Week 1 HW: LungLite — Principles, Practices, and Governance
🌬️ Project Idea: LungLite (AI + Breath Microfluidics + Cell-Free Synbio)
1) Biological engineering application/tool + why
LungLite is a low-cost, noninvasive breath monitoring system that uses a microfluidic disposable cartridge.
The cartridge contains freeze-dried cell-free synthetic biology reactions to detect breath biomarkers associated with airway inflammation and oxidative stress.
A smartphone camera reads the cartridge’s color/fluorescence pattern and an AI model interprets the result.
The tool is intended to help users monitor lung health over time—especially people with asthma, COPD risk, and high pollution exposure—and provide early warning signals of inflammation before severe symptoms appear.
LungLite leverages cell-free synthetic biology to detect breath biomarkers safely and efficiently. Instead of using live engineered cells, it employs freeze-dried transcription-translation (TX-TL) systems with non-replicating DNA circuits that respond to molecules associated with airway inflammation and oxidative stress. When a user exhales into the microfluidic cartridge, these engineered circuits trigger colorimetric or fluorescent signals proportional to biomarker levels. The sealed cartridge design, combined with built-in post-reaction neutralization, ensures safety, while AI algorithms analyze the visual output to provide an accurate, real-time readout of lung health. This integration of synthetic biology, microfluidics, and AI enables a low-cost, noninvasive tool for continuous monitoring, especially in high-risk environments or populations with limited access to traditional respiratory diagnostics.
Why this matters:
Current lung monitoring tools like spirometers often require strong forced exhalation and are not always accessible, comfortable, or usable for children, elderly people, or individuals in low-resource settings.
This problem is also deeply personal to me because I grew up around severe air pollution in Delhi, where “bad air days” are normal and respiratory symptoms are common. LungLite is motivated by the idea that people in high-exposure environments should be able to track early signs of inflammation easily and affordably—before symptoms become severe.


Initially worked on an AI-powered diagnostic tool for lung cancer. During this opportunity, I pivoted the design to focus on the Present Idea: a low-cost, noninvasive breath test that uses a microfluidic cartridge to track early signs of lung inflammation.
LungLite goal:
breathe → cartridge reacts → phone reads

References-
Cell free systems:
- https://pmc.ncbi.nlm.nih.gov/articles/PMC11920963/
- https://www.mdpi.com/1422-0067/25/16/9109
- https://www.nature.com/articles/s41467-021-25233-y
- https://www.mdpi.com/1420-3049/29/8/1878
- Biochemical Preparation of Cell Extract for Cell-Free Protein Synthesis without Physical Disruption
DNA Circuits:
- https://www.mdpi.com/2079-4991/11/11/2955
- https://pmc.ncbi.nlm.nih.gov/articles/PMC10997190/
- https://www.science.org/doi/10.1126/sciadv.ady8165
- https://ietresearch.onlinelibrary.wiley.com/doi/full/10.1049/iet-nbt.2014.0056
2) Governance/policy goals for an ethical future
Because LungLite sits at the intersection of bioengineering + consumer health + AI, it raises issues in biosecurity, lab safety, privacy, equity, and responsible health claims.
The governance goal is to ensure LungLite contributes to an ethical future by preventing harm while promoting constructive public health benefits.
Policy Goal A — Enhance Biosecurity
- Sub-goal A1: Prevent incidents
Prevent misuse of cartridge biology (DNA templates, cell-free reagents) for harmful applications. - Sub-goal A2: Help respond
Ensure traceability and safe reporting if unsafe use or distribution occurs.
Policy Goal B — Foster Lab Safety
- Sub-goal B1: Prevent incidents
Ensure safe handling, manufacturing, and disposal of cartridges and reagents. - Sub-goal B2: Help respond
Ensure protocols exist for spills, exposure, or improper disposal.
Policy Goal C — Protect the Environment
- Sub-goal C1: Prevent incidents
Ensure cartridges and reagents do not introduce living organisms into waste streams. - Sub-goal C2: Help respond
Ensure recall, disposal, and remediation pathways if materials are found to persist or contaminate waste streams.
