Week 2 HW: DNA Read, Write, & Edit
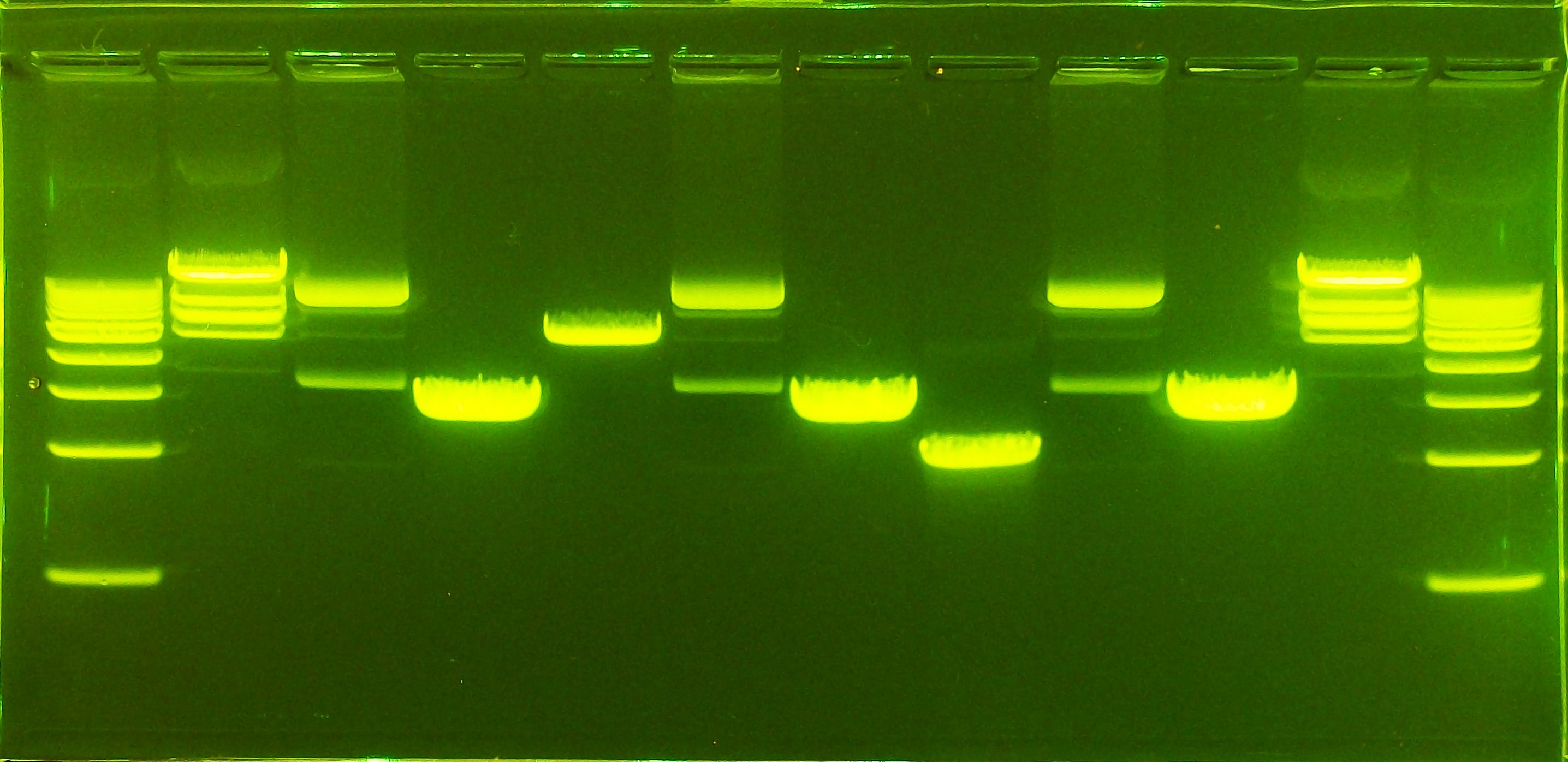

Part 1: Benchling & In-silico Gel Art
I was made a random gel art. It took me one hour just to understand and make this lol :v I use several enzymes and this is it
