Week 2 HW: DNA Read, Write, and Edit
Part 3: DNA Design Challenge

3.1 Protein of interest:
I’m interested in exploring JAK1 protein that is implicated in autoimmune diseases. I chose this by finding which are the priority protein targets for treatment of conditions like Vitiligo. Link to the protein from Uniprot: https://rest.uniprot.org/uniprotkb/P23458.fasta
Protein Sequence:
>sp|P23458|JAK1_HUMAN Tyrosine-protein kinase JAK1 OS=Homo sapiens OX=9606 GN=JAK1 PE=1 SV=2
MQYLNIKEDCNAMAFCAKMRSSKKTEVNLEAPEPGVEVIFYLSDREPLRLGSGEYTAEEL
CIRAAQACRISPLCHNLFALYDENTKLWYAPNRTITVDDKMSLRLHYRMRFYFTNWHGTN
DNEQSVWRHSPKKQKNGYEKKKIPDATPLLDASSLEYLFAQGQYDLVKCLAPIRDPKTEQ
DGHDIENECLGMAVLAISHYAMMKKMQLPELPKDISYKRYIPETLNKSIRQRNLLTRMRI
NNVFKDFLKEFNNKTICDSSVSTHDLKVKYLATLETLTKHYGAEIFETSMLLISSENEMN
WFHSNDGGNVLYYEVMVTGNLGIQWRHKPNVVSVEKEKNKLKRKKLENKHKKDEEKNKIR
EEWNNFSYFPEITHIVIKESVVSINKQDNKKMELKLSSHEEALSFVSLVDGYFRLTADAH
HYLCTDVAPPLIVHNIQNGCHGPICTEYAINKLRQEGSEEGMYVLRWSCTDFDNILMTVT
CFEKSEQVQGAQKQFKNFQIEVQKGRYSLHGSDRSFPSLGDLMSHLKKQILRTDNISFML
KRCCQPKPREISNLLVATKKAQEWQPVYPMSQLSFDRILKKDLVQGEHLGRGTRTHIYSG
TLMDYKDDEGTSEEKKIKVILKVLDPSHRDISLAFFEAASMMRQVSHKHIVYLYGVCVRD
VENIMVEEFVEGGPLDLFMHRKSDVLTTPWKFKVAKQLASALSYLEDKDLVHGNVCTKNL
LLAREGIDSECGPFIKLSDPGIPITVLSRQECIERIPWIAPECVEDSKNLSVAADKWSFG
TTLWEICYNGEIPLKDKTLIEKERFYESRCRPVTPSCKELADLMTRCMNYDPNQRPFFRA
IMRDINKLEEQNPDIVSEKKPATEVDPTHFEKRFLKRIRDLGEGHFGKVELCRYDPEGDN
TGEQVAVKSLKPESGGNHIADLKKEIEILRNLYHENIVKYKGICTEDGGNGIKLIMEFLP
SGSLKEYLPKNKNKINLKQQLKYAVQICKGMDYLGSRQYVHRDLAARNVLVESEHQVKIG
DFGLTKAIETDKEYYTVKDDRDSPVFWYAPECLMQSKFYIASDVWSFGVTLHELLTYCDS
DSSPMALFLKMIGPTHGQMTVTRLVNTLKEGKRLPCPPNCPDEVYQLMRKCWEFQPSNRT
SFQNLIEGFEALLK
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