Policy Goal D — Other considerations
- Minimize costs and burdens to stakeholders
- Ensure feasibility for student prototyping and future scaling
- Do not unnecessarily impede legitimate research
- Promote constructive applications (public health monitoring, pollution health impacts)
3) Governance actions
Option 1: Technical Safety-by-Design
(Cell-free only + built-in kill chemistry + non-replicating DNA templates)
Idea
Many biosensors rely on living engineered organisms or wet reagents that could survive handling errors. LungLite instead commits to a cell-free-only architecture, using non-replicating DNA and post-reaction neutralization so the cartridge cannot become a biological propagation risk.
Design
Actors: student researchers, academic labs, cartridge designers, manufacturers.
Key elements:
- Use commercially available or lab-prepared TX-TL cell-free extract
- Use DNA templates without replication machinery
- Add nuclease or denaturing reagents in a sealed “waste chamber” that activates after the reaction
- Design the cartridge as a sealed unit so users cannot access wet reagents directly
- Provide clear disposal instructions (trash-safe, not drain)
- Include a QR code for standardized disposal instructions and recall notices
Assumptions
- Cell-free systems are safe enough for consumer-adjacent use
- DNA templates cannot be easily repurposed into harmful functions
- Cartridge sealing prevents tampering and accidental exposure
- Neutralization chemistry is robust across temperature/humidity variation
Risks of failure
- Users could physically open the cartridge, mishandle reagents, or bypass neutralization
- Poor sealing could cause leakage
- DNA templates could be shared and repurposed outside intended use
Risks of “success”
- Widespread adoption could normalize at-home “bio reaction kits” without safety literacy
- Overconfidence in “bio-safe” claims could reduce careful oversight and institutional review
Option 2: Distribution + Supply Chain Controls
(DNA sequence screening + controlled reagent distribution + batch traceability)
Purpose
Even if the platform is designed safely, misuse risk increases when synbio components are distributed widely. This option adds governance at the distribution layer, aiming to prevent malicious acquisition or repurposing of DNA templates and reagents.
Design
Actors: DNA synthesis companies, cartridge manufacturers, distributors, university procurement offices, and potentially regulators.
Key elements:
- DNA template sequences are screened using existing industry DNA synthesis screening norms
- Cartridges sold with batch numbers, manufacturer ID, and basic traceability
- Reagent supply chain restricted to verified vendors
Assumptions
- Screening reliably catches harmful sequences
- Vendors cooperate and screening is consistently implemented
- Traceability meaningfully deters malicious use
- Legitimate users will tolerate additional friction
Risks of failure
- DIY synthesis or black-market sources bypass screening
- Screening could generate false positives and slow benign development
- Increased cost and friction could reduce adoption in low-resource communities
Risks of “success”
- Centralization of power in a small number of vendors could limit open science
- Smaller labs, students, and global south researchers could be excluded due to cost and access barriers
- Overly broad screening could suppress legitimate respiratory health research
Option 3: Responsible Health Claims + Data Governance
(Limit medical claims + privacy-by-design + transparency)
Aim
Even if the biology is safe, LungLite could still cause harm through false reassurance, panic, biased AI outputs, or privacy breaches. This option focuses on preventing digital harms and misleading health interpretation.
Design
Actors: app developers, product companies, IRBs/ethics boards (if research), privacy regulators, public health agencies, and clinical collaborators.