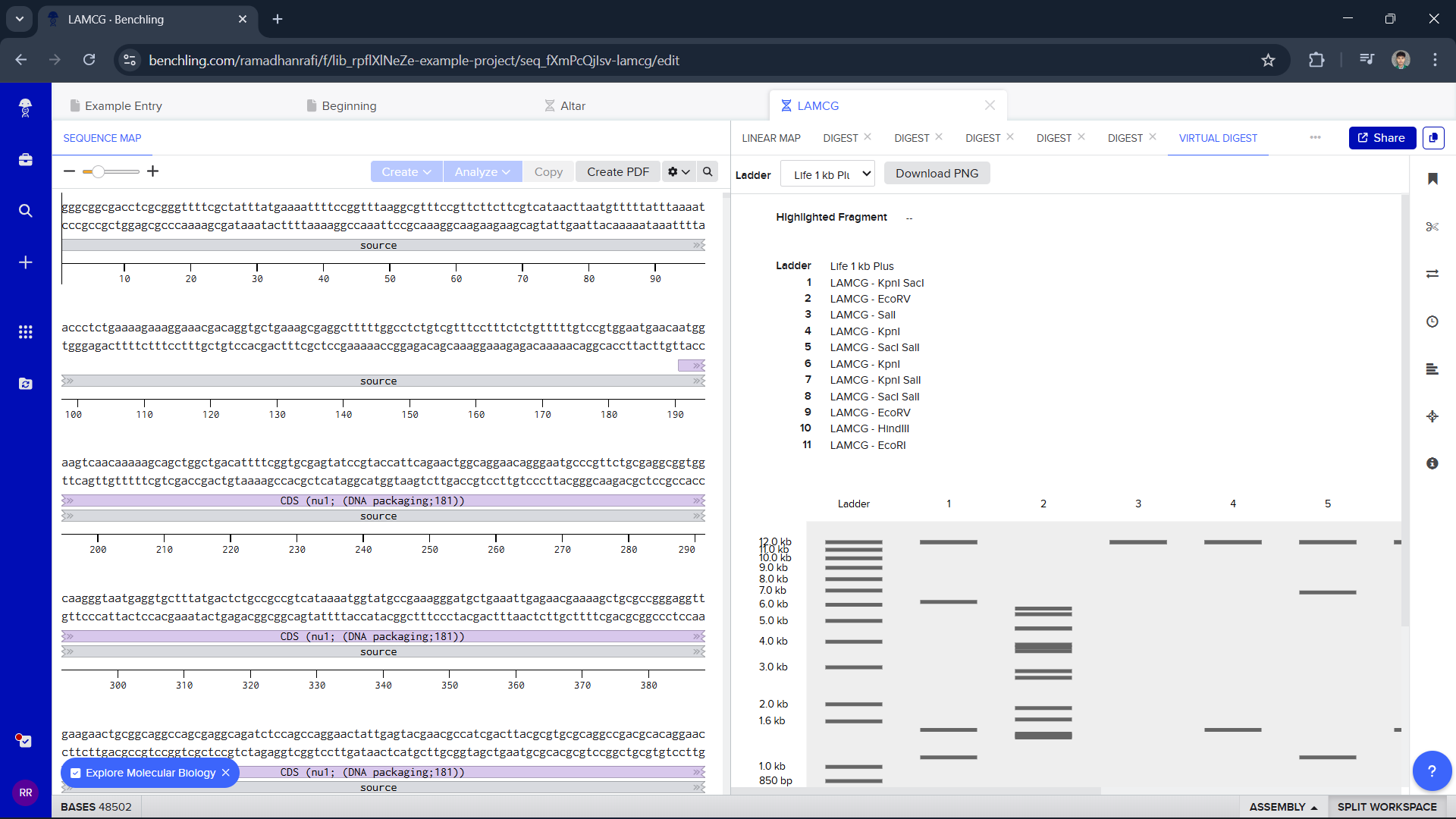

Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
I can’t perform this gel art in the lab because I don’t have the tools to make it, but it’s fun to see what I can make :)
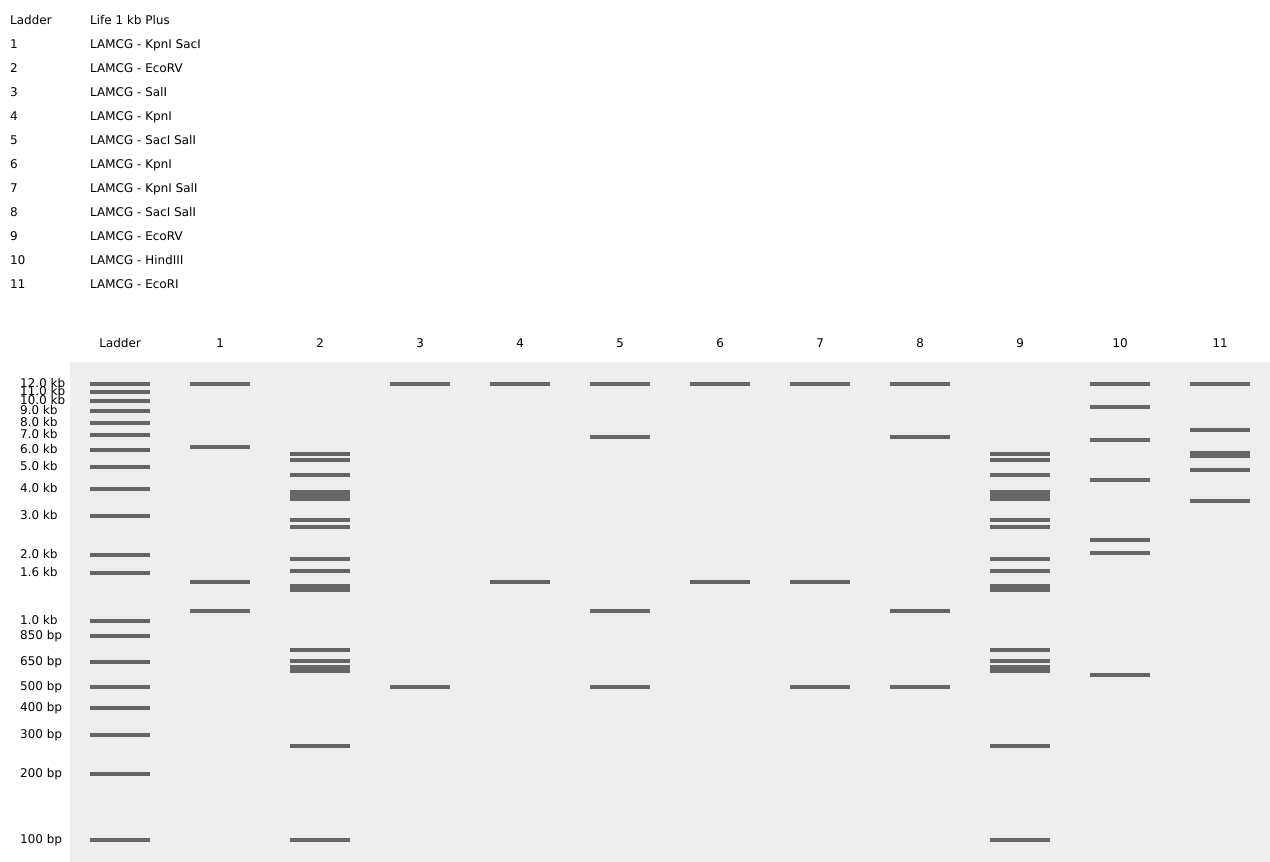