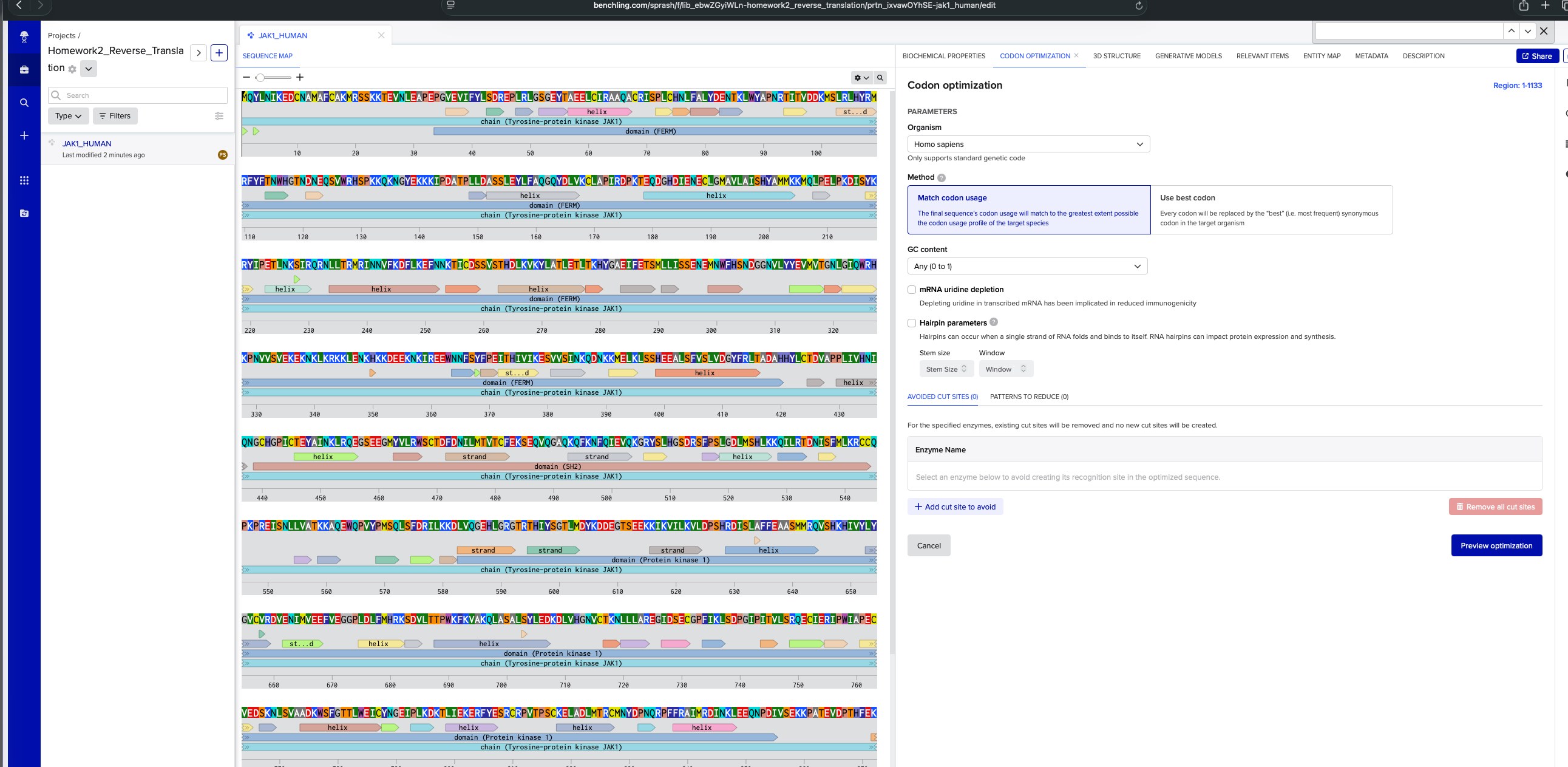

DNA Sequence for homosapien using codon optimization tool in benchling:
ATGCAGTACCTGAACATCAAGGAGGACTGCAACGCCATGGCCTTCTGCGCCAAGATGAGAAGCAGCAAGAAGACCGAGGTGAACCTGGAGGCCCCCGAGCCCGGCGTGGAGGTGATCTTCTACCTGAGCGACAGAGAGCCCCTGAGGCTGGGCAGCGGCGAGTACACCGCCGAGGAGCTGTGCATCAGAGCCGCCCAGGCCTGCAGGATCAGCCCCCTGTGCCACAACCTGTTCGCCCTGTACGACGAGAACACCAAGCTGTGGTACGCCCCCAACAGGACCATCACCGTGGACGACAAGATGAGCCTGAGGCTGCACTACAGGATGAGGTTCTACTTCACCAACTGGCACGGCACCAACGACAACGAGCAGAGCGTGTGGAGGCACAGCCCCAAGAAGCAGAAGAACGGCTACGAGAAGAAGAAGATCCCCGACGCCACCCCCCTGCTGGACGCCAGCAGCCTGGAGTACCTGTTCGCCCAGGGCCAGTACGACCTGGTGAAGTGCCTGGCCCCCATCAGGGACCCCAAGACCGAGCAGGACGGCCACGACATCGAGAACGAGTGCCTGGGCATGGCCGTGCTGGCCATCAGCCACTACGCCATGATGAAGAAGATGCAGCTGCCCGAGCTGCCCAAGGACATCAGCTACAAGAGGTACATCCCCGAGACCCTGAACAAGAGCATCAGGCAGAGGAACCTGCTGACCAGAATGAGGATCAACAACGTGTTCAAGGACTTCCTGAAGGAGTTCAACAACAAGACCATCTGCGACAGCAGCGTGAGCACCCACGACCTGAAGGTGAAGTACCTGGCCACCCTGGAGACCCTGACCAAGCACTACGGCGCCGAGATCTTCGAGACCAGCATGCTGCTGATCAGCAGCGAGAACGAGATGAACTGGTTCCACAGCAACGACGGCGGCAACGTGCTGTACTACGAGGTGATGGTGACCGGCAACCTGGGCATCCAGTGGAGGCACAAGCCCAACGTGGTGAGCGTGGAGAAGGAGAAGAACAAGCTGAAGAGGAAGAAGCTGGAGAACAAGCACAAGAAGGACGAGGAGAAGAACAAGATCAGGGAGGAGTGGAACAACTTCAGCTACTTCCCCGAGATCACCCACATCGTGATCAAGGAGAGCGTGGTGAGCATCAACAAGCAGGACAACAAGAAGATGGAGCTGAAGCTGAGCAGCCACGAGGAGGCCCTGAGCTTCGTGAGCCTGGTGGACGGCTACTTCAGGCTGACCGCCGACGCCCACCACTACCTGTGCACCGACGTGGCCCCCCCCCTGATCGTGCACAACATCCAGAACGGCTGCCACGGCCCCATCTGCACCGAGTACGCCATCAACAAGCTGAGACAGGAGGGCAGCGAGGAGGGCATGTACGTGCTGAGGTGGAGCTGCACCGACTTCGACAACATCCTGATGACCGTGACCTGCTTCGAGAAGAGCGAGCAGGTGCAGGGCGCCCAGAAGCAGTTCAAGAACTTCCAGATCGAGGTGCAGAAGGGCAGGTACAGCCTGCACGGCAGCGACAGGAGCTTCCCCAGCCTGGGCGACCTGATGAGCCACCTGAAGAAGCAGATCCTGAGGACCGACAACATCAGCTTCATGCTGAAGAGGTGCTGCCAGCCCAAGCCCAGAGAGATCAGCAACCTGCTGGTGGCCACCAAGAAGGCCCAGGAGTGGCAGCCCGTGTACCCCATGAGCCAGCTGAGCTTCGACAGGATCCTGAAGAAGGACCTGGTGCAGGGCGAGCACCTGGGCAGAGGCACCAGGACCCACATCTACAGCGGCACCCTGATGGACTACAAGGACGACGAGGGCACCAGCGAGGAGAAGAAGATCAAGGTGATCCTGAAGGTGCTGGACCCCAGCCACAGGGACATCAGCCTGGCCTTCTTCGAGGCCGCCAGCATGATGAGGCAGGTGAGCCACAAGCACATCGTGTACCTGTACGGCGTGTGCGTGAGGGACGTGGAGAACATCATGGTGGAGGAGTTCGTGGAGGGCGGCCCCCTGGACCTGTTCATGCACAGGAAGAGCGACGTGCTGACCACCCCCTGGAAGTTCAAGGTGGCCAAGCAGCTGGCCAGCGCCCTGAGCTACCTGGAGGACAAGGACCTGGTGCACGGCAACGTGTGCACCAAGAACCTGCTGCTGGCCAGGGAGGGCATCGACAGCGAGTGCGGCCCCTTCATCAAGCTGAGCGACCCCGGCATCCCCATCACCGTGCTGAGCAGGCAGGAGTGCATCGAGAGGATCCCCTGGATCGCCCCCGAGTGCGTGGAGGACAGCAAGAACCTGAGCGTGGCCGCCGACAAGTGGAGCTTCGGCACCACCCTGTGGGAGATCTGCTACAACGGCGAGATCCCCCTGAAGGACAAGACCCTGATCGAGAAGGAGAGGTTCTACGAGAGCAGATGCAGGCCCGTGACCCCCAGCTGCAAGGAGCTGGCCGACCTGATGACCAGGTGCATGAACTACGACCCCAACCAGAGGCCCTTCTTCAGAGCCATCATGAGGGACATCAACAAGCTGGAGGAGCAGAACCCCGACATCGTGAGCGAGAAGAAGCCCGCCACCGAGGTGGACCCCACCCACTTCGAGAAGAGGTTCCTGAAGAGGATCAGGGACCTGGGCGAGGGCCACTTCGGCAAGGTGGAGCTGTGCAGATACGACCCCGAGGGCGACAACACCGGCGAGCAGGTGGCCGTGAAGAGCCTGAAGCCCGAGAGCGGCGGCAACCACATCGCCGACCTGAAGAAGGAGATCGAGATCCTGAGGAACCTGTACCACGAGAACATCGTGAAGTACAAGGGCATCTGCACCGAGGACGGCGGCAACGGCATCAAGCTGATCATGGAGTTCCTGCCCAGCGGCAGCCTGAAGGAGTACCTGCCCAAGAACAAGAACAAGATCAACCTGAAGCAGCAGCTGAAGTACGCCGTGCAGATCTGCAAGGGCATGGACTACCTGGGCAGCAGGCAGTACGTGCACAGGGACCTGGCCGCCAGGAACGTGCTGGTGGAGAGCGAGCACCAGGTGAAGATCGGCGACTTCGGCCTGACCAAGGCCATCGAGACCGACAAGGAGTACTACACCGTGAAGGACGACAGGGACAGCCCCGTGTTCTGGTACGCCCCCGAGTGCCTGATGCAGAGCAAGTTCTACATCGCCAGCGACGTGTGGAGCTTCGGCGTGACCCTGCACGAGCTGCTGACCTACTGCGACAGCGACAGCAGCCCCATGGCCCTGTTCCTGAAGATGATCGGCCCCACCCACGGCCAGATGACCGTGACCAGACTGGTGAACACCCTGAAGGAGGGCAAGAGGCTGCCCTGCCCCCCCAACTGCCCCGACGAGGTGTACCAGCTGATGAGGAAGTGCTGGGAG