Key elements:
- Position LungLite initially as wellness monitoring, not a medical diagnostic
- Focus on trend tracking rather than absolute disease classification
- Provide clear disclaimers (“not a diagnosis; seek medical care if symptoms worsen”)
- Use local-first processing: results computed on-device when possible
- Require informed consent for any cloud upload or model improvement
- Provide opt-out for data sharing
- Publish model limitations and performance across demographics
- Align product claims with existing regulatory distinctions between wellness tools and regulated diagnostic devices
Assumptions
- Users understand “monitoring” vs “diagnosis”
- Privacy measures meaningfully reduce harm
- AI transparency improves trust and responsible use
- The model will generalize across different phones, lighting, and populations
Risks of failure
- Users may treat outputs as diagnoses and delay care
- Data leaks could expose sensitive health data
- Model bias could cause false reassurance or false alarms in specific groups
- Smartphone hardware variability could distort readings
Risks of “success”
- A widely adopted breath-health dataset could become commercially valuable and exploited
- Insurers/employers/schools could pressure people to share breath scores (coercive screening)
- “Wellness” framing could still function as a de facto diagnostic
4) Scoring matrix (1 = best, 3 = worst; n/a allowed)
| Does the option: | Option 1 | Option 2 | Option 3 |
|---|---|---|---|
| Enhance Biosecurity | |||
| • By preventing incidents | 1 | 1 | 2 |
| • By helping respond | 2 | 1 | 2 |
| Foster Lab Safety | |||
| • By preventing incident | 1 | 2 | 2 |
| • By helping respond | 2 | 2 | 2 |
| Protect the environment | |||
| • By preventing incidents | 1 | 2 | 2 |
| • By helping respond | 2 | 2 | 2 |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 1 | 3 | 2 |
| • Feasibility? | 1 | 2 | 1 |
| • Not impede research | 1 | 3 | 1 |
| • Promote constructive applications | 1 | 2 | 1 |
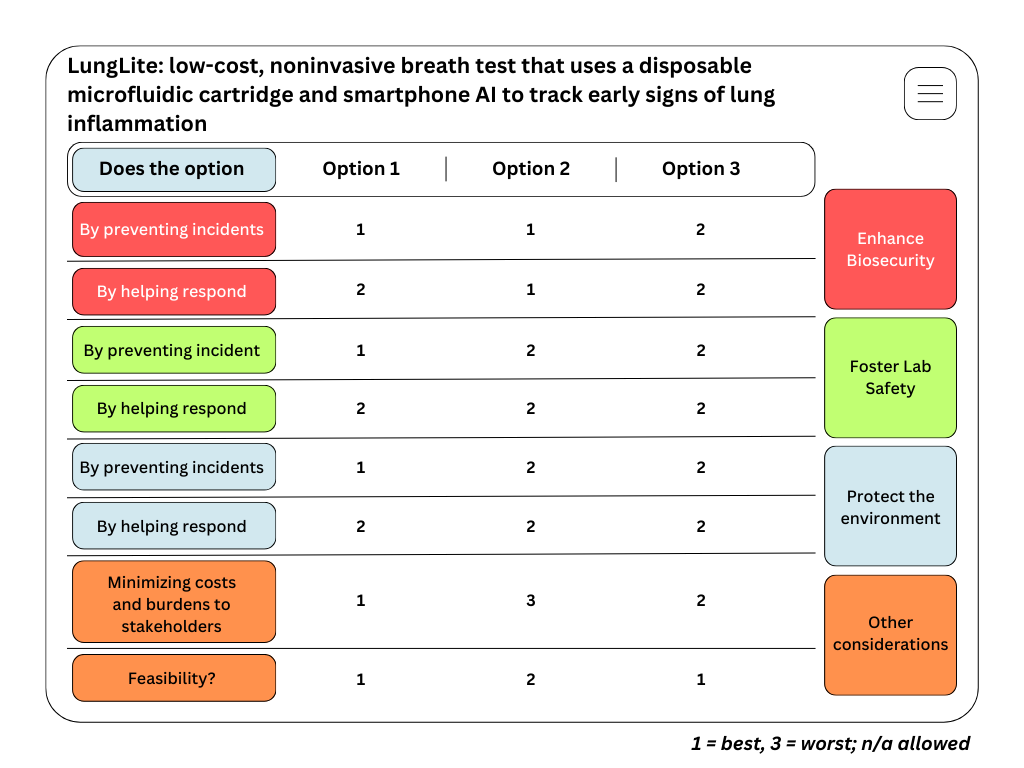

Figure: LungLite governance scoring matrix
Scoring justification:
- Option 1 reduces biological risk at the source and does not rely heavily on enforcement.
- Option 2 is strongest on biosecurity response, but worst on cost, equity, and research openness.
- Option 3 is strongest for AI/privacy harms but does not fully address upstream biosecurity.
There are few environmental concerns regarding this device like: packaging waste at scale, there might be low environmental risk regarding cell-free extracts, small risks associated with chemicals and dyes. Mitigation can be: minimal-material design, sealed leak-proof cartridge, and take-back/clinic disposal at scale.