Part 3: DNA Design Challenge
I use Uniprot to get the sequence of the insulin protein
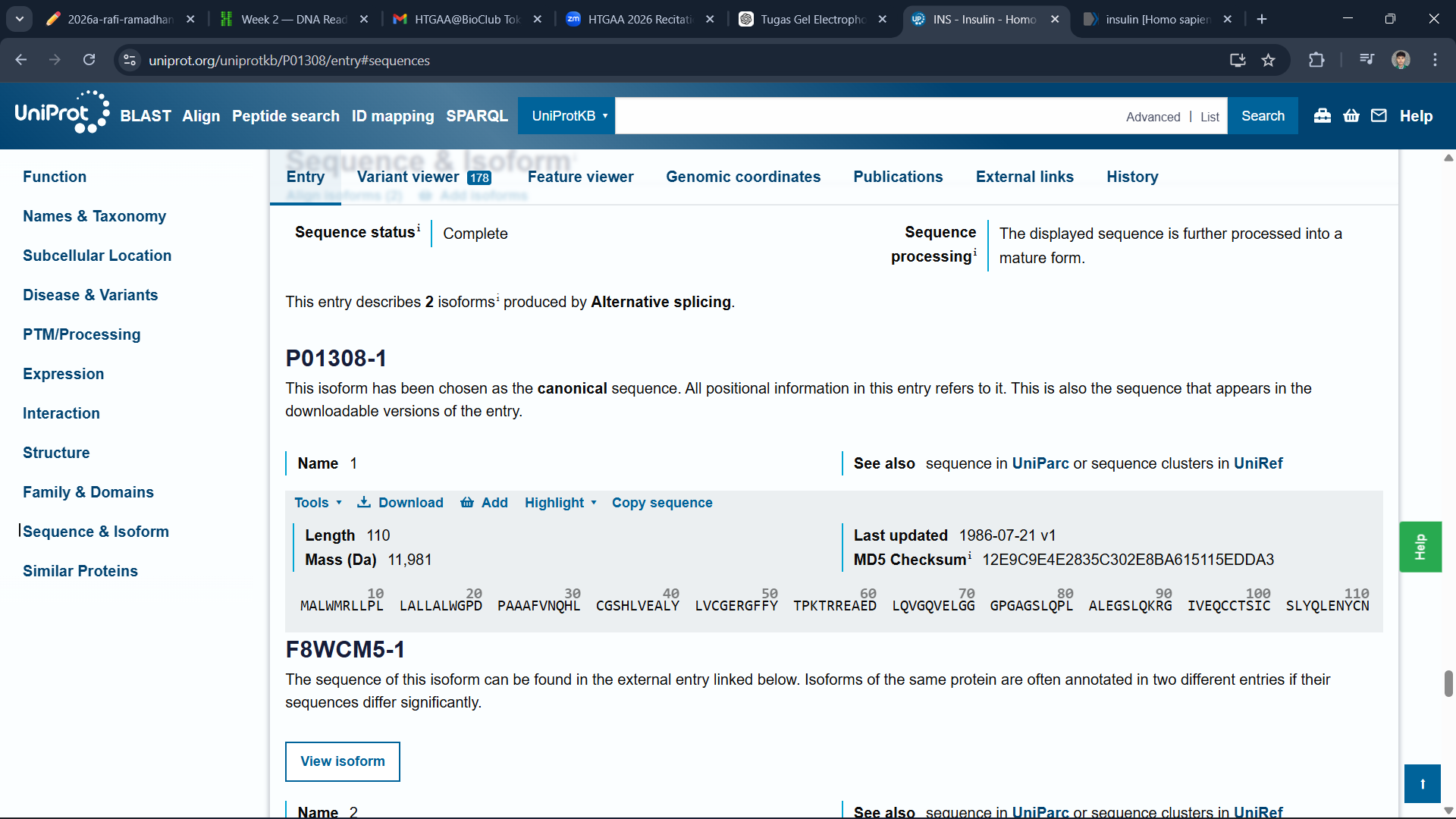

After copying the sequence, I use bioinformatics.org to reverse the protein sequence into a DNA sequence
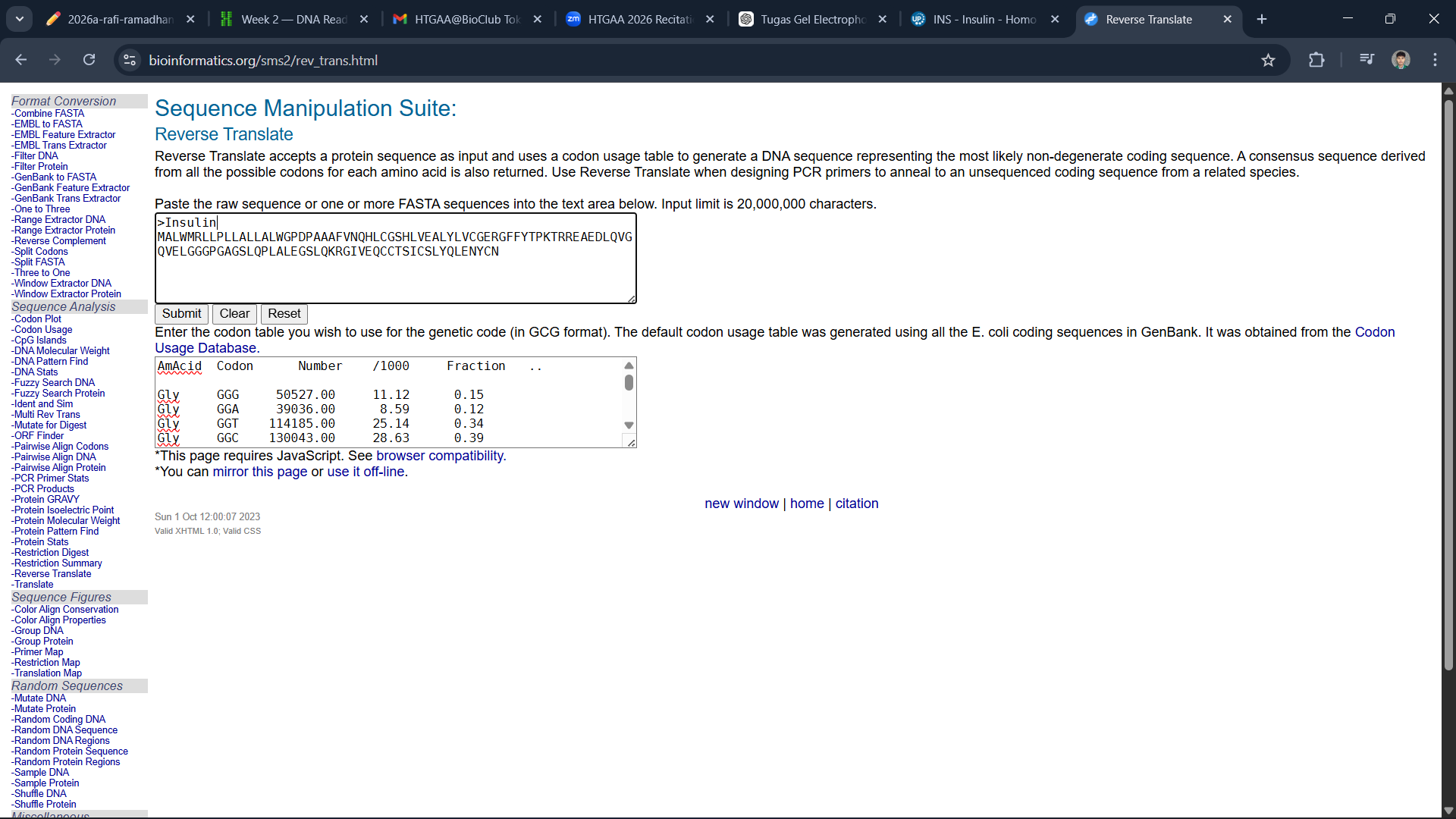