3.4. You have a sequence! Now what? What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
At first, Transcribe the DNA sequence to mRNA by inserting the DNA into living cells. The DNA template strand is read 3’ to 5’ and mRNA is synthesized 5’ to 3’. The resulting mRNA contains 5’ UTR, start codon (AUG), coding sequence, stop codon, and 3’ UTR.
Next is Translation. The ribosome binds the mRNA, it recognizes the start codon (AUG), reads the codons sequentially, tRNA delivers corresponding amino acids. Peptide bonds are formed and it stops at stop codon (UAA, UAG, UGA). The polypeptide then folds, may undergo post-translational modification and could get either secreted or remain intracellular.
3.5. [Optional] How does it work in nature/biological systems? Describe how a single gene codes for multiple proteins at the transcriptional level.
Single genes code for multiple proteins through the process of alternate splicing where different combination of exons are expressed.
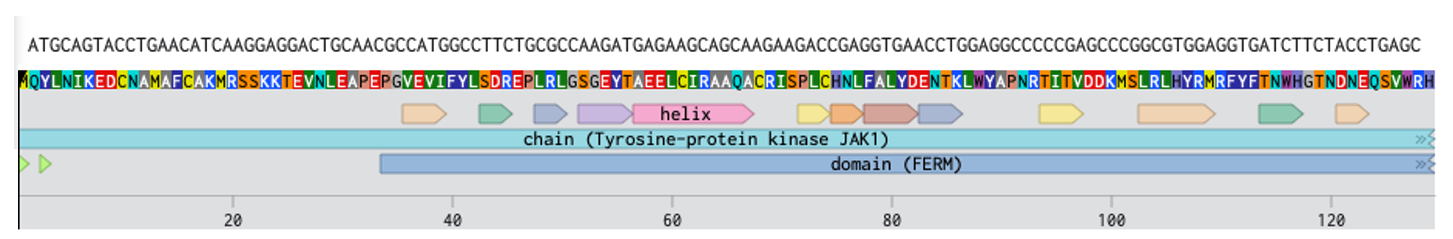
Superimposed DNA -> AA
Part 4: Prepare a Twist DNA Synthesis Order

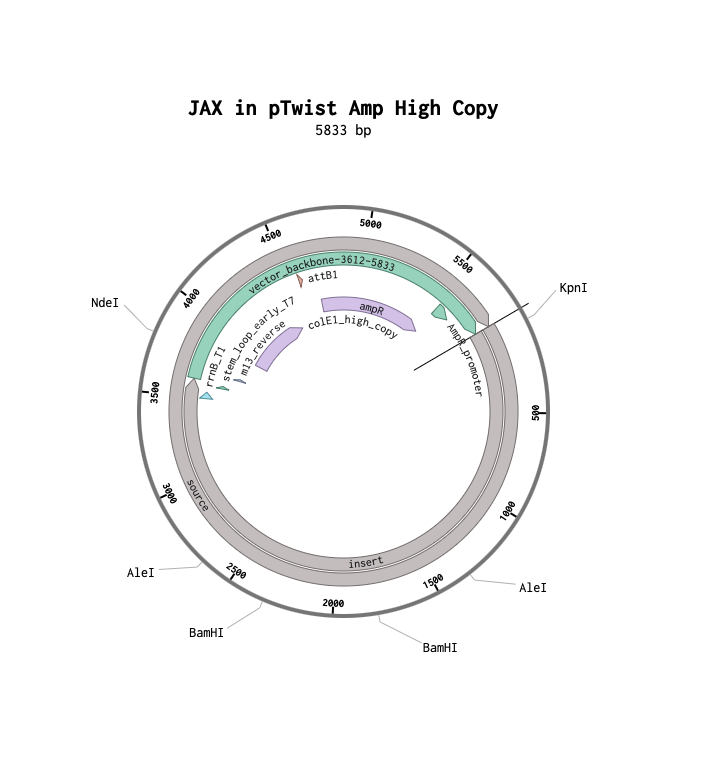
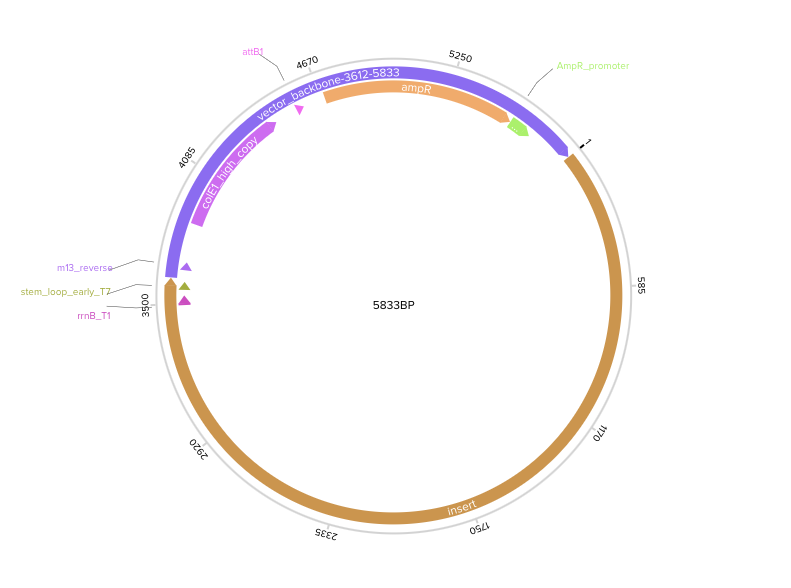
My annotated expression cassette
(JAX1 protein)[https://benchling.com/s/seq-kczgpR5k4eKn4BIEZYrC?m=slm-XqgAw2klZciZdJUaxluG]
Final Plasmid
Part 5: DNA Read/Write/Edit
Given my project for Vitiligo to develop a genetic circuit that detects reactive oxidative stress (ROS) and turns on genetic mechanisms to protect melanocytes, I would build the following bioengineering pipeline.
- DNA READ → Understand endogenous ROS regulatory DNA
- DNA WRITE → Build synthetic ROS-responsive circuit