5) Prioritized strategy
Recommended strategy
I believe we should prioritize Option 1 + Option 3 as the core approach now, and adopt a lightweight version of Option 2 only once scaling and commercialization begins.
Why Option 1 is essential
Option 1 addresses the biggest safety and biosecurity concern,i.e, distributing engineered biological systems into homes. By committing to cell-free synthetic biology only, LungLite becomes safer, easier to dispose of, and easier to govern ethically.
Why Option 3 is equally critical
Even if the biology is safe, LungLite can still cause harm through:
- false reassurance
- panic from false positives
- privacy breaches
- biased AI outputs
Option 3 reduces these risks through responsible messaging, careful AI design, and privacy-by-design.
Where Option 2 fits
Option 2 becomes more important once LungLite is manufactured at scale. Heavy supply chain restrictions too early could:
- block student prototyping
- increase costs
- reduce equitable access
- slow research innovation
So the staged approach is:
- Option 1 + Option 3 now
- Option 2 later (commercialization / mass distribution)
Tradeoffs considered
- Safety vs accessibility
- Innovation vs security
- User empowerment vs medical risk
- Privacy vs model improvement
Audience for recommendation
This governance strategy is best targeted at:
- MIT/university lab leadership
- future consumer product manufacturers
- public health agencies
- privacy regulators
6) What I Learned
Ethical concerns that arose
- Dual-use risk
- AI harm
- Privacy
- Equity
- Regulatory gray zone
- Coercion risk (monitoring becomes surveillance)
Governance actions proposed to address these
- Use cell-free systems only and avoid living organisms
- Seal cartridges and neutralize biological material post-test
- Implement privacy-by-design + local-first processing
- Avoid medical claims until clinically validated
- Keep manufacturing scalable and affordable
- Add anti-coercion safeguards (minimize retention, discourage third-party access)
Week 2 Lecture Prep
Homework Questions — Professor Jacobson
1) DNA polymerase error rate, genome comparison, and how biology handles the discrepancy
Nature’s machinery for copying DNA is DNA polymerase. High-fidelity replicative DNA polymerases (with proofreading) have an error rate of approximately:
~1 error per 1,000,000 to 10,000,000 base pairs
Comparison to the human genome
The human genome is approximately:
~3,200,000,000 base pairs
If replication relied only on polymerase accuracy:
At 1 error per 1,000,000 bp:
3,200,000,000 / 1,000,000 = 3,200 errors per genome replicationAt 1 error per 10,000,000 bp:
3,200,000,000 / 10,000,000 = 320 errors per genome replication
So even “high-fidelity” polymerase alone would still introduce hundreds to thousands of mistakes each time the genome is copied.
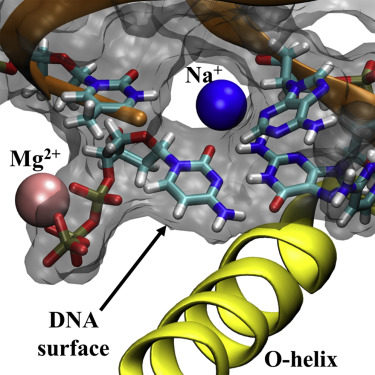

DNA polymerase’s shape precisely fits correct base pairs and uses a conformational “proofreading” motion to minimize misincorporation. https://www.sciencedirect.com/science/article/pii/S0969212615002695
How biology deals with the discrepancy
Biology reduces the final mutation rate using multiple layers of error correction:
- Polymerase proofreading removes many misincorporated bases during replication.
- Mismatch repair (MMR) fixes errors missed by proofreading.
- Base excision repair (BER) fixes chemically damaged bases.
- Nucleotide excision repair (NER) removes bulky lesions.