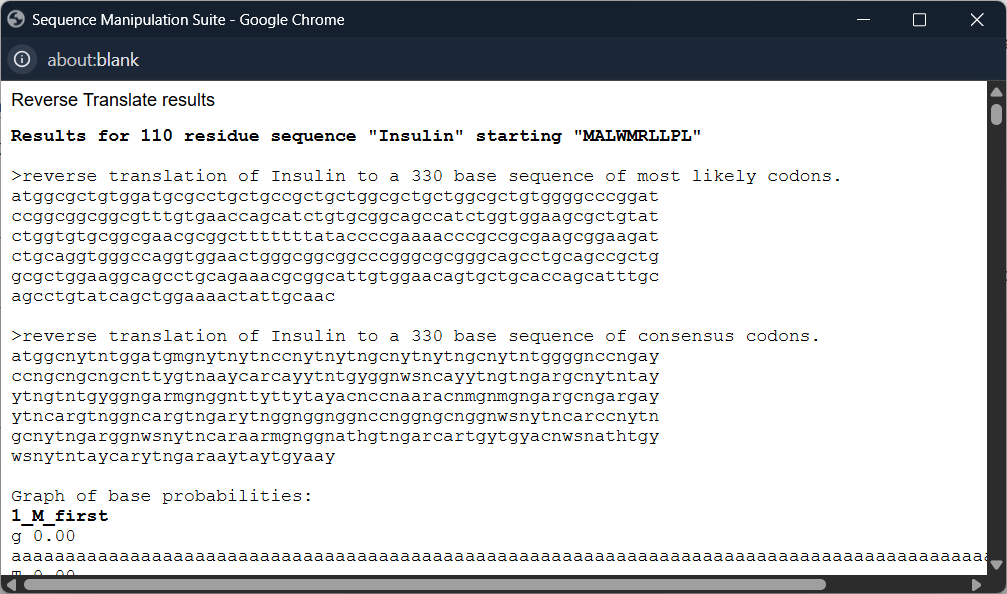

Since all organisms have different codon preferences, I optimized the DNA sequence obtained from Homo sapiens using the VectorBuilder website and excluded the enzymes BsaI, BsmBI, and BbsI.