- DNA EDIT → Insert circuit into melanocytes safely
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I would sequence the regulatory DNA that controls the oxidative stress response in melanocytes. My goal is to build a system that detects oxidative stress (ROS) in melanocytes and turn on protective genes. I would want to sequence:
The endogenous antioxidant response pathway genes in melanocytes
• Especially NRF2 (NFE2L2)
• KEAP1
• Antioxidant genes like HMOX1 (HO-1), NQO1, GCLC
Out of these, I would choose the NRF2, which is the master regulator of oxidative stress response. It activates antioxidant genes when ROS levels increase. If I understand the exact DNA sequence in melanocytes, I can:
• Identify natural promoter elements
• Find antioxidant response elements (AREs)
• Detect polymorphisms that affect stress response
• Design a synthetic circuit that mimics or enhances this pathway
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Also answer the following questions:
Is your method first-, second- or third-generation or other? How so?
What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
Since this is targeted sequencing of specific loci, I would use 2nd-gen, short-read sequencing (Illumina sequencing). Illumina sequencing has high accuracy, is suitable for variant detection. If I were studying structural variants or large rearrangements, I might use PacBio or Oxford Nanopore (third-gen). For my application, short-read high accuracy is ideal.
What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
Input: Genomic DNA extracted from melanocytes.
- DNA Extraction
- Lysis: Break open cells to release genomic DNA — ensures access to genetic material.