Together, these systems reduce the effective mutation rate to roughly:
~1 error per 1,000,000,000 to 10,000,000,000 bp per cell division
That means across one human genome replication, the final result is typically on the order of:
~0.3 to 3 mutations per cell division
🧪 Homework Questions — Dr. LeProust
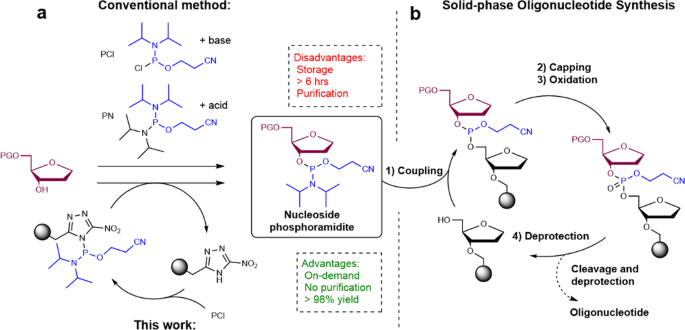
1) What’s the most commonly used method for oligo synthesis currently?
The most commonly used method is:
Solid-phase phosphoramidite DNA synthesis
This is the standard chemistry used by most commercial oligo suppliers. It works by building a DNA strand one nucleotide at a time on a solid support (like a bead, column, or array surface) using repeated cycles of:
- deprotection
- coupling
- capping
- oxidation
2) Why is it difficult to make oligos longer than ~200 nt via direct synthesis?
Direct synthesis becomes difficult past ~200 nucleotides because:
A) The yield drops exponentially with length
Each synthesis step has less than 100% efficiency, so errors compound as the oligo gets longer.Even in an optimistic scenario, most strands are truncated or incorrect.
B) Errors accumulate
Long oligos contain more:
- deletions (from incomplete coupling)
- substitutions (from incorrect incorporation)
- depurination damage (especially A/G under acidic conditions)
- truncated fragments
C) Purification becomes difficult and expensive
Separating a perfect 200-mer from 199-mer and 198-mer fragments is hard, so cost and complexity increase quickly.
3) Why can’t you make a 2000 bp gene via direct oligo synthesis?
Because the yield would collapse to essentially zero and the error rate would be unusable.
A) Yield becomes extremely low
B) The error rate becomes unacceptable
Even the rare full-length molecules would almost always contain:
- substitutions
- deletions
- truncations
- damaged bases
So you would not get a clean, correct 2000 bp product.
What is done instead in practice?
Instead of direct synthesis, genes are made by:
- synthesizing shorter oligos (usually 60–200 nt)
- assembling them into longer DNA using methods like:
- Gibson Assembly
- PCR-based assembly
- Golden Gate
- Ligase Cycling Assembly (LCA)
- then sequence-verifying clones to find a correct one
📄 HW by Dr. George Church — Grant Application (Devised)
Project Title
LungLite: A Room-Temperature, Breath-to-Color Microfluidic Cartridge Powered by Cell-Free Synthetic Biology and Smartphone AI for At-Home Lung Inflammation Monitoring
1) Abstract
Chronic respiratory disease affects hundreds of millions globally, yet lung health monitoring remains clinic-centered, effort-dependent, and inaccessible for many populations. Existing tools such as spirometry require strong forced exhalation and proper technique, while lab tests for inflammation and oxidative stress are expensive and slow.
I propose LungLite, a low-cost breath monitoring system that combines breath condensation microfluidics, freeze-dried cell-free synthetic biology, and smartphone computer vision + AI. Users breathe into a disposable cartridge that captures breath condensate and routes it through multiple reaction zones. Each zone contains a freeze-dried cell-free reaction that produces a colorimetric/fluorescent signal in response to oxidative stress and inflammation-associated breath chemistry.
A smartphone reader standardizes illumination, quantifies reaction outputs, and uses machine learning to interpret a multi-zone “fingerprint” into a trend score. LungLite is designed for safe, scalable, room-temperature storage and distribution and aims to enable daily lung health monitoring outside specialized medical centers.
2) Specific Aims
Aim 1 — Engineer a breath-to-fluid microfluidic cartridge
Hypothesis: A passive, low-cost cartridge can consistently convert breath into a defined liquid sample volume and deliver it to reaction zones with minimal variability.
Outcome: consistent fluid delivery across users and breathing conditions.
Aim 2 — Develop a multi-zone freeze-dried cell-free synbio sensing panel
Hypothesis: freeze-dried cell-free reactions can be stabilized at room temperature and produce reproducible outputs when rehydrated.