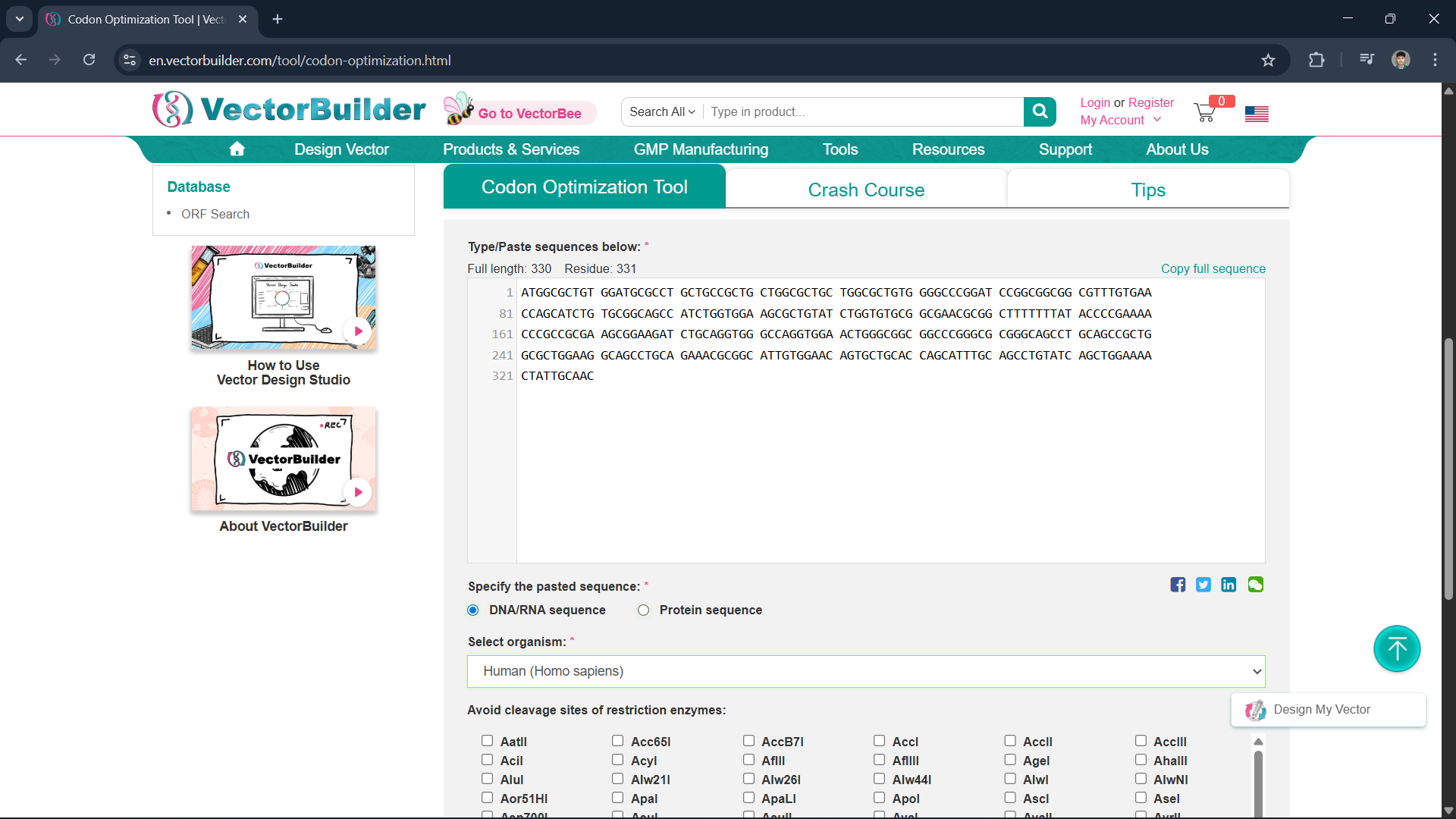

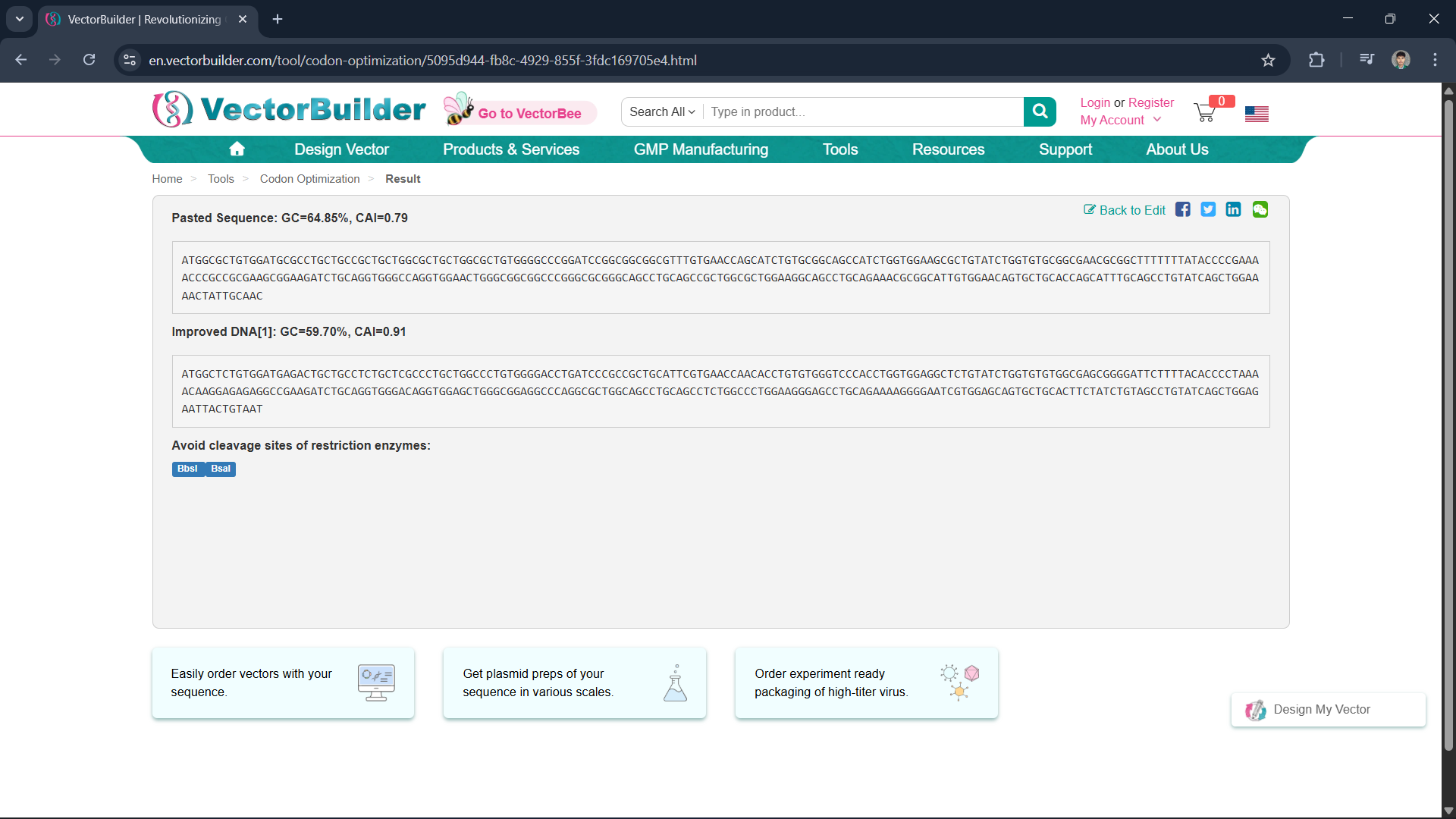

Insulin protein can be produced from DNA sequences using recombinant DNA technology, either through a cell-based system (E. coli) or a cell-free system. The optimized insulin DNA sequence is inserted into a plasmid. The plasmid is then inserted into E. coli cells and transcribed within the cells by RNA polymerase. Ribosomes then read the mRNA in the form of codons and translate it into amino acid chains. The resulting polypeptide chains then undergo folding and form a complete protein structure.
