Week 2 HW: DNA Read, Write, & Edit
Part 1: Benchling & In-silico Gel Art


Info
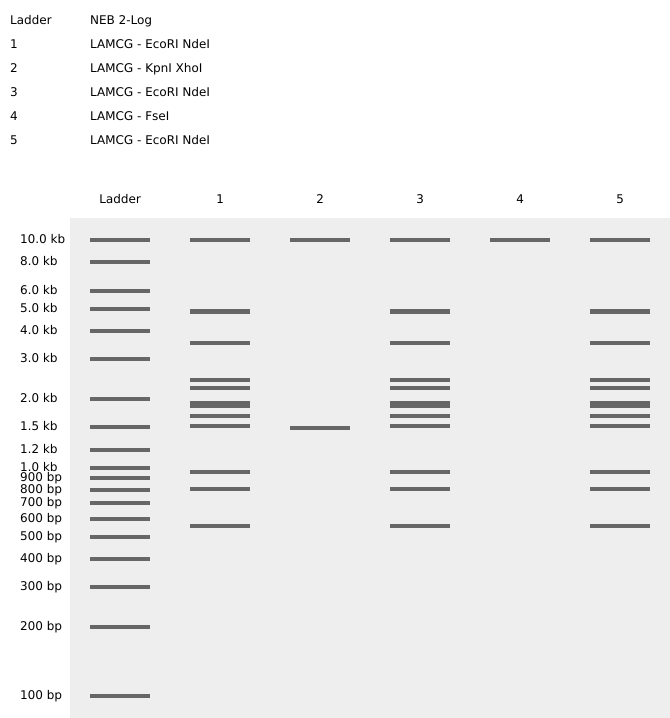
This is a picture of the gel art I designed on Benchling. The bands in the 1-6 ladders create the word “Hi” on completion. The restriction enzymes used on the Lambda DNA are listed above the diagram.
Part 3: DNA Design Challenge
3.1. Choose your protein.
The protein that I chose is the human testis-determining factor. I chose this protein because I find it interesting how one gene plays such a big role in sex differentiation and is the largest factor in deciding how a human embryo will grow. It is interesting to think about how one type of protein encoded by one gene in the human genome can spark significant change in the entire development process of humans. This sequence was taken from the NCBI:
>NP_003131.1 sex-determining region Y protein [Homo sapiens]
MQSYASAMLSVFNSDDYSPAVQENIPALRRSSSFLCTESCNSKYQCETGENSKGNVQDRVKRPMNAFIVWSRDQRRKMALENPRMRNSEISKQLGYQWKMLTEAEKWPFFQEAQKLQAMHREKYPNYKYRPRRKAKMLPKNCSLLPADPASVLCSEVQLDNRLYRDDCTKATHSRMEHQLGHLPPINAASSPQQRDRYSHWTKL
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
> reverse translation of NP_003131.1 sex-determining region Y protein [Homo sapiens] to a 612 base sequence of most likely codons.
ATGCAGAGCTATGCGAGCGCGATGCTGAGCGTGTTTAACAGCGATGATTATAGCCCGGCGGTGCAGGAAAACATTCCGGCGCTGCGCCGCAGCAGCAGCTTTCTGTGCACCGAAAGCTGCAACAGCAAATATCAGTGCGAAACCGGCGAAAACAGCAAAGGCAACGTGCAGGATCGCGTGAAACGCCCGATGAACGCGTTTATTGTGTGGAGCCGCGATCAGCGCCGCAAAATGGCGCTGGAAAACCCGCGCATGCGCAACAGCGAAATTAGCAAACAGCTGGGCTATCAGTGGAAAATGCTGACCGAAGCGGAAAAATGGCCGTTTTTTCAGGAAGCGCAGAAACTGCAGGCGATGCATCGCGAAAAATATCCGAACTATAAATATCGCCCGCGCCGCAAAGCGAAAATGCTGCCGAAAAACTGCAGCCTGCTGCCGGCGGATCCGGCGAGCGTGCTGTGCAGCGAAGTGCAGCTGGATAACCGCCTGTATCGCGATGATTGCACCAAAGCGACCCATAGCCGCATGGAACATCAGCTGGGCCATCTGCCGCCGATTAACGCGGCGAGCAGCCCGCAGCAGCGCGATCGCTATAGCCATTGGACCAAACTG
3.3. Codon optimization.
Optimizing a codon sequence can have various impacts. Codon optimization replaces less-favored codons in a specific organism with more common codons. An optimized codon sequences has a higher efficiency in translation which then leads to higher levels of protein expression. In addition, an optimized codon improves the stability of the mRNA since they are more likely to be recognized by tRNAs. Overall, an optimized codon is more likely to have increased protein expression.
