Week 6 HW: Genetic Circuits Part I: Assembly Technologies

Assignment: DNA Assembly
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion High-Fidelity PCR Master Mix consists of Phusion DNA Polymerase, deoxynucleotides, and a reaction buffer (including MgCl2). The Phusion DNA Polymerase is a high-fidelity enzyme that is used to synthesize new, complementary nucleotides to the 3’ end of a DNA strand. Deoxynucleotides are present within the master mix to be added to the cloned DNA strand. The reaction buffer facilitates enzymatic function and stabilizes the DNA polymerase, allowing the PCR reaction to proceed smoothly.
- What are some factors that determine primer annealing temperature during PCR?
The annealing temperature for a PCR reaction the primer base composition (proportions of A, T, G, and C), primer concentration, primer length, and ionic reaction environment.
There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR Restriction Enzyme Digest Purpose Create millions of copies of a DNA segment (amplify the DNA segment). Cut DNA at a specific site. Active Enzymes DNA Polymerase Restriction Endonucleases Targetting Defined by synthetic primers. Defined by palindromic recognition sequences. Temperature Alternates between denaturation, annealing, and extension temperatures (95°C, 55°C, 72°C, respectively). Usually a steady incubation at 37°C. Result High-concentration amplicons of a single size Different sized fragments of DNA based on the number of sites Input DNA Minimal amount of DNA template High concentrations of DNA How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
We ensure that the DNA sequences that we have digested and PCR-ed are appropriate for Gibson cloning through preparing the correct conditions for Gibson cloning. The two DNA inserts we create must have identical ends that will overlap with one another. This is to ensure that they will stick together during Gibson assembly. In addition, the DNA used must be treated with DpnI and a cleanup kit (column purification) to remove the original template DNA and leftover salts/enzymes.
- How does the plasmid DNA enter the E. coli cells during transformation?
The most common method for plasmid DNA to enter E. Coli cells during transformation is chemical transformation. In this method, cells are treated with calcium chloride which neutralizes the negative charges on the DNA and cell membrane to bring them closer to one another. Then, the cells are cooled down and suddenly heated to 42°C. This sudden change in temperature creates a pressure difference between the inside and outside of the cell, forming temporary pores in the membrane which allow the entry of the plasmid DNA into the cytoplasm.
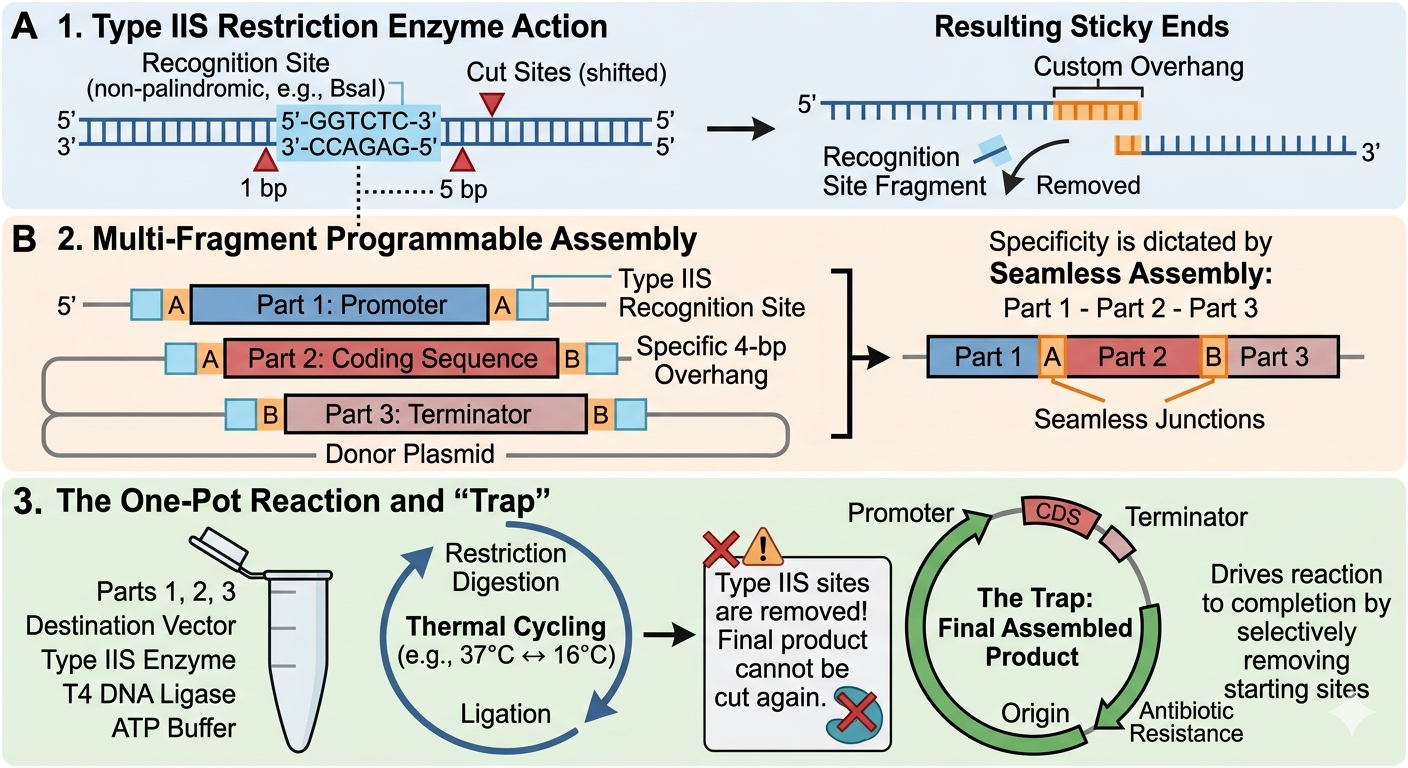
- Describe another assembly method in detail (such as Golden Gate Assembly)
Golden Gate Assembly (GGA) is a highly efficient molecular cloning method that allows for the simultaneous assembly of multiple DNA fragments into a single piece. GGA relies on Type IIS restriction enzymes. These enzymes recognize specific DNA sequences but cut at a defined distance outside of that sequence. This allows the user to design complementary overhangs with any 4-base-pair sequence on different fragments so that they are forced to assemble in a specific order and orientation. Researchers can also design this process to cut off and discard the recognition site. By having the restriction digest and ligation happen at the same time, the DNA is sealed within its final, correct form without the possibility of being cut again. Golden Gate assembly allows for the restriction digest and ligation to happen all in one tube with the highly efficient, seamless and programmable arrangement of multiple segments of DNA.