Week 2 HW: DNA Read, Write, and Edit
HTGAA 2026 – DNA Gel Art Lab Report
Lab Partner: Alexandra Valdepeñas
Objective
The objective of this laboratory exercise was to create DNA get art using available restriction enzyme, prepare restriction digests, cast and run agarose gel, perform electrophoresis and compare observed DNA fragment patterns to the envisioned gel design.
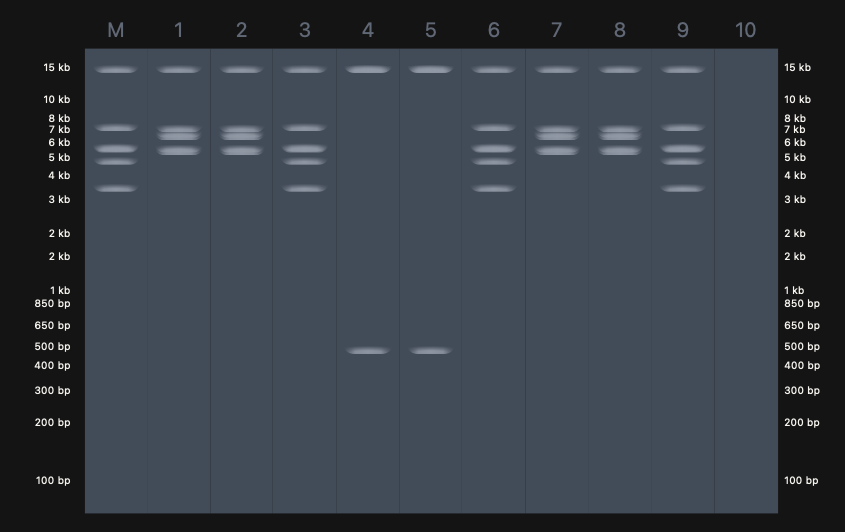
Part 0: We agreed to the following design:

Part I: Restriction Digest Setup
Restriction enzyme reactions were prepared using Lamda DNA, CutSmart buffer, and specific enzymes (EcoRI, BamHI, SalI) according to the virtual digest table.
Digest Reaction Map
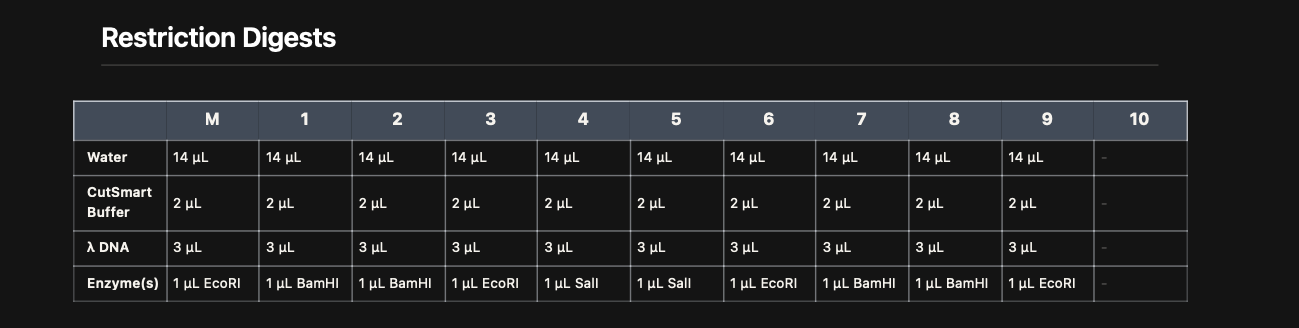

Each reaction contained:
- 14 µL water\
- 2 µL CutSmart buffer\
- 1 µL restriction enzyme
- 3 µL λ DNA\
The reagents were stored in an ice bucket and carefully pipetted.
Part II: Gel Preparation
Agarose was weighed and dissolved in buffer by heating until fully melted, then poured into a casting tray with comb inserted.


Part III: Sample Preparation
Digested DNA samples were carefully mixed with loading dye


The total reaction volume was 20 µL.
Part IV: Loading and Electrophoresis
The gel was placed into the electrophoresis chamber and submerged in buffer. Samples were loaded carefully into wells using micropipettes.




The chamber was connected to a power supply and run until adequate separation occurred.


Expected Result (Design)
The predicted fragment pattern based on known λ DNA restriction maps is shown below.