- Protein digestion: Remove histones and other proteins — prevents contamination and improves downstream efficiency.
- DNA purification: Isolate clean DNA — required for accurate fragmentation and library prep.
- Fragmentation: Mechanical (sonication) or enzymatic shearing: Break DNA into smaller pieces because sequencing platforms cannot read very long DNA directly. Target size ~300–500 bp: Optimal length for Illumina read chemistry and efficient cluster formation.
- End Repair: Generate blunt ends: Fix overhangs or damaged ends after fragmentation ensures uniform ends for adapter ligation.
- 4. A-tailing: Add single 3′ A overhang: Creates compatible ends for T-overhang adapters which increases ligation specificity and efficiency.
- Adapter Ligation
- Ligate Illumina adapters with index barcodes:
- Adapters allow DNA fragments to bind to the flow cell.
- Barcodes enable multiplexing of multiple samples in one sequencing run.
- PCR Amplification: Enrich adapter-ligated fragments: Selectively amplifies properly ligated DNA — increases library concentration and removes unligated fragments.
- Cluster Generation
- Bind to flow cell: Adapter sequences hybridize to complementary oligos on the flow cell surface which immobilizes fragments.
- Bridge amplification: Creates clonal clusters of identical DNA fragments — amplifies signal strength for accurate base detection during sequencing.
Illumina uses Sequencing by Synthesis (SBS):
1. Add fluorescently labeled reversible terminator nucleotides.
2. DNA polymerase incorporates ONE base.
3. Lasers excite fluorophores.
4. Camera records color → identifies base.
5. Terminator removed.
6. Cycle repeat, where each cycle = one base.
What is the output of your chosen sequencing technology?
The output is FASTQ files which contain Sequence reads, Quality scores (Phred scores). This can be used to align to reference genome and perform variant calling
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
I would synthesize a ROS-sensing genetic circuit that:
- Detects oxidative stress
- Activates antioxidant genes
- Protects melanocytes from damage
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
Also answer the following questions:
I would use Phosphoramidite DNA synthesis + Gene Assembly (Commercial gene synthesis, e.g., Twist Bioscience)
What are the essential steps of your chosen sequencing methods?
What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
I would use chemical DNA synthesis.
DNA Synthesis Workflow
- Solid-phase phosphoramidite synthesis: Chemically builds short DNA oligos base by base on a solid support to generate designed sequences.
- Oligo cleavage: Releases synthesized oligos from the solid support and removes protecting groups to obtain usable DNA.
- Oligo assembly (Gibson or enzymatic assembly): Joins overlapping oligos into a full-length gene construct.
- Cloning into plasmid backbone: Inserts the assembled gene into a plasmid for propagation and expression in cells.
- Sequence verification: Confirms the final DNA sequence is correct and free of synthesis errors.
Limitations
- Oligos longer than 200 nt are error-prone: Chemical synthesis efficiency decreases with length, increasing mutation rates.
- Repetitive elements reduce fidelity: Repeats promote misalignment during synthesis and assembly.
- GC-rich sequences are difficult: High GC content can impair synthesis and cloning efficiency.
- Cost increases with length: Longer constructs require more synthesis and validation steps.
- Cloning validation required: Errors introduced during synthesis must be identified by sequencing.
5.3 DNA Edit
(i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
- Edit melanocyte genomic DNA at a safe harbor locus (e.g., AAVS1).
- This allows stable insertion of a therapeutic construct without disrupting essential endogenous genes.
- Insert a ROS-responsive antioxidant genetic circuit. This enables melanocytes to activate protective genes only during oxidative stress, reducing unnecessary expression.
- Optionally enhance endogenous NRF2 pathway activity. Strengthening the native antioxidant response may improve resilience to oxidative damage implicated in vitiligo.
(ii) What technology or technologies would you use to perform these DNA edits and why?
I’ll use CRISPR-Cas9 genome editing. This system enables targeted DNA modification at precise genomic locations.
- Design a single guide RNA targeting the chosen safe harbor locus. The guide RNA determines the exact genomic cut site and ensures specificity.
- Provide a donor DNA template containing the ROS-sensing circuit with homology arms. Homology arms direct precise insertion via homology-directed repair.
- Deliver Cas9, guide RNA, and donor DNA into melanocytes ex vivo. Editing cells outside the body improves safety and allows selection of correctly modified cells.
- Screen and sequence edited clones. Verification ensures accurate insertion and absence of unintended mutations.
Also answer the following questions:
How does your technology of choice edit DNA? What are the essential steps?
