Radi KHODR — HTGAA Spring 2026

About me
I am passionate about interdisciplinary applications in the field of biotechnology. I worked on the developement of a bioremediation strategy of “forever pollutants” (also known as ‘PFAS’).


I am passionate about interdisciplinary applications in the field of biotechnology. I worked on the developement of a bioremediation strategy of “forever pollutants” (also known as ‘PFAS’).
Week 1 HW: Principles and Practices
(Image source : here) Biological Engineering Governance Analysis 1. The Scientific Application General Context Per- and polyfluoroalkyl substances (PFAS) represent one of the most intractable environmental contamination challenges of the 21st century. Comprising over 12,000 synthetic compounds characterized by extraordinarily stable carbon–fluorine (C–F) bonds — among the strongest in organic chemistry (bond dissociation energy ~544 kJ/mol) — PFAS have earned the designation “forever chemicals” for their near-complete resistance to biological, chemical, and photolytic degradation under ambient conditions. Their widespread use in aqueous film-forming foams (AFFF), food packaging, non-stick coatings, and industrial surfactants since the 1940s has resulted in ubiquitous environmental distribution: PFAS are now detectable in Arctic ice cores, deep ocean sediments, human breast milk, and the bloodstreams of wildlife on every continent. 1
Week 2 HW: DNA Read/Write/Edit
My homework Homework Questions from Professor Jacobson: [Lecture 2 slides] Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy? The error rate of a polymerase is 1:10^6 error rate with a 3.2 billion nucleotide in the human genome. In biology, base pairing and proofreading/erro correction mechanisms and layers exist to minimize mutations inside the genome. How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest? Proteins are strings of amino acids, there are 20 typical amino acids encoded by codons of 3 nucleotides. The genetic material in mRNA is written in codons, a set of specific consecutive triplets of nucleotides. Because RNA contains four different bases (A, U, C, G), there are 4³ = 64 possible codons. These 64 codons are more than enough to specify the 20 amino acids, resulting in some redundancy (multiple codons can code for the same amino acid). Additionally, three special codons function as stop signals that mark the end of the protein sequence. Also That being said, there are an enormous number of possible DNA sequences that can encode the same average human protein due to redundancy in the genetic code. Since most amino acids are encoded by multiple codons, a ~400-amino-acid protein could theoretically be encoded by roughly 3400 (~10190) different DNA sequences. However, in practice, many of these sequences do not work well because synonymous codons affect translation efficiency, mRNA stability and structure, splicing, protein folding, and overall gene expression. So while the amino acid sequence may be the same, the cellular outcome can differ significantly. Homework Questions from Dr. LeProust: [Lecture 2 slides] What’s the most commonly used method for oligo synthesis currently? Chemical synthesis (phosphoramidite method) Why is it difficult to make oligos longer than 200nt via direct synthesis? Why can’t you make a 2000bp gene via direct oligo synthesis? Since oligo synthesis is done by chemical synthesis, bases are added one at a time, increasing the number of errors thus as teh fragment size increases. Thus uligos longer than ~200 nt are difficult because each nucleotide addition has a small error rate, so errors accumulate with length—making direct synthesis of something like a 2000 bp gene impractical due to low yield and high mutation frequency. Homework Question from George Church: [Lecture 2 slides] Choose ONE of the following three questions to answer; and please cite AI prompts or paper citations used, if any.