Outcome: 6–12 zone panel with internal controls and reproducible readouts.
Aim 3 — Build a smartphone reader + AI pipeline
Hypothesis: smartphone imaging + AI normalization improves reliability and interpretability.
Outcome: trend score + confidence + invalid-test detection.
3) Significance
LungLite could enable:
- noninvasive monitoring
- high-frequency measurement
- accessibility for children and low-resource settings
- room-temperature distribution
- population-level monitoring during wildfire smoke events
4) Innovation
- Cell-free synbio in a consumer cartridge
- Fingerprint sensing rather than single biomarker
- AI as a reliability layer (normalization + invalid detection + confidence)
5) Technical Approach and Work Plan (12 months)
- Months 1–2: breath capture + condensation
- Months 2–4: routing + zone array
- Months 3–7: freeze-dry stabilization
- Months 5–8: phone reader + illumination
- Months 7–10: AI training + invalid detection
- Months 10–12: validation + usability
6) Expected Deliverables
- disposable cartridge (6–12 zones)
- freeze-dried reaction panel + controls
- smartphone reader dock
- AI pipeline
- validation report
- product pathway plan
7) Risk Analysis and Mitigation
- biomarkers variable → fingerprint + controls + AI
- stability issues → sealed packaging + desiccant
- diagnostic misuse → wellness framing + disclaimers
- privacy misuse → local-first + opt-in + deletion
8) Safety, Ethics, and Governance Plan
- cell-free only
- sealed cartridges
- built-in neutralization
- sequence screening at synthesis
- traceability if scaling begins
- bias testing + transparency
- no disease claims until validated
9) Team and Resources
Cross-disciplinary team spanning:
- microfluidics
- cell-free synbio
- optics + computer vision
- ML
- product design
10) Long-Term Vision and Commercialization
- reusable reader + disposable cartridges
- room-temperature shipping
- low-cost manufacturing (paper microfluidics)
- Year 1: wellness monitoring
- Year 2+: clinical validation + regulated pathway
HW Review Papers — Week Summary Notes
1) DNA Sequencing at 40 (Shendure, J., Balasubramanian, S., Church, G. et al. https://doi.org/10.1038/nature24286)
Idea
DNA sequencing has gone through multiple revolutions and now functions as a universal molecular measurement tool — not just a way to read genomes.
Key points
- In ~40 years, sequencing scaled from kilobases → first human genome → millions of genomes
- Sequencing is no longer only for genomes; it is now used to measure:
- gene expression (RNA-seq)
- chromatin state (ATAC-seq, ChIP-seq)
- lineage tracing
- somatic mutations
- molecular interactions
- Costs dropped dramatically due to next-generation sequencing (NGS)
- Authors argue sequencing’s long-term impact may rival the microscope
Key message
We have become extremely good at reading DNA at massive scale, speed, and low cost.
2) DNA Synthesis Technologies to Close the Gene Writing Gap (2023), Hoose, A., Vellacott, R., Storch, M. et al. https://doi.org/10.1038/s41570-022-00456-9
Focus
We still cannot write DNA as efficiently as we can read it — and this is a major bottleneck for synthetic biology.
Key points
- Synthetic DNA is essential for:
- synthetic biology
- gene therapy
- DNA data storage
- nanotechnology
- Current chemical synthesis struggles beyond ~200 base pairs
- Long DNA synthesis is expensive and error-prone
- New approaches aiming to scale DNA writing include:
- enzymatic (template-independent) synthesis
- microarray-based synthesis + assembly
- rolling circle amplification
- molecular assembly + cloning pipelines
- As DNA writing becomes easier, regulation and oversight become more important
3) Recombineering and MAGE (2021), Wannier T, et al. Nat Rev Methods Primers, https://www.ncbi.nlm.nih.gov/pmc/articles/PMC9083505/
Core idea
Recombineering and MAGE enable precise, scarless, multiplex genome editing without requiring toxic double-strand breaks (DSBs).
Why traditional editing is limiting
Older editing methods (ZFNs, TALENs, CRISPR with DSBs):
- rely on double-strand breaks
- DSBs can be toxic (especially in bacteria)
- repair