- Cas9 introduces a double-strand break at the target site. The break initiates cellular DNA repair mechanisms.
- Cells repair the break using homology-directed repair when a donor template is present. This enables precise integration of the therapeutic circuit.
- The edited locus now contains the ROS-responsive construct. The inserted sequence becomes a permanent part of the melanocyte genome.
What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
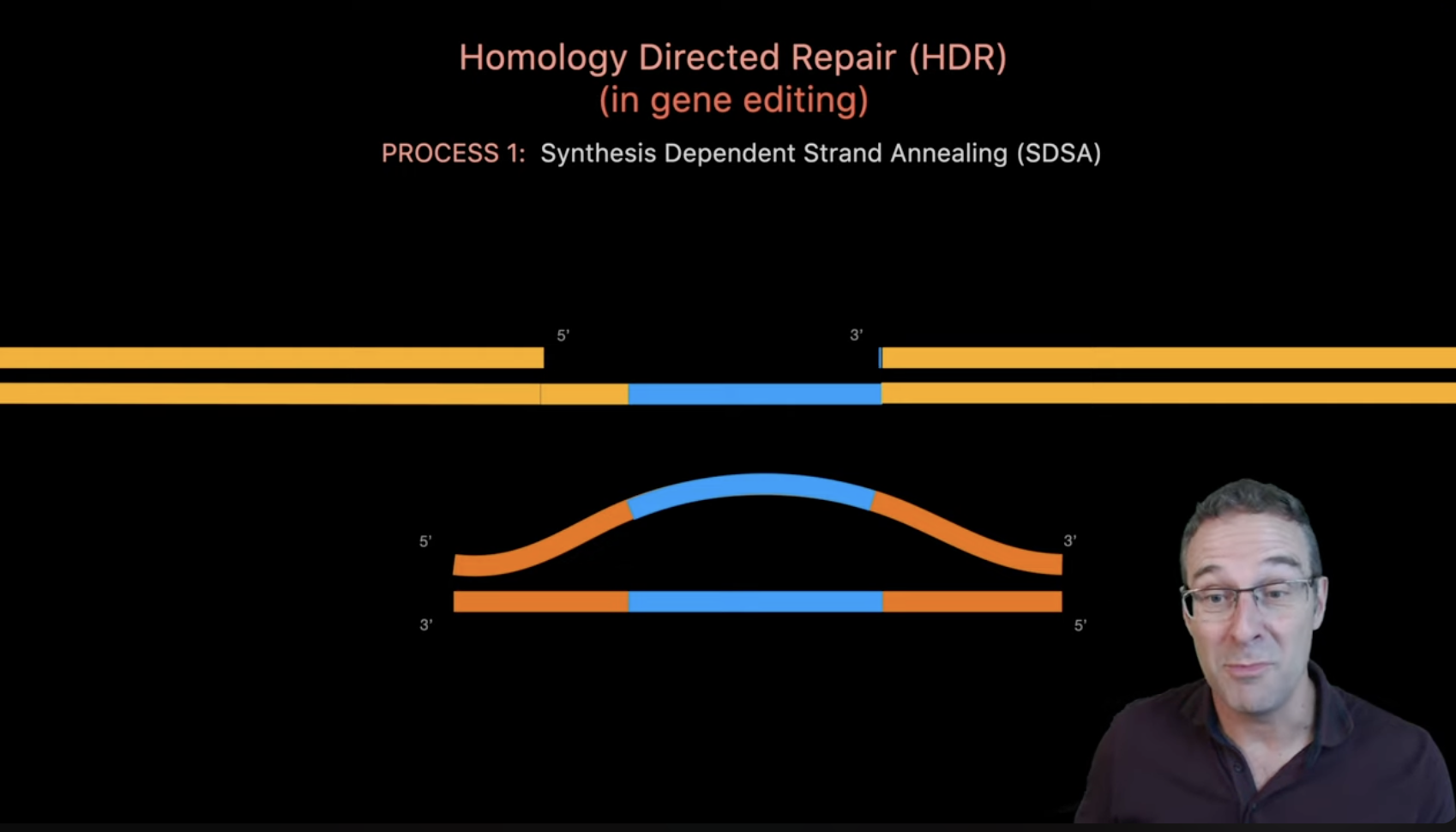
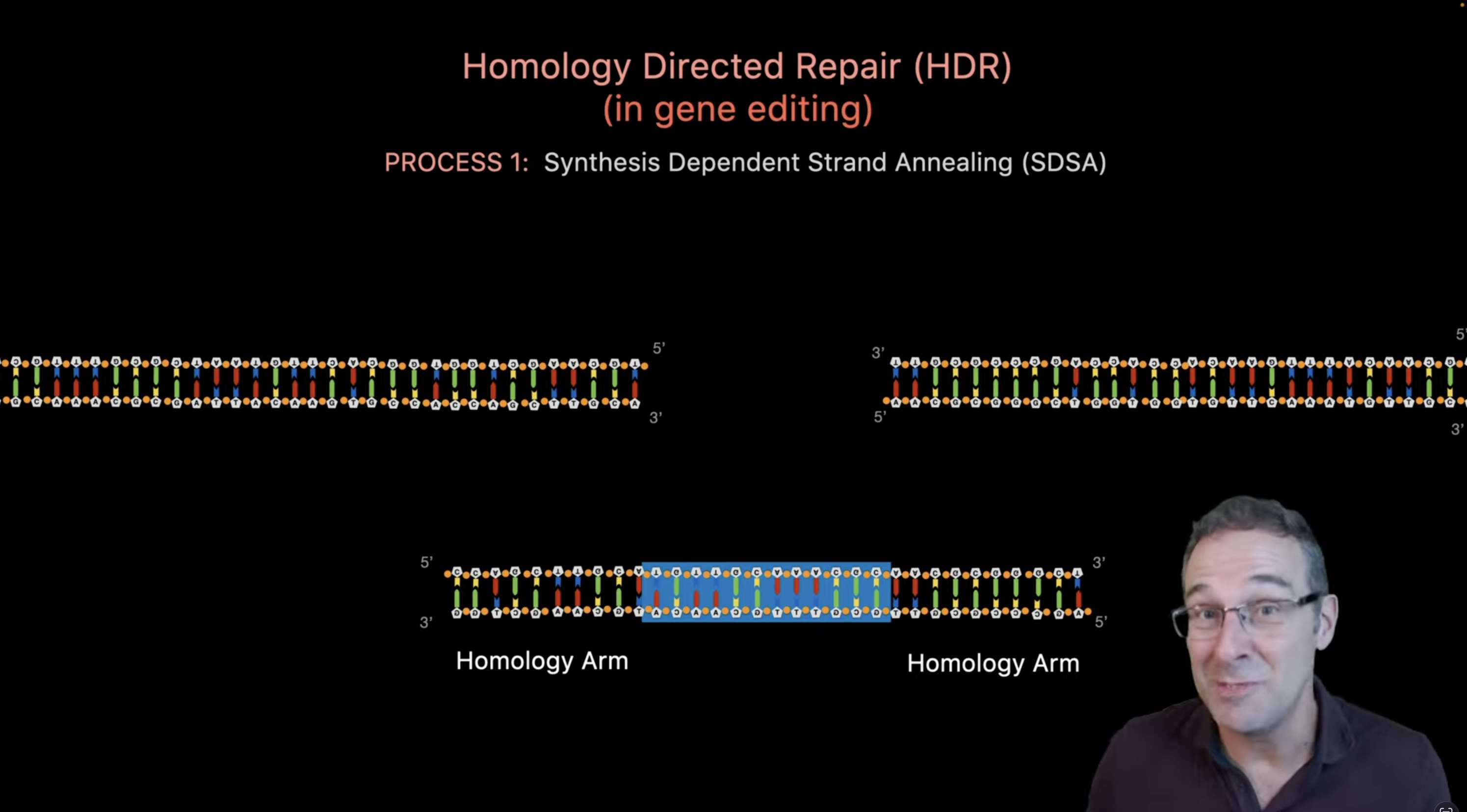




 (Source)[https://www.youtube.com/watch?v=cLMo6DYdJRE]
(Source)[https://www.youtube.com/watch?v=cLMo6DYdJRE]
- Cas9 protein or expression plasmid. Provides the nuclease that cuts DNA
- Single guide RNA (sgRNA). Directs Cas9 to the target locus
- Donor DNA template with homology arms. Supplies the sequence to be inserted.
- Cultured melanocytes: Serve as the editable cellular substrate.
What are the limitations of your editing methods (if any) in terms of efficiency or precision?
- Homology-directed repair efficiency can be low. Precise insertion may occur in only a fraction of treated cells.
- Off-target edits are possible. Imperfect guide specificity may lead to unintended genomic changes.
- Delivery into primary melanocytes can be challenging. Some cell types are difficult to transfect efficiently.
- Permanent genomic edits raise regulatory and safety concerns. Long-term monitoring would be required before clinical use.