> Optimized codon sequence of NP_003131.1 sex-determining region Y protein to Humans (Homo sapiens).
ATGCAGTCCTATGCCTCCGCCATGCTGAGCGTGTTTAACAGTGATGACTACTCCCCAGCCGTGCAGGAGAACATCCCAGCCCTGAGACGCAGCAGCTCATTCCTGTGTACCGAGTCTTGCAACTCCAAGTACCAGTGCGAGACCGGCGAGAACAGTAAGGGAAACGTGCAGGATCGCGTGAAAAGGCCCATGAACGCTTTCATCGTGTGGAGCCGCGATCAGAGGAGGAAGATGGCCCTGGAGAATCCCAGGATGCGGAACAGCGAAATCTCCAAGCAGCTGGGCTACCAGTGGAAGATGCTGACCGAGGCCGAGAAGTGGCCATTTTTCCAGGAGGCACAGAAGCTGCAGGCCATGCACAGAGAGAAGTACCCCAATTACAAGTACAGACCCAGAAGAAAGGCCAAAATGCTGCCTAAGAACTGTTCCCTGCTGCCCGCCGACCCAGCCTCCGTGCTGTGCTCTGAAGTCCAGCTGGACAACCGCCTGTACAGAGACGACTGTACCAAGGCCACCCACTCCCGCATGGAACACCAGCTGGGGCACCTGCCCCCCATTAATGCCGCATCCTCCCCCCAGCAGCGCGACCGGTACAGCCACTGGACAAAGCTG
3.4. You have a sequence! Now what?
To produce this protein from my DNA, we can use a multitude of both cell-dependent and cell-free methods.
We can living cells as “factories” for our proteins. We can design a plasmid containing our TDF-encoding gene that also has a promoter, ribosome binding site, and antibiotic resistance marker. Through transforming a host cell (like Escherichia coli) with this DNA, we can then induce expression of the TDF protein within the cells. Upon harvesting and purifying the protein, we can then have a batch of TDF proteins.
If a cell-free system was preferable, we could combine ribosomes, tRNAs, polymerases, amino acids + nucleotides, and an energy system with our DNA to create our protein in a test tube. This method would be much faster.
Part 4: Prepare a Twist DNA Synthesis Order
Info
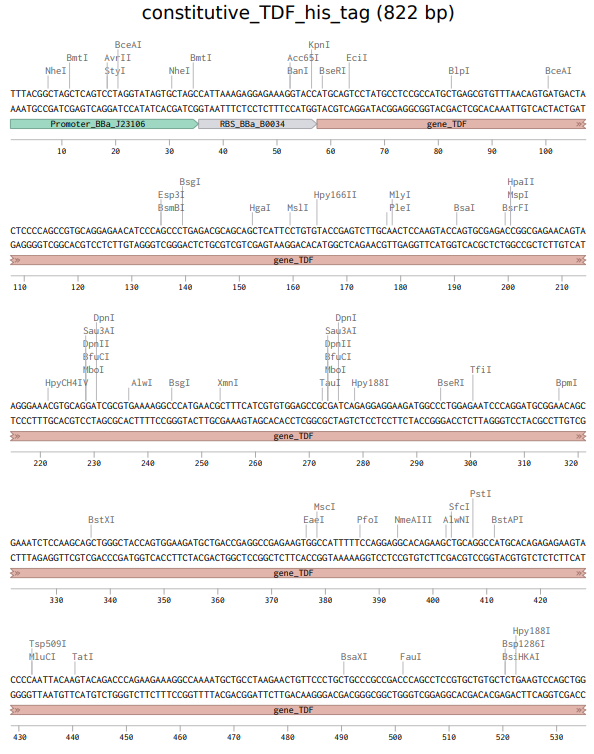
The above photo is the sequence of the expression cassette to express TDF proteins. This photo was taken in Benchling.