 (Image source : here)
(Image source : here)
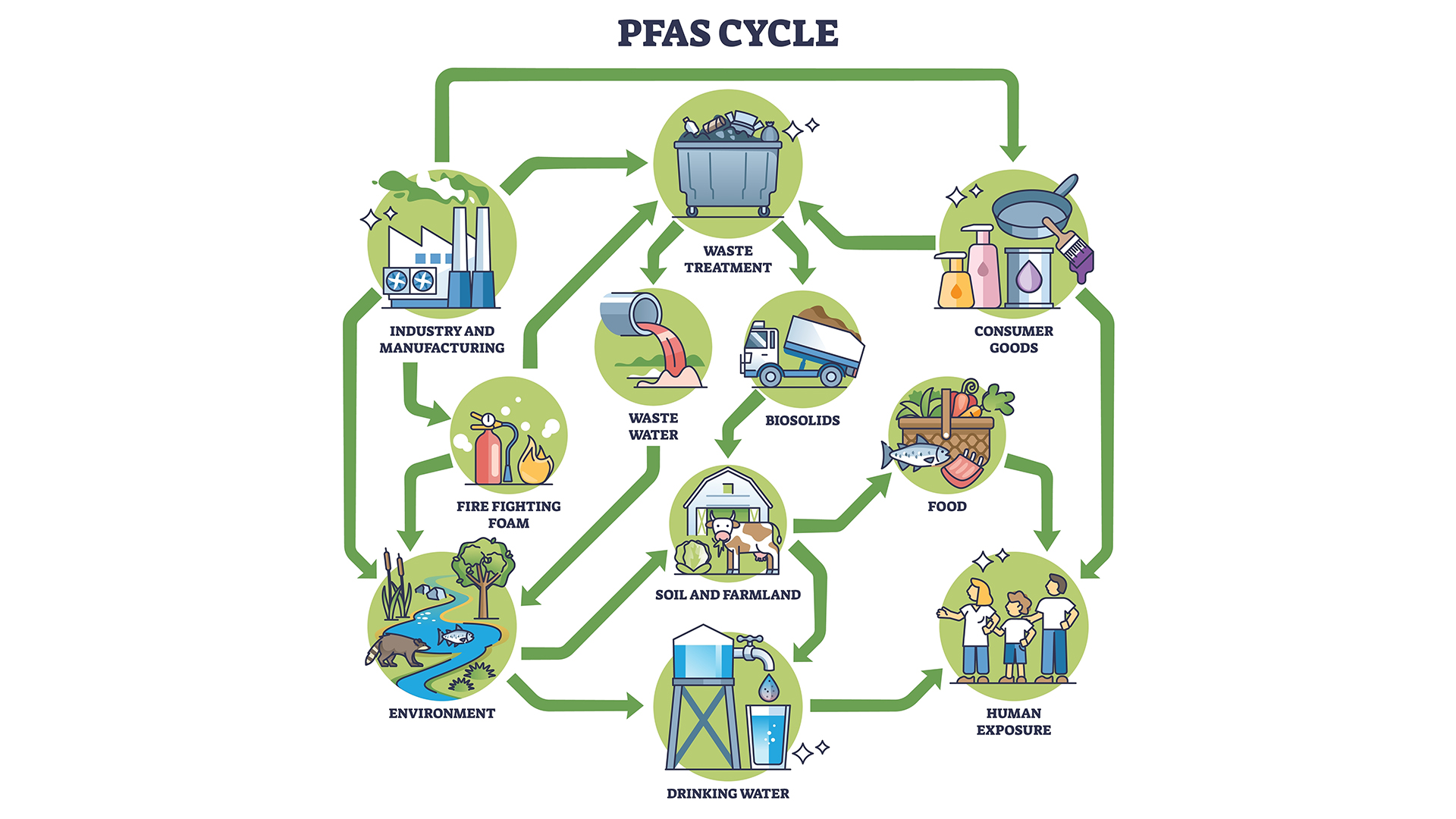
Per- and polyfluoroalkyl substances (PFAS) represent one of the most intractable environmental contamination challenges of the 21st century. Comprising over 12,000 synthetic compounds characterized by extraordinarily stable carbon–fluorine (C–F) bonds — among the strongest in organic chemistry (bond dissociation energy ~544 kJ/mol) — PFAS have earned the designation “forever chemicals” for their near-complete resistance to biological, chemical, and photolytic degradation under ambient conditions. Their widespread use in aqueous film-forming foams (AFFF), food packaging, non-stick coatings, and industrial surfactants since the 1940s has resulted in ubiquitous environmental distribution: PFAS are now detectable in Arctic ice cores, deep ocean sediments, human breast milk, and the bloodstreams of wildlife on every continent. 1
In this context, there is a need for PFAS depollution. One specific solution is “Bioremediation”, i.e. the use of biological components for the break-down and degradation of PFAS
Describe at least three governance actions. Each action may involve different actors and mechanisms.
Primary Actor(s):
Primary Actor(s):
Primary Actor(s):
“Pollution from PFAS could cost €440 billion on the European Union by 2050, if current contamination continues unchecked” 2
Score each governance action against the policy goals.
Scoring Key:
| Ban of PFAS | Engineer biological material | Trust Fund | |
|---|---|---|---|
| Enhance Biosecurity | 1 | 3 | N/A |
| Foster Lab Safety | 1 | 3 | 3 |
| Protect the environment | 1 | 1 | 1 |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 3 | 3 | 1 |
| • Feasibility? | 3 | 1 | 1 |
Top Priority Action(s):
Rationale:
This option allows to advantage on the scientific level and bypassing the regulatory and legal hurdles that can frain the developement of scientific applications. Debates are still being done at a national and international levels concerning the feasibility and necessity of banning PFAS and even some experts argue that such a generalistic decision could do more harm than good by ignoring the fact that some PFAS are needed and they aren’t all troublesome.
Key Trade-offs Considered:
Choose ONE of the following three questions to answer; and please cite AI prompts or paper citations used, if any.