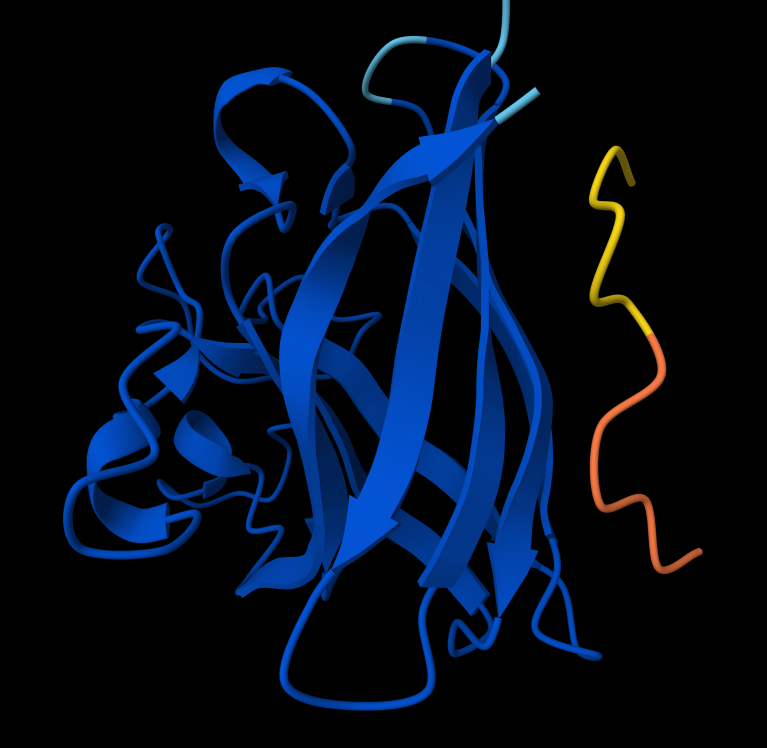

Interaction with N-terminus WRYPVAGLGWAE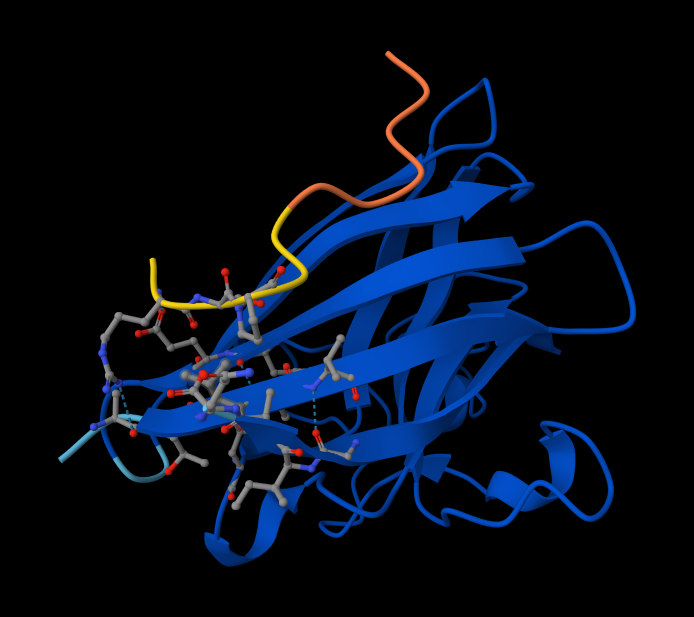

- Peptide 5: FLYRWLPSRRGG
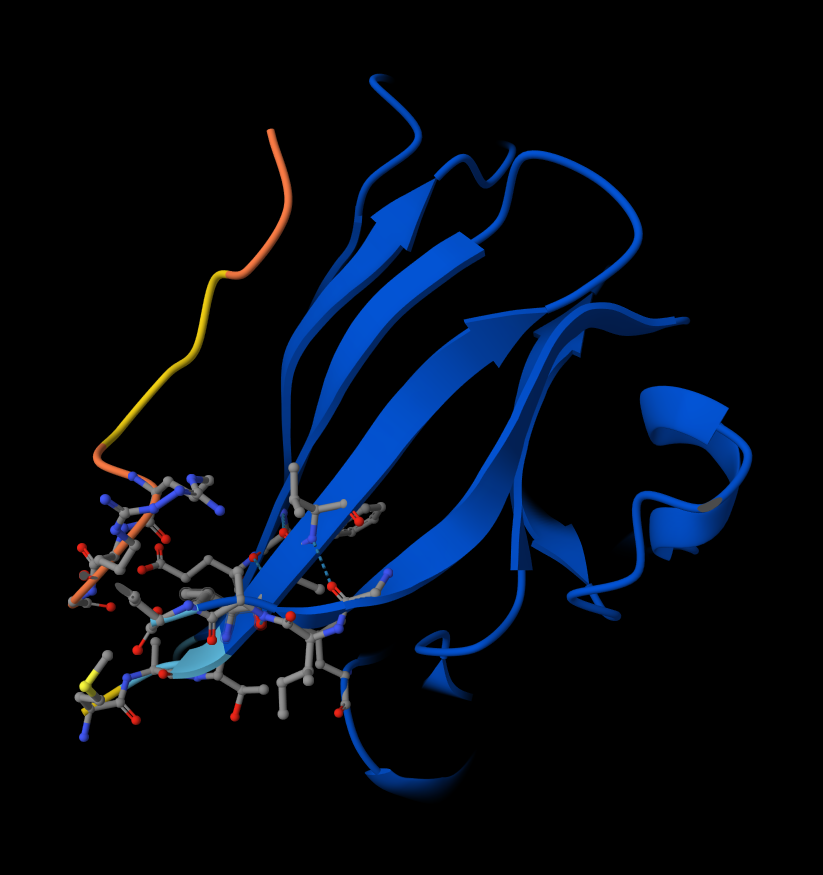

- Record the ipTM score and briefly describe where the peptide appears to bind. Does it localize near the N-terminus where A4V sits? Does it engage the β-barrel region or approach the dimer interface? Does it appear surface-bound or partially buried?