Info
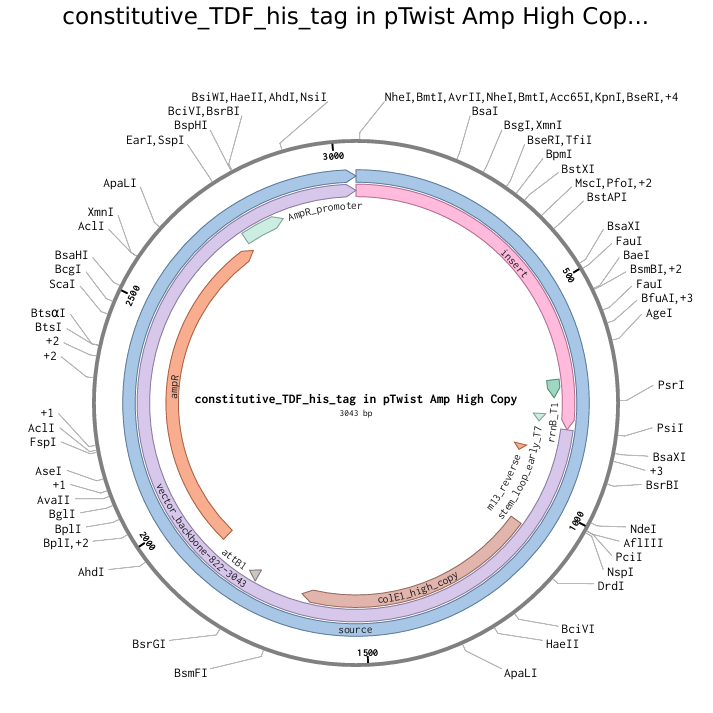
The above photo is a pTwist Amp High Copy plasmid backbone with the TDF-expressing insert (shown by “insert”). This photo was taken in Twist Bioscience.
Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
I would want to sequence the genome of a lactose intolerant person. I want to sequence this genome to better understand what genes are implicated in the reduced expression of lactase in lactose-intolerant patients.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
To sequence my DNA, I would use Illumina Whole-Genome Sequencing. I would use this method as it can give a complete, highly accurate view of someone’s entire DNA sequence. This is a second-generation sequencing method that can sequence millions of short fragments in parallel and uses PCR amplification. For our input, we must extract blood/saliva and purify the genomic DNA of these cells. Then, we shear the DNA into ~200-500 bp fragments using enzymes. Once we ligate synthetic adapters to both ends of the fragments, we can then PCR amplify the adapter-ligated fragments. The prepared DNA can be combined with complementary oligos from a flow cell to generate clusters. This method uses a sequencing-by-synthesis method. By adding fluorescently labled nucleotides, we can use a camera to record which color/nucleotide attached to the sequence. After removing the nucleotide chemically and repeating the process with each type of nucleotide, we can then generate a raw FASTQ file.
DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
I would like use synthesize a sequence of DNA that can work in a cell-free system and detect certain molecules present in a disease. This circuit will detect specific molecules from a pathogen like malaria. I want to create this biosensor to give impoverished areas a way to detect disease with cheap methods.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
I will perform this DNA synthesis using Gibson Assembly and molecular cloning. I can order the parts of the genetic circuit and assemble them accordingly using Gibson Assembly. After assembling the plasmid, the plasmid can be amplified through transformation, cloning in a bacterial cell, and purification. The limitations of this method is that it can be slower due to colony screening and sequencing verification and could include failed ligations, wrong inserts, etc. It also requires that I have already synthesized DNA.
DNA Edit
(i) What DNA would you want to edit and why?
One application for editing human DNA is to cure lactose intolerance. By editing the genome of a lactose intolerant human to be able to produce lactase, we can cure his lactose intolerance and allow him to consume foods with lactose. This application of gene editing is just one example of how synthetic biology can be leveraged to solve common human disorders.


(ii) What technology or technologies would you use to perform these DNA edits and why?
Lactose intolerance is commonly caused due to reduced expression of the lactase enzyme. To fix this, one could modify regulatory variants near the LCT gene in the human genome. The best way of doing this uses base editing systems (a CRIPSR-derived technology) to convert one base into another at a targetted location. This process works by a guide RNA first directing a Cas protein to a specific DNA sequence. Upon finding this DNA sequence, the Cas protein binds to the target site and a fused enzyme will then chemically convert the base into another base. After the cell’s repair mechanisms fix the strand, there will be a single-letter DNA change.
To prepare to preform this DNA edit, you must first identify which regulatory variant is implicated in adult lactase expression. Then, you have to prepare a sequence-specific guide RNA, base editor protein, and delivery system (maybe viral?). As a note, this system will target human intestinal stem cells in vivo. This method, however, carries various limitations. Since it is in vivo, delivery will be extremely difficult and all cells will not be edited (mosaic editing). In addition, this editing method may cause unintended edits in other areas of the genome.