This design reflects: - Specific fragment sizes unique to EcoRI, BamHI, and SalI digests\
- Clear separation of fragments across molecular weight ranges\
- Distinct band counts per lane corresponding to enzyme cut frequency
Observed Result
The final gel image obtained experimentally is shown below.
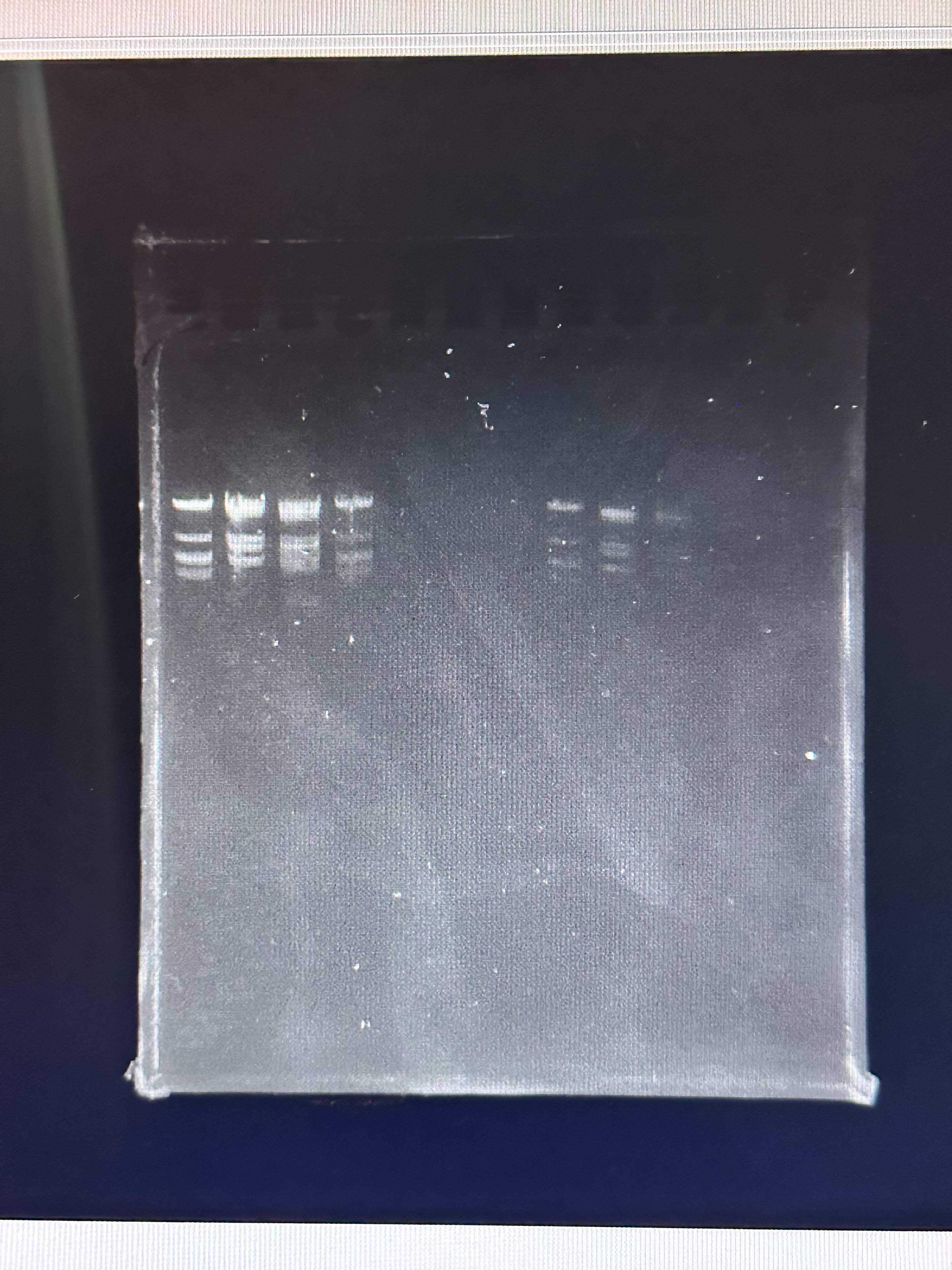

Compare it with our desired art:


Comparison and Analysis
Differences Observed
- Band Intensity Variation
- Some expected bands were faint or absent.
- Uneven intensity suggests incomplete digestion or uneven DNA loading.
- Missing or Merged Bands
- Some closely sized fragments appear merged.
- Likely due to insufficient gel resolution or short run time.
- Smearing
- Slight smearing suggests partial degradation, overloading, or suboptimal buffer conditions.
- Incomplete Digestion
- If enzyme activity was suboptimal (temperature, incubation time, or enzyme degradation), uncut or partially cut DNA would appear as unexpected higher molecular weight bands.
- Gel Concentration Effects
- Agarose percentage affects resolution. If not optimized for fragment size range, smaller fragments may not separate clearly.
- Running Conditions
- Voltage too high can cause band distortion.
- Insufficient run time reduces separation between fragments.
Why the Final Result Did Not Fully Match the Desired Design
The discrepancy between the predicted and observed gel likely results from a combination of:
- Partial restriction enzyme digestion
- Pipetting variability affecting enzyme concentration
- Inconsistent loading volumes
- Gel percentage not optimized for expected fragment sizes
- Limited electrophoresis duration
- Potential enzyme inactivation or improper incubation conditions
The theoretical design assumes complete digestion, perfect stoichiometry, and optimal gel resolution. In practice, small deviations in enzymatic efficiency, buffer composition, or electrophoresis parameters produce visible differences in band clarity and separation.
Conclusion
The experiment successfully demonstrated restriction digestion and agarose gel electrophoresis. While the observed banding pattern approximated the expected design, experimental variability led to differences in band intensity and resolution. These discrepancies highlight the importance of precise enzymatic handling, incubation control, gel optimization, and electrophoresis parameters in molecular biology workflows.