Part 4: Prepare a Twist DNA Synthesis Order
To build a plasmid, I use the sequence of E.coli and edit it in Benchling and Twist
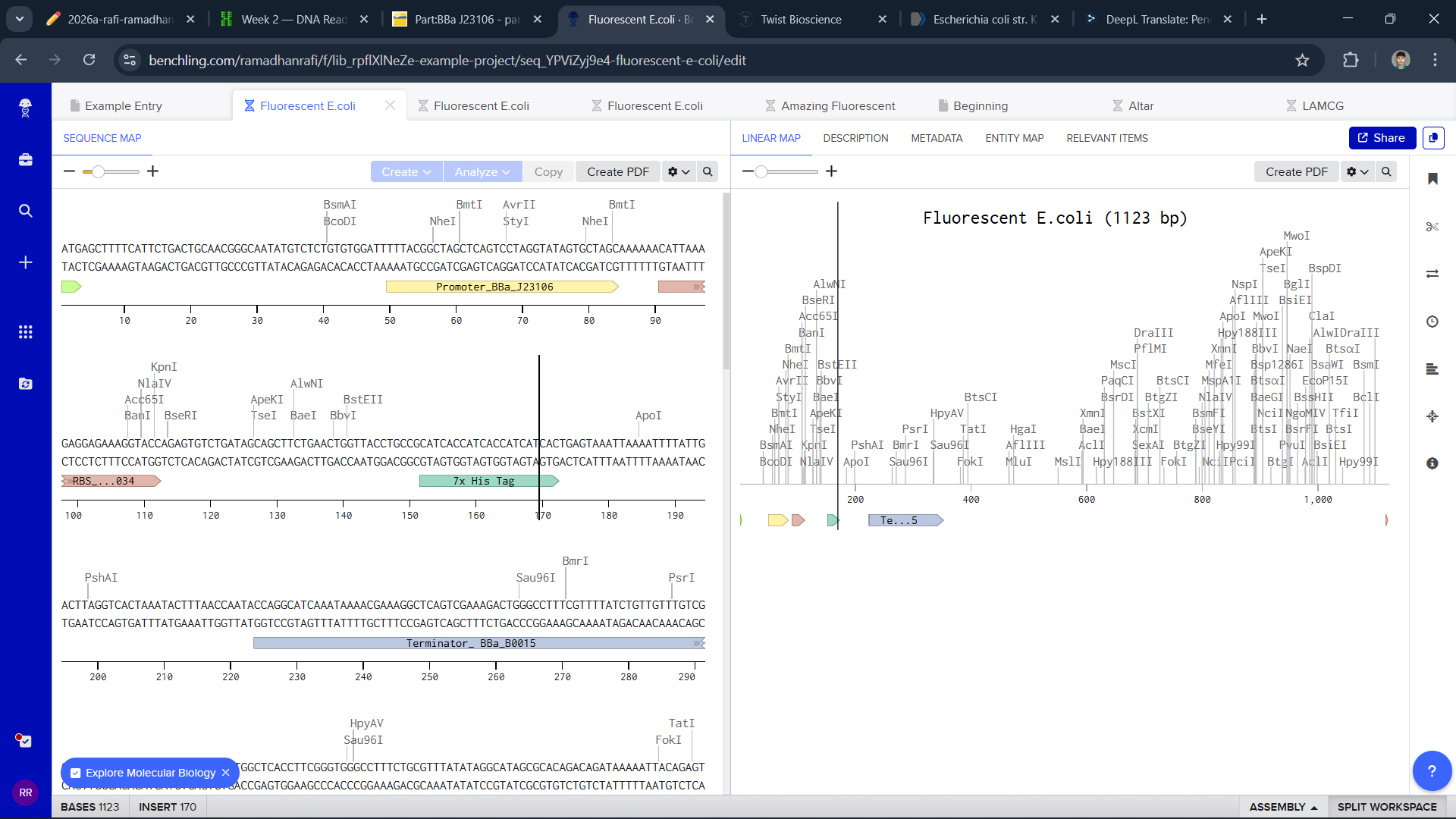

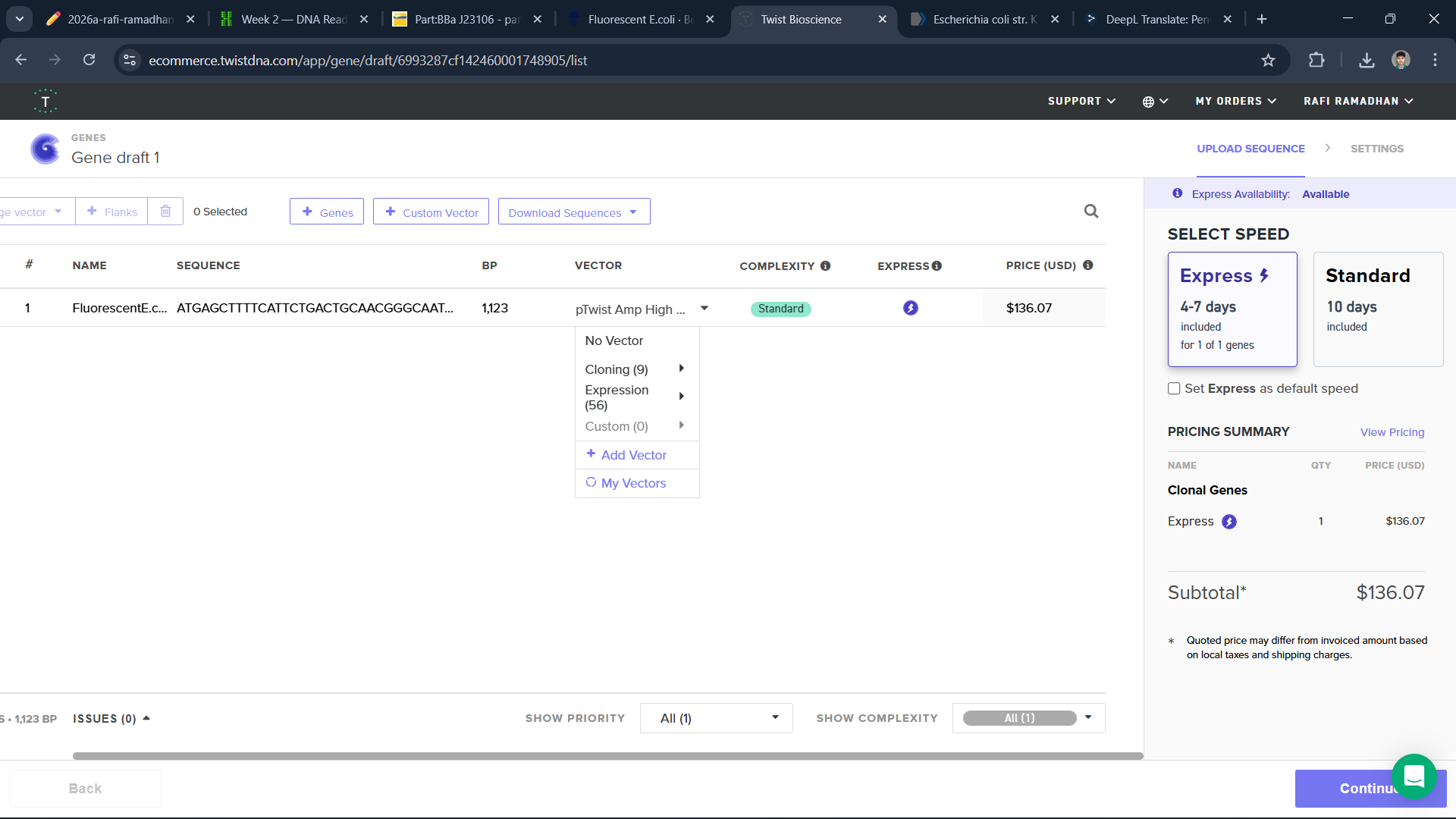

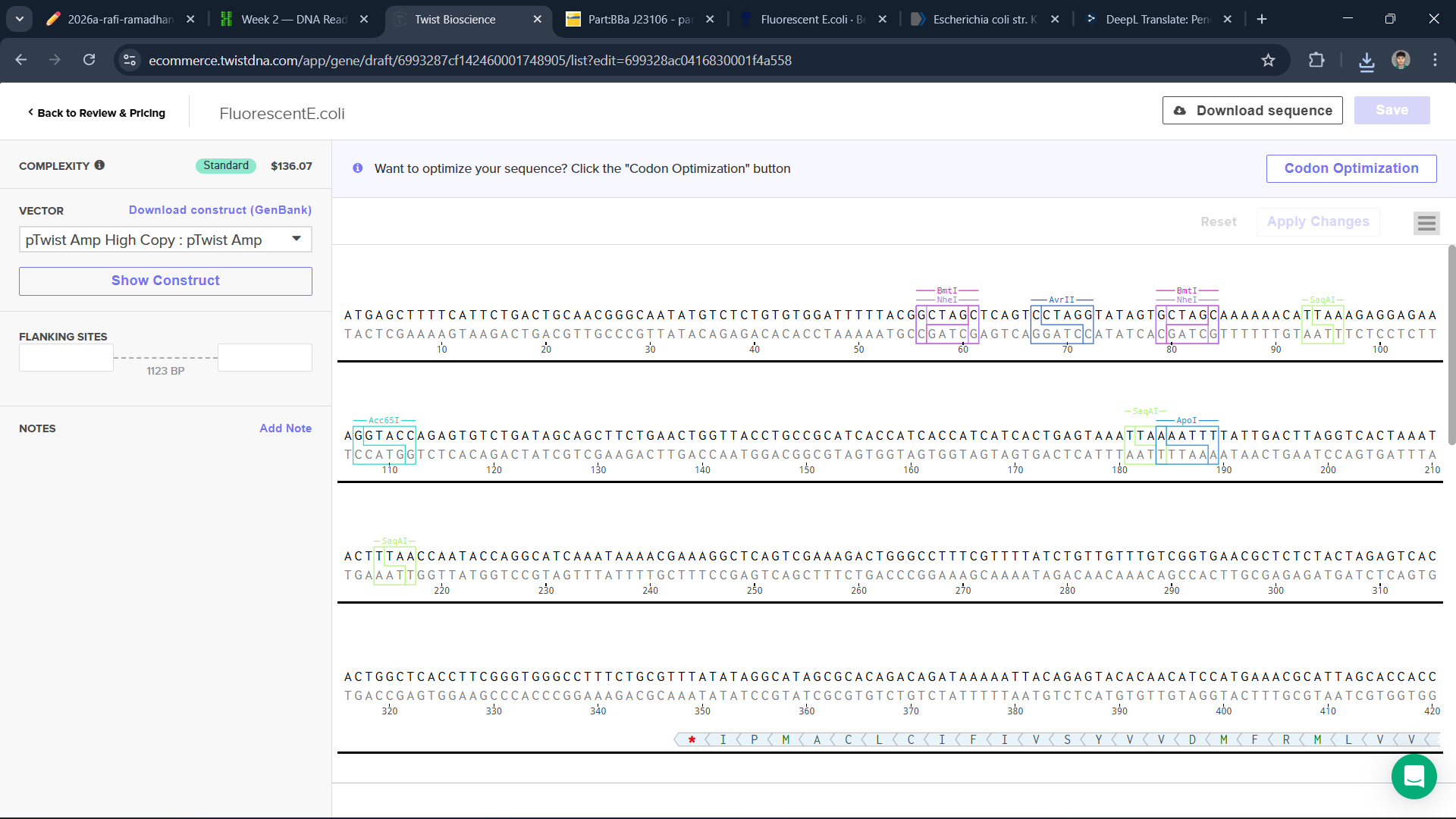

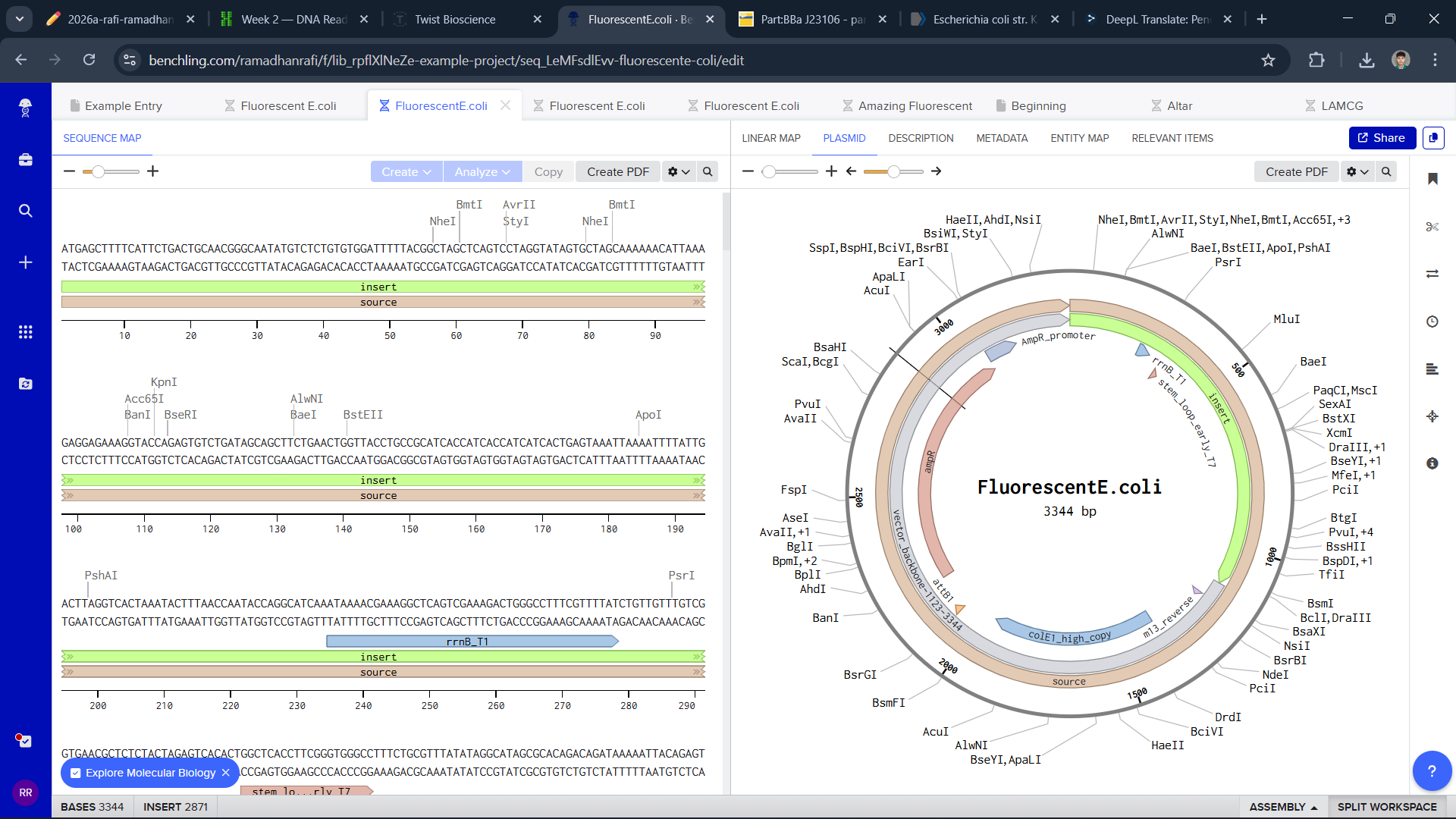
 I think this is a good plasmid that I made for the first time
I think this is a good plasmid that I made for the first time
Part 5: DNA Read/Write/Edit
5.1 DNA Read
I want to analyze the DNA of the LRRK2 gene associated with Parkinson’s disease in order to identify the mutations that cause the disease and determine the appropriate treatment therapy.

I want to use second-generation technology (Next Generation Sequencing/NGS) because it is widely used and accurate. Here, I will input pure DNA from the extraction, prepare the sample with extraction and PCR amplification, read the bases using software to convert the signals into A, T, C, G, and eventually produce the base sequence output.
5.2 DNA Write
I want to synthesize anti-inflammatory protein expression plasmid DNA for use in nano research. The technology used is chemical DNA synthesis with the following process:
- Sequence design on a computer
- Base synthesis one by one
- Deprotection
- Purification
- Assembly (if long)
- Cloning into a plasmid
5.3 DNA Edit
I really want to try editing the Parkinson’s and Alzheimer’s genes in humans to help people maintain their memory, or perhaps create drought-resistant genes in rice to preserve food supplies. The main editing technology used is CRISPR-Cas9. Guide RNA will detect the target DNA and then CAS9 will cut the DNA and repair the cell, which will then be repaired by the body. Required inputs:
- Cas9 plasmid
- Guide RNA
- Template DNA
- Target cells However, there are several limitations here, namely ethical issues and the extremely high cost involved.