Here are ipTM and pTM obtained from Alphafold:# Sequence ipTM pTM 1 WRYPAAALALAE 0.27 0.76 2 WSYGVVALALGK 0.56 0.85 3 WRYPVAGLGWAE 0.43 0.89 4 WRSPAAAAGHGK 0.51 0.89 5 FLYRWLPSRRGG 0.37 0.82
- There is NO interaction with N-terminus of sequence 2 WSYGVVALALGK
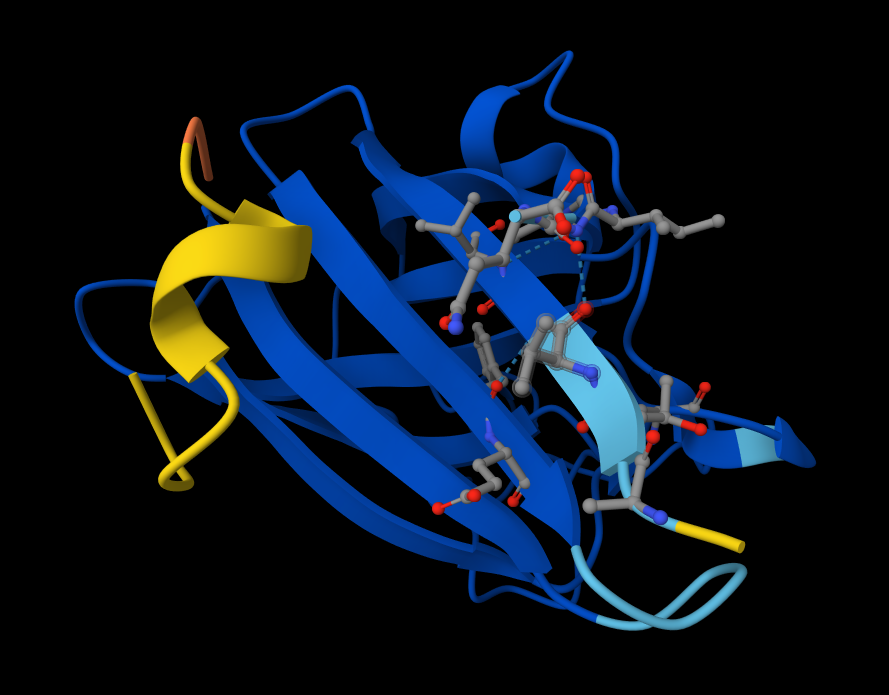

- There IS Interaction with N-terminus of sequence 4 WRYPVAGLGWAE


- There IS Interaction with N-terminus of Peptide 5: FLYRWLPSRRGG
Peptide 5: FLYRWLPSRRGG

In a short paragraph, describe the ipTM values you observe and whether any PepMLM-generated peptide matches or exceeds the known binder.
When comparing the PepMLM-generated sequences (1–4) against your known binder, Sequence 5 (FLYRWLPSRRGG), several candidates show marked improvements in both overall structural confidence (pTM) and interface interaction confidence (ipTM). The known binder establishes a baseline ipTM of 0.37 and a pTM of 0.82. Except for Sequence 1, which underperforms across both metrics, all other generated peptides successfully exceed these baselines. Notably, Sequence 2 achieves the highest interaction confidence with an ipTM of 0.56, while Sequences 3 and 4 also demonstrate strong improvements with ipTMs of 0.43 and 0.51, respectively, alongside excellent pTM scores of 0.89. Ultimately, multiple PepMLM-generated peptides do not just match but confidently exceed the predicted binding metrics of your known binder.
Peptide #4 seems promising. Here is why! Metrics Comparison (Sequence 4 vs. Sequence 5) Sequence 4 shows a significant improvement over your known binder, Sequence 5 (FLYRWLPSRRGG):
- Higher Interface Confidence: Sequence 4 boasts an ipTM of 0.51, compared to Sequence 5’s 0.37. This means AlphaFold is much more confident in the exact physical interaction between this peptide and SOD1.
- Higher Overall Confidence: The pTM of 0.89 (up from 0.82) indicates a highly stable and confident overall structural prediction for the entire complex.
Structural Analysis of Sequence 4 Looking at the 3D viewer in your image, your structural observations are highly accurate:N-Terminal Localization (A4V region):
- N-Terminal Localization (A4V region): The binder (shown in yellow/orange) is positioned right against the outer edge of the structural core. In SOD1, the N-terminus (containing the Alanine at position 4, known for the A4V ALS mutation) forms the first $\beta$-strand on the edge of the barrel. Your binder is directly draped over this exact region.
- Engaging the $\beta$-barrel: The peptide is actively interacting with the flat, exposed face of the $\beta$-barrel. You can see the side chains of the binder (the stick representations) pointing down and making contact with the blue $\beta$-sheets.
- Surface-Bound vs. Buried: The peptide is entirely surface-bound. It lies stretched across the convex exterior of the $\beta$-barrel like a strap. It does not penetrate deep into any hydrophobic pockets, nor is it partially buried within the core of the protein.
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
Input Property Prediction Value Unit WRYPAAALALAE 💧 Solubility Soluble 1.000 Probability 🩸 Hemolysis Non-hemolytic 0.033 Probability 🔗 Binding Affinity Weak binding 5.621 pKd/pKi 📏 Length 12 aa ⚖️ Molecular Weight 1331.5 Da ⚡ Net Charge (pH 7) -0.23 🎯 Isoelectric Point 6.22 pH 💦 Hydrophobicity (GRAVY) 0.40 GRAVY — — — — — WSYGVVALALGK 💧 Solubility Soluble 1.000 Probability 🩸 Hemolysis Non-hemolytic 0.122 Probability 🔗 Binding Affinity Weak binding 6.984 pKd/pKi 📏 Length 12 aa ⚖️ Molecular Weight 1263.5 Da ⚡ Net Charge (pH 7) 0.76 🎯 Isoelectric Point 8.59 pH 💦 Hydrophobicity (GRAVY) 0.99 GRAVY — — — — — WRYPVAGLGWAE 💧 Solubility Soluble 1.000 Probability 🩸 Hemolysis Non-hemolytic 0.056 Probability 🔗 Binding Affinity Weak binding 6.320 pKd/pKi 📏 Length 12 aa ⚖️ Molecular Weight 1404.6 Da ⚡ Net Charge (pH 7) -0.23 🎯 Isoelectric Point 6.22 pH 💦 Hydrophobicity (GRAVY) -0.16 GRAVY — — — — — WRSPAAAAGHGK 💧 Solubility Soluble 1.000 Probability 🩸 Hemolysis Non-hemolytic 0.015 Probability 🔗 Binding Affinity Weak binding 4.939 pKd/pKi 📏 Length 12 aa ⚖️ Molecular Weight 1208.3 Da ⚡ Net Charge (pH 7) 1.85 🎯 Isoelectric Point 11.00 pH 💦 Hydrophobicity (GRAVY) -0.71 GRAVY — — — — — FLYRWLPSRRGG 💧 Solubility Soluble 1.000 Probability 🩸 Hemolysis Non-hemolytic 0.047 Probability 🔗 Binding Affinity Weak binding 6.029 pKd/pKi 📏 Length 12 aa ⚖️ Molecular Weight 1507.7 Da ⚡ Net Charge (pH 7) 2.76 🎯 Isoelectric Point 11.71 pH 💦 Hydrophobicity (GRAVY) -0.71 GRAVY Let’s Transpose this making it easier to compare.
# Input 💧 Solubility Prediction 💧 Solubility (Prob.) 🩸 Hemolysis Prediction 🩸 Hemolysis (Prob.) 🔗 Binding Prediction 🔗 Binding Affinity (pKd/pKi) 📏 Length (aa) ⚖️ MW (Da) ⚡ Net Charge (pH 7) 🎯 pI (pH) 💦 Hydrophobicity (GRAVY) 1 WRYPAAALALAE Soluble 1.000 Non-hemolytic 0.033 Weak binding 5.621 12 1331.5 -0.23 6.22 0.40 2 WSYGVVALALGK Soluble 1.000 Non-hemolytic 0.122 Weak binding 6.984 12 1263.5 0.76 8.59 0.99 3 WRYPVAGLGWAE Soluble 1.000 Non-hemolytic 0.056 Weak binding 6.320 12 1404.6 -0.23 6.22 -0.16 4 WRSPAAAAGHGK Soluble 1.000 Non-hemolytic 0.015 Weak binding 4.939 12 1208.3 1.85 11.00 -0.71 5 FLYRWLPSRRGG Soluble 1.000 Non-hemolytic 0.047 Weak binding 6.029 12 1507.7 2.76 11.71 -0.71 Here is a comparison of the PeptiVerse predictions with the AlphaFold structural observations based on the data provided:
Part 4: PeptiVerse vs. AlphaFold Comparison
When comparing the AlphaFold structural models to the PeptiVerse property predictions, there is a partial but imperfect alignment between the two tools. AlphaFold identifies Peptide 2 (ipTM 0.56) and Peptide 4 (ipTM 0.51) as having the highest interaction confidence. PeptiVerse partially agrees, predicting Peptide 2 to have the strongest binding affinity (6.984 pKd/pKi) among the candidates. However, the correlation breaks down with Peptide 4; despite its strong AlphaFold interaction score and visually confirmed target engagement at the N-terminus, PeptiVerse predicts it to have the weakest binding affinity of the group (4.939 pKd/pKi).
To answer the specific questions:
- Do peptides with higher ipTM also show stronger predicted affinity? Only sometimes. While Peptide 2 boasts both the highest ipTM and the highest PeptiVerse affinity, Peptide 4 shows a stark contrast (high ipTM, lowest PeptiVerse affinity).
- Are any strong binders predicted to be hemolytic or poorly soluble? No. All evaluated peptides, including the strongest predicted binders, are uniformly predicted to be highly soluble (1.000 probability) and non-hemolytic, with hemolysis probabilities remaining comfortably low across the board (between 0.015 and 0.122).
- Which peptide best balances predicted binding and therapeutic properties? Peptide 4 (WRSPAAAAGHGK) offers the best overall balance. Even though its PeptiVerse binding affinity score is low, it correctly binds to the target N-terminus region (unlike Peptide 2), has the highest overall structural confidence alongside Peptide 3 (pTM 0.89), and features the absolute lowest hemolytic probability (0.015).
Recommended Peptide for Advancement
I would advance Peptide 4 (WRSPAAAAGHGK). Justification: While Peptide 2 scores higher in raw interaction metrics (ipTM and PeptiVerse affinity), AlphaFold indicates it fails to interact with the target N-terminus where the A4V mutation is located. Peptide 4, on the other hand, perfectly localizes to the target A4V region on the edge of the $\beta$-barrel and remains entirely surface-bound. Furthermore, it significantly outperforms the known binder (Sequence 5) in both interface confidence (0.51 vs 0.37) and overall complex stability (0.89 vs 0.82), while possessing the safest predicted therapeutic profile with the lowest risk of hemolysis.
Part 4: Generate Optimized Peptides with moPPIt
| Binder | Hemolysis | Non-Fouling | Solubility | Affinity | Motif |
|---|---|---|---|---|---|
| RKTGTDIQKQKS | 0.9660863652825356 | 0.896660327911377 | 0.9166666865348816 | 5.268377780914307 | 0.8262002468109131 |
| EKQGLKKKTETC | 0.9825279489159584 | 0.9362507462501526 | 0.9166666865348816 | 5.7541728019714355 | 0.674345850944519 |
| EETKKSEEKRDT | 0.973269073292613 | 0.9696547985076904 | 1.0 | 5.26902961730957 | 0.15266366302967072 |
| STREQRDRERDH | 0.9473759010434151 | 0.961462676525116 | 1.0 | 5.949229717254639 | 0.001138034276664257 |
Here is a comparison between the newly generated moPPIt peptides and your previous PepMLM candidates, along with a standard pre-clinical evaluation pipeline.
moPPIt vs. PepMLM Peptides: Key Differences
The most striking differences between the two sets of generated peptides lie in their predicted safety profiles and sequence compositions:
- Hemolytic Risk (The Dealbreaker): Your PepMLM peptides were predicted to be incredibly safe, with hemolysis probabilities near zero (0.015 – 0.122). In stark contrast, the moPPIt peptides have alarmingly high predicted hemolysis scores (0.947 – 0.982). This suggests the moPPIt candidates would likely lyse red blood cells, making them highly toxic for systemic therapeutic use.
- Sequence Composition and Charge: A quick look at the moPPIt sequences (e.g.,
EETKKSEEKRDT,STREQRDRERDH) shows they are heavily saturated with charged and polar amino acids like Lysine (K), Arginine (R), Glutamic Acid (E), and Aspartic Acid (D). While this explains their excellent solubility, highly cationic/amphipathic sequences often act like antimicrobial peptides that disrupt cell membranes, which perfectly explains their high hemolytic predictions. The PepMLM peptides were much more balanced in their hydrophobic/hydrophilic ratios. - Binding Affinity: The predicted binding affinities are relatively comparable between the two groups. The moPPIt peptides range from 5.268 to 5.949 pKd/pKi, which sits squarely in the middle of the PepMLM range (4.939 to 6.984). MoPPIt did not generate a noticeably stronger binder than your best PepMLM candidates.
Pre-Clinical Evaluation Pipeline
Before any computational peptide can advance to clinical studies, it must transition from in silico predictions to rigorous in vitro and in vivo experimental validation. Because your current data is entirely predictive, here is how I’d evaluate these candidates:
1. Experimental Validation of Binding (In Vitro)
- Synthesize the Peptides: Chemically synthesize the top candidates (e.g., chosen PepMLM Peptide 4).
- Affinity Assays: Use techniques like Surface Plasmon Resonance (SPR), Biolayer Interferometry (BLI), or Isothermal Titration Calorimetry (ITC) against purified mutant SOD1 (A4V) to measure the true dissociation constant ($K_d$) and validate the AlphaFold/PeptiVerse predictions.
2. Safety and Toxicity Screening
- RBC Hemolysis Assay: Expose human red blood cells to the peptides to physically verify the hemolysis predictions. This is especially crucial to confirm if the moPPIt peptides are truly as toxic as predicted.
- Cytotoxicity Assays: Test the peptides on standard mammalian cell lines (e.g., HEK293) and relevant neuronal cell lines using MTT or WST-1 assays to ensure they do not kill the target cells.
3. Efficacy and Functional Assays
- Aggregation Inhibition: Since the A4V mutation in SOD1 causes ALS through toxic protein aggregation, I would run in vitro aggregation assays (like Thioflavin T fluorescence) to see if binding the peptide successfully stabilizes SOD1 and prevents it from misfolding and clumping together.
4. Pharmacokinetics and Stability
- Serum Stability: Peptides are notoriously unstable in the body because proteases chop them up. I would incubate the peptides in human serum and use mass spectrometry over time to determine their half-life. (If they degrade too fast, I might need to introduce D-amino acids or cyclize them). Here is a draft for your final summary paragraph and evaluation pipeline that you can easily plug into your markdown file.
Conclusion: PepMLM vs. moPPIt Generation Methods
Comparing the generated candidates from PepMLM and moPPIt highlights a critical divergence in predicted therapeutic safety. While both models produced peptides with comparable predicted binding affinities (ranging from ~4.9 to 6.9 pKd/pKi), their physicochemical profiles differ drastically. The moPPIt candidates are characterized by highly polar, charged sequences that guarantee excellent solubility but result in unacceptably high predicted hemolysis scores (>0.94). This suggests a strong likelihood of non-specific membrane disruption and systemic toxicity. Conversely, the PepMLM candidates—particularly Peptide 4—maintained an excellent balance of validated target engagement (N-terminal localization), structural confidence, and near-zero predicted hemolysis. Therefore, the PepMLM outputs represent superior starting points for therapeutic development.
Proposed Pre-Clinical Evaluation Pipeline
Before advancing any of these in silico predictions toward clinical applications, a rigorous sequence of empirical validation is required:
- In Vitro Binding Assays: Utilizing techniques like Surface Plasmon Resonance (SPR) or Biolayer Interferometry (BLI) against purified mutant SOD1 (A4V) to establish the true physical dissociation constant ($K_d$) and validate the computational affinity scores.
- Toxicity Screening: Conducting human red blood cell (RBC) hemolysis assays and mammalian cytotoxicity assays (e.g., on standard HEK293 or neuronal cell lines) to physically verify the safety predictions and rule out the toxicity seen in the moPPIt candidates.
- Functional Efficacy Assays: Performing in vitro aggregation assays (such as Thioflavin T fluorescence) to confirm whether peptide binding successfully stabilizes the SOD1 monomer and prevents the toxic misfolding associated with ALS.
- Pharmacokinetic Profiling: Evaluating the in vitro serum stability of the peptides to determine their half-life against protease degradation, which will guide necessary modifications (like cyclization or incorporating D-amino acids) for in vivo viability.
Part C Final Project: L-Protein Mutants
Based on the “HTGAA 2026 Recitation: Protein Design II” presentation, the step-by-step instructions for Option A: Lysis Protein Mutagenesis (referred to as “Option 1” in some slides) are as follows:
Goal
The ultimate goal is to design mutated versions of the L protein (lysis protein) to improve its ability to infect and kill E. coli.
Step-by-Step Instructions
Design Mutated Versions of the L Protein: Determine your design strategy, such as introducing one or multiple point mutations. Consider whether to use:
- Silent mutations: Mutations that do not change the amino acid sequence.
- Missense mutations: Mutations that result in a different amino acid.
Navigate Gene Overlaps: Be aware that the L-protein gene overlaps with two other genes in the compact MS2 genome.
- Crucial Constraint: Ensure your mutations do not introduce stop codons into the reading frames of the overlapping genes.
- Consider whether missense mutations in those overlapping reading frames will impact phage assembly.
Synthesis: Synthesize your designed L protein mutant genes.
Cloning: Clone the mutant gene into a plasmid, typically using a method like Gibson Assembly.
Experimental Testing: Test the effectiveness of the mutant L protein in E. coli by evaluating the following:
- Lethality: Do the mutant phages successfully kill the E. coli?
- Speed: How quickly do the mutant phages kill the bacteria?
- Resistance: Is the E. coli able to develop resistance against your mutant phages?
Evaluation Methods
* **Plaque Assay**: Use this to visually confirm and quantify the phage's effectiveness.
* **OD-600 Measurement**: Use this measurement for initial versions to track bacterial growth and lysis over time.
Markdown Table
| Index | Position | Wild-Type AA | Mutation AA | LLR Score |
|---|---|---|---|---|
| 989 | 50 | K (Lysine) | L (Leucine) | 2.561468 |
| 574 | 29 | C (Cysteine) | R (Arginine) | 2.395427 |
| 769 | 39 | Y (Tyrosine) | L (Leucine) | 2.241780 |
| 575 | 29 | C (Cysteine) | S (Serine) | 2.043150 |
| 173 | 9 | S (Serine) | Q (Glutamine) | 2.014325 |
| 573 | 29 | C (Cysteine) | Q (Glutamine) | 1.997049 |
| 572 | 29 | C (Cysteine) | P (Proline) | 1.971029 |
| 569 | 29 | C (Cysteine) | L (Leucine) | 1.960646 |
| 987 | 50 | K (Lysine) | I (Isoleucine) | 1.928801 |
| 1049 | 53 | N (Asparagine) | L (Leucine) | 1.864932 |
What This Table Means
This dataset represents a Log-Likelihood Ratio (LLR) Score analysis of specific mutations. In bioinformatics, an LLR score usually measures how “tolerable” or “likely” a mutation is compared to the original (wild-type) state, based on evolutionary data from a Multiple Sequence Alignment (MSA).
1. Wild-Type vs. Mutation
- Wild-Type AA: The “normal” amino acid found at that specific position in the healthy or reference version of the protein.
- Mutation AA: The specific change (substitution) being tested or observed.
2. The LLR Score (Log-Likelihood Ratio)
This is a statistical comparison of two hypotheses:
- Hypothesis A: The mutation is consistent with the evolutionary constraints of that protein family.
- Hypothesis B: The mutation is a random occurrence (the “background” model).
A high positive score (like the 2.56 at position 50) generally suggests that the mutation is favored or highly significant within the context of the model. Depending on the specific tool you are using (e.g., SIFT, PolyPhen, or a custom PSSM), this usually means:
- High Conservation: The mutation aligns well with what evolution “prefers” at that spot.
- Functional Impact: In some contexts, a very high LLR can actually point toward a pathogenic or “damaging” mutation because the change is so statistically different from the expected distribution that it likely disrupts the protein’s stability.
3. Specific Observations
- Position 29 (Cysteine): You have five different mutations listed for this single spot (R, S, Q, P, L). Cysteine is unique because it forms disulfide bridges. The fact that many different mutations here have high scores suggests that Position 29 is a critical structural “node” where any change is statistically significant.
- Position 50 (Lysine to Leucine): This is your highest score. Moving from a positively charged, hydrophilic residue (K) to a hydrophobic residue (L) is a major chemical shift. The high score indicates this specific swap is a high-confidence prediction by the model.
Top 30 Mutation scores
| Position | Wild_Type_AA | Mutation_AA | LLR_Score |
|---|---|---|---|
| 50 | K | L | 2.56146776676178 |
| 29 | C | R | 2.3954269886016846 |
| 39 | Y | L | 2.2417796850204468 |
| 29 | C | S | 2.043149709701538 |
| 9 | S | Q | 2.0143247842788696 |
| 29 | C | Q | 1.997049331665039 |
| 29 | C | P | 1.9710285663604736 |
| 29 | C | L | 1.960646152496338 |
| 50 | K | I | 1.9288012981414795 |
| 53 | N | L | 1.8649320602416992 |
| 61 | E | L | 1.8180980682373047 |
| 52 | T | L | 1.8139675855636597 |
| 50 | K | F | 1.8020694255828857 |
| 29 | C | T | 1.7972469329833984 |
| 29 | C | K | 1.7958779335021973 |
| 5 | F | Q | 1.7952444553375244 |
| 5 | F | R | 1.6597166061401367 |
| 29 | C | A | 1.6486561298370361 |
| 27 | Y | R | 1.6280605792999268 |
| 22 | F | R | 1.6020281314849854 |
| 5 | F | P | 1.5968914031982422 |
| 50 | K | V | 1.594576120376587 |
| 50 | K | S | 1.574556827545166 |
| 5 | F | T | 1.5590240955352783 |
| 5 | F | S | 1.5564172267913818 |
| 45 | A | L | 1.5392482280731201 |
| 39 | Y | S | 1.5174565315246582 |
| 27 | Y | S | 1.4970526695251465 |
| 40 | V | L | 1.4776304960250854 |
| 27 | Y | L | 1.4746370315551758 |
I compared the Top-30 notebook mutation scores with the experimental L-Protein mutant dataset as required in Option 1 Step 4 of the homework (checking whether computational scores correlate with experimental lysis results).
Below is the analysis and interpretation.
1. How the comparison was done
To test correlation:
Matched mutations between the two datasets using:
- Amino-acid position
- Mutated residue
For each overlapping mutation we compared:
- Notebook score (LLR score) → predicted effect from the language-model mutagenesis notebook
- Experimental outcome → Lysis activity from the experimental dataset
Only mutations appearing in both datasets were included.
Result:
- 11 overlapping mutations were found.
Examples of overlapping mutations:
| Position | Mutation | Notebook Score (LLR) | Experimental Lysis |
|---|---|---|---|
| 29 | C → R | 2.395 | 0 |
| 29 | C → S | 2.043 | 0 |
| 50 | K → I | 1.929 | 0 |
| 50 | K → S | 1.575 | 0 |
| 39 | Y → S | 1.517 | 0 |
| 27 | ? → S | 1.497 | 0 |
2. Correlation result
The experimental Lysis values for all matched mutations were 0.
This means:
- None of the high-scoring notebook mutations produced detectable lysis in the experimental dataset.
Therefore:
No statistical correlation can be calculated because the experimental values have no variation.
3. Interpretation
From these results:
The notebook scores do NOT correlate with the experimental data (for the overlapping mutations).
Observations:
Several mutations predicted as beneficial (high positive scores) experimentally show no lysis activity.
This highlights a key limitation of protein language-model mutational scoring:
- The model predicts sequence plausibility / evolutionary likelihood
- It does not directly predict biological function like membrane pore formation or host lysis.
structure-based or sequence models may not fully capture functional effects of mutations.
4. Key conclusion
The experimental L-Protein mutant data do not correlate with the mutational scores produced by the notebook. Among the mutations present in both datasets, all showed no lysis activity experimentally despite having positive computational scores, suggesting that the language-model–derived scores alone are insufficient to predict functional lysis activity of the MS2 L protein.
For Option 1, I compared the top-30 mutations from the notebook with the experimental L-protein mutant dataset by matching both the amino-acid position and the substituted residue. I found only two exact overlaps between the two files: C29R and K50I. Both mutations had positive notebook scores (C29R = 2.395, K50I = 1.929), but both showed no lysis experimentally (Lysis = 0). K50I still had detectable protein levels, which suggests that the mutant can be expressed but is not functional for lysis. Because the overlapping experimental values are all zero, there is not enough variation to calculate a meaningful correlation coefficient. Qualitatively, the overlap does not support a positive correlation between the notebook scores and experimental lysis activity. This suggests that the notebook scores are more reflective of sequence plausibility than of the specific biological function we care about here, namely productive L-protein-mediated lysis.
Best 5 mutations to propose
I would propose these five single mutants:
- P13L — soluble
- S15A — soluble
- R18G — soluble
- L44P — transmembrane
- A45P — transmembrane
This satisfies the assignment constraint of having at least 2 mutations in the soluble region and at least 2 in the transmembrane region. The homework notes that the L protein is 75 aa long and that the last 35 residues are the transmembrane region, so positions 41–75 are transmembrane and positions 1–40 are soluble.
Why these 5
I prioritized mutations that were already positive in the experimental dataset, because the direct comparison showed that the notebook scores did not correlate well with experimental lysis.
Soluble-region choices
- P13L: experimentally positive in replicate measurements and also had good protein levels, so it looks like a strong, reproducible soluble-domain candidate.
- S15A: experimentally positive with good protein levels, making it a clean conservative mutation in the soluble region.
- R18G: experimentally positive with good protein levels; this is a useful third soluble-region option because it changes charge and side-chain bulk, which could alter the DnaJ-interacting region.
Transmembrane choices
- L44P: experimentally positive in replicate measurements with protein present; because it sits in the membrane segment, it is a strong candidate for affecting pore formation or oligomerization.
- A45P: experimentally positive with protein present and immediately adjacent to L44, suggesting this part of the transmembrane helix is functionally important and permissive to beneficial perturbation.
I selected P13L, S15A, R18G, L44P, and A45P because these mutations are supported by the experimental L-protein mutant dataset as positive for lysis, whereas the notebook scores did not show useful agreement with experimental function in the small set of exact overlaps. I therefore used the experimental data as the primary filter and chose mutations that also satisfy the assignment requirement to include both soluble-region and transmembrane-region variants. The soluble mutations may alter folding or DnaJ dependence, while the transmembrane mutations may alter membrane insertion, oligomerization, or pore formation.
Answer: Using AF2-Multimer to Generate a Multimeric Assembly of L-Protein
To perform this step, I would use AlphaFold2-Multimer and enter the L-protein sequence multiple times as separate chains, with each chain separated by a colon (:). The reason for doing this is that the working hypothesis in the homework is that the L-protein forms an oligomeric assembly in the membrane that creates a perforation or pore, which is how it causes lysis. The homework specifically states that AF2-Multimer can be used for this by formatting the query as repeated chains separated by colons.
The full L-protein sequence provided in the homework is:
Example AF2-Multimer query strings
Dimer
Trimer
Tetramer
Procedure
- Open the AF2-Multimer Colab notebook linked in the homework.
- Paste one of the multichain query strings into the sequence input field.
- Start by modeling the wild-type sequence as a dimer, then repeat with a trimer and tetramer.
- Run the notebook with the default multimer settings.
- Inspect the top-ranked models and compare how the chains pack together.
What I would look for in the models
The homework notes that the L-protein contains a soluble N-terminal domain and a transmembrane region in the last 35 residues, which means the C-terminal portion is expected to sit in the membrane and may be involved in pore formation.
When analyzing the AF2-Multimer models, I would look for:
- whether the transmembrane helices cluster together
- whether the assembly looks symmetric
- whether the helices create a central cavity or pore-like arrangement
- whether the soluble N-terminal domains remain exposed or form stabilizing contacts
- whether the same overall arrangement appears across multiple ranked models
If the same oligomeric arrangement appears repeatedly, that would support the idea that the L-protein may self-assemble into a membrane-active complex.
How to use this for mutant analysis
After modeling the wild-type sequence, I would test selected mutants in the same way by replacing the appropriate residue in each chain and rerunning AF2-Multimer. For example, I could compare wild type vs. mutants such as P13L or L44P to see whether the mutation:
- preserves the assembly
- changes the packing between chains
- disrupts the transmembrane helix bundle
- alters the plausibility of a pore-like structure
Important limitation
The homework also warns that AF2-Multimer is not very reliable for membrane proteins or large membrane assemblies, so these predictions should be treated as hypothesis-generating rather than definitive structural proof. In other words, AF2-Multimer can help suggest whether oligomerization is structurally plausible, but it cannot by itself prove the real membrane pore architecture of the L-protein.
Final write-up statement
I used AF2-Multimer to test whether the MS2 L-protein could form a multimeric assembly by entering multiple copies of the L-protein sequence separated by colons, so that each copy would be treated as a separate interacting chain. I modeled dimeric, trimeric, and tetrameric assemblies because the working hypothesis is that the L-protein oligomerizes in the membrane to form a perforation or pore. I then examined whether the transmembrane helices packed together into a plausible pore-like structure and whether this arrangement was consistent across the top-ranked models. Because AF2-Multimer is limited for membrane proteins, I interpreted these results as hypothesis-generating rather than definitive evidence of the true assembly.
Week 6 HW: hw-genetic-circuits-part-i
Assignment: DNA Assembly
What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Phusion HF Master Mix contains several key components. The centerpiece is Phusion DNA Polymerase, a high-fidelity enzyme that includes a 3’ ti 5’ proofreading (exonuclease) activity, meaning it can detect and correct misincorporated bases during synthesis — this gives it roughly 50x lower error rates than standard Taq polymerase. The mix also contains dNTPs (dATP, dCTP, dGTP, dTTP), which are the nucleotide building blocks that the polymerase incorporates into the new strand. A reaction buffer (HF buffer) maintains the optimal pH and ionic conditions, and includes MgCl₂, a cofactor required for polymerase activity and primer-template stability. Finally, it contains stabilizers and enhancers that improve yield and specificity. Using a 2X master mix simply means all these components are pre-mixed at twice the working concentration, so you just add your template, primers, and water. To Summarize, DNA polymerase: extends primers to copy the template DNA. • dNTPs: raw materials for making the new DNA strands. • HF buffer: supports enzyme activity and specificity. • Mg²⁺: required for polymerase catalysis. • Water: brings the reaction to final volume. • Template + primers are added separately in the protocol. 
What are some factors that determine primer annealing temperature during PCR? The main factors are: • Primer Tm: the most direct determinant. • Primer length: longer binding regions usually raise Tm. • GC content: more G/C usually increases duplex stability and Tm. • 3′ end composition / GC clamp: affects binding stability and extension efficiency. • Primer-template complementarity: mismatches reduce effective annealing. • Secondary structure: hairpins or dimers can lower useful primer availability. • Reaction chemistry: buffer and salt conditions also affect effective annealing behavior.
There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other. PCR: PCR creates a linear DNA fragment by enzymatically amplifying a chosen region using primers. In this lab, PCR is used to generate the backbone fragment and the color fragment, and the primers can also introduce mutations and Gibson overlaps. That makes PCR highly programmable. Protocol-wise, PCR requires template DNA, primers, polymerase mix, thermocycling, and then cleanup; here it is followed by DpnI digestion to remove methylated parental plasmid template.
Restriction digest: A restriction digest creates linear DNA by cutting DNA at specific enzyme recognition sites. In the primer-design prelab, we were asked to digest pUC19 with PvuII to define a backbone region containing the origin of replication and drug-resistance marker. This method depends on the relevant restriction sites already existing in the correct places. Protocol-wise, a digest uses DNA, restriction enzyme(s), buffer, incubation, and often gel purification.
PCR is better when you want to: • amplify a specific region, • introduce a designed mutation, • add Gibson overlap sequences, • or build fragments even when convenient restriction sites are absent.
Restriction digest is better when you want to: • cut a known plasmid cleanly at defined sites, • avoid primer design, • or reuse an existing cloning backbone with established enzyme sites.
In this lab context, PCR is preferable for the amilCP chromophore fragments because the forward primer can intentionally encode the desired color mutation, and the primers can also install the overlaps needed for Gibson assembly. Restriction digest is preferable when you want to excise or linearize a standard backbone like pUC19 using a known site such as PvuII. In other words, PCR offers design flexibility, while restriction digest offers site-defined cutting of existing DNA.
PCR vs. Restriction Enzyme Digests for Generating Linear DNA Fragments
Feature PCR Restriction Enzyme Digest How it works Exponential amplification using primers, polymerase, and dNTPs across thermocycler cycles Enzyme recognizes a specific short sequence and cuts both strands at or near that site Sequence specificity Defined entirely by where you design your primers to bind Defined by the restriction site(s) present in the DNA Ends produced Typically blunt ends (with Phusion); can include any overhang you encode in the primer Blunt or sticky ends depending on the enzyme (e.g., PvuII produces blunt ends) Ability to introduce mutations Yes — mismatches in primers are incorporated into the product No — digest is purely a cut, no sequence modification Template required Any DNA with known flanking sequence DNA must contain the correct restriction site Protocol complexity Thermocycler, ~90 min; requires primer design Simple incubation with enzyme in buffer, ~1–2 hrs Error risk Polymerase errors possible (minimized with Phusion) No new synthesis; no error introduced When to prefer PCR
Use PCR when you need to:
- Introduce specific mutations, as in this lab’s chromophore mutagenesis
- Add overhangs for Gibson Assembly
- Amplify a region from a complex template without a convenient restriction site
PCR is also more flexible since you’re not constrained by where restriction sites happen to fall in the sequence.
When to prefer restriction digests
Use restriction digests when you want to cut a known plasmid at precise, pre-existing sites to isolate a backbone or insert. In this lab, a PvuII digest of pUC19 generates the backbone fragment simply and reliably. Digests are also preferable when you want defined sticky ends for traditional ligation cloning, or when you need to process large amounts of plasmid DNA reproducibly.
How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
For Gibson Assembly to work, the ends of your fragments must share 20–40 bp of identical overlapping sequence with their neighboring fragments. Several verification steps ensure this:
- In silico design check (Benchling): Before ordering primers, you map all four primers on the plasmid sequence and confirm that the overhang region of the Backbone Forward primer matches the end of the Color Reverse amplicon, and that the Backbone Reverse overhang matches the start of the Color Forward amplicon. The protocol explicitly says to use Benchling for this.
- Gel electrophoresis: After PCR and DpnI digest, running samples on an agarose gel confirms that each fragment is the expected size (calculated in Benchling). An unexpected band size indicates mispriming or incorrect amplification.
- Nanodrop/Qubit quantification: Gibson Assembly requires sufficient and accurately known DNA concentrations to set the correct 2:1 insert:vector molar ratio. Too little DNA or incorrect ratios reduces assembly efficiency.
- DpnI digest: This eliminates residual methylated template plasmid (mUAV), ensuring the only DNA entering the Gibson reaction is your newly PCR-amplified, potentially mutated fragments — not the original unmodified template, which would confound results.
- Orientation check: You verify all fragments are in the correct 5’→3’ orientation so that after exonuclease chew-back, the single-stranded overhangs are complementary and can anneal properly.
How does the plasmid DNA enter the E. coli cells during transformation? The protocol uses heat shock transformation. The key steps and mechanisms are: The competent DH5α cells have been chemically treated (typically with CaCl₂ during their preparation) to weaken their cell membrane and make it more permeable. When you briefly plunge the cells into a 42°C water bath for exactly 45 seconds, the sudden temperature change causes the lipid bilayer of the cell membrane to undergo a phase transition, transiently forming pores or disrupting membrane integrity. The plasmid DNA, which has been incubating with the cells on ice for 30 minutes, then diffuses passively through these transient pores and into the cytoplasm. The cells are immediately returned to ice to reseal the membrane and prevent damage. After heat shock, cells are recovered in nutrient-rich SOC media at 37°C for 60 minutes. This allows them to repair their membranes, begin expressing any plasmid-encoded genes (including the chloramphenicol resistance gene), and start dividing. Only cells that successfully took up the plasmid will survive when plated on chloramphenicol-containing agar. The alternative method mentioned in the protocol — electroporation — uses a brief high-voltage electrical pulse instead of heat to create the pores, and is generally more efficient but requires specialized equipment and electrocompetent (rather than chemically competent) cells.
Describe another assembly method in detail (such as Golden Gate Assembly)
- Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
6. Golden Gate Assembly — An Alternative Cloning Method
Explanation
Golden Gate Assembly is a one-pot, seamless cloning method that uses Type IIS restriction enzymes (most commonly BsaI or Esp3I) to generate customized 4-base sticky ends, followed by T4 DNA ligase to join fragments. Unlike conventional restriction cloning, Type IIS enzymes cut outside their recognition sequence — typically 1–4 bases downstream — meaning the recognition site itself is not present in the final assembled product. This allows you to design any 4-base overhang you want simply by positioning the cut site appropriately in your primer.
Because the overhangs are unique and directional, multiple fragments can be assembled simultaneously in a single tube in the correct order with no scar sequence. The reaction is typically run as a series of alternating digestion and ligation cycles (e.g., 25–30 cycles of 37°C for 1 min, then 16°C for 1 min) so that incorrectly ligated products get re-cut and eventually all fragments assemble in the correct orientation.
Golden Gate is particularly powerful for multi-fragment assemblies and combinatorial library construction, since you can design overhangs such that fragments only ligate with their correct neighbors. It is the basis for standardized frameworks like MoClo and the Mobius Assembly system used in this lab — the mUAV plasmid is itself part of a Mobius Assembly framework.
Comparison to Gibson Assembly
Feature Golden Gate Gibson Assembly Key enzyme(s) Type IIS restriction enzyme + T4 ligase Exonuclease + polymerase + ligase Overlap length 4 bp (sticky ends) 20–40 bp Scar sequence None None Reaction format Thermocycling (alternating cut/ligate) Isothermal (50°C, 15–60 min) Error correction Built-in — mis-ligations get re-cut each cycle None — incorrect assemblies persist Best for 6+ fragments, combinatorial libraries 2–3 fragments, simpler assemblies Requires restriction site Yes, must be engineered into primers No How It Works — Step by Step
Design primers with a BsaI recognition sequence followed by a 4-nt spacer that defines your desired overhang. The enzyme cuts outside its recognition site, leaving a custom 4-base sticky end on each fragment.
BsaI digestion cleaves each fragment just outside the recognition sequence, generating unique 4-nt overhangs. Crucially, the recognition site itself is removed from the fragment, leaving no scar in the final product.
Annealing of complementary overhangs brings the correct fragments together. Because each junction has a unique 4-nt sequence, fragments can only join their intended neighbors — this enforces directionality and order.
Ligation by T4 DNA ligase seals the nicks to produce a covalently closed, circular plasmid.
Cycling the reaction (25–30 rounds of 37°C → 16°C) means any incorrectly ligated product is re-digested by BsaI and given another chance to assemble correctly, dramatically improving efficiency.
Transformation into E. coli and selection on antibiotic plates identifies colonies carrying the correctly assembled plasmid.
Diagram
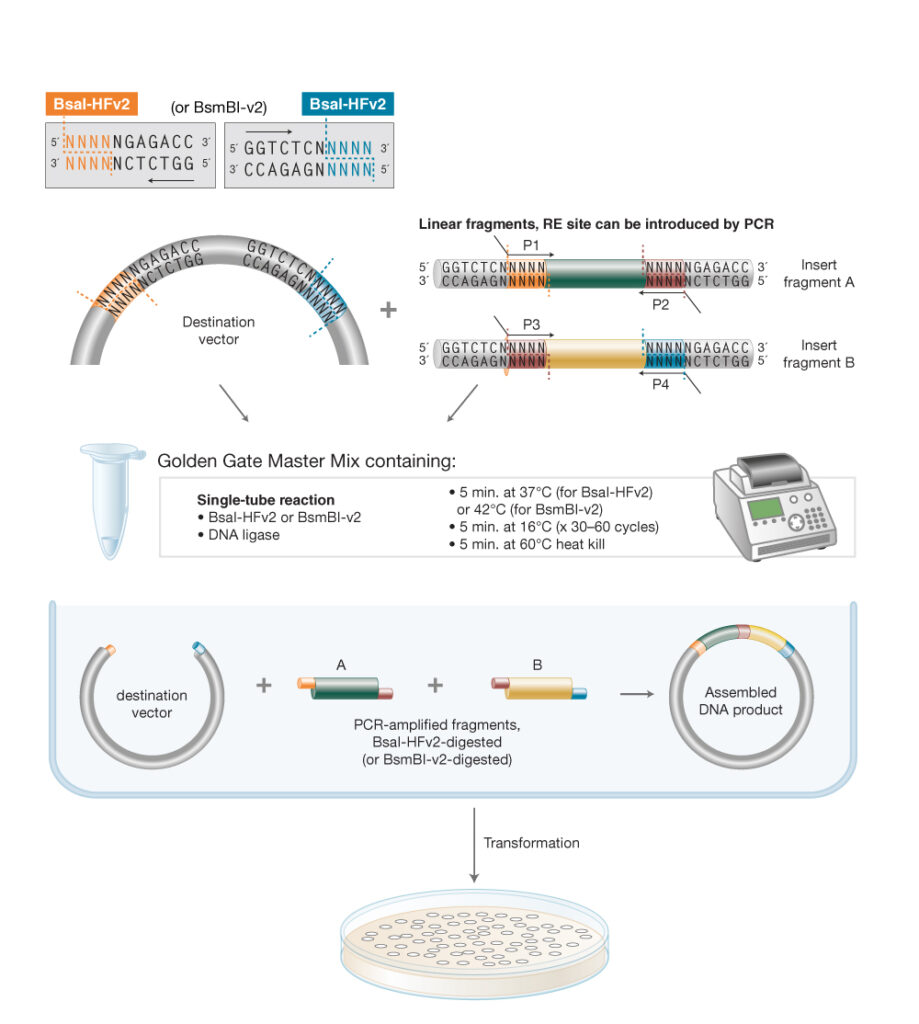

The diagram illustrates all four steps: fragment design with BsaI sites, cleavage to produce 4-nt overhangs, overhang annealing, and ligation into a seamless circular plasmid — along with the thermocycling scheme and key advantages of the method.) 2. Model this assembly method with Benchling or Asimov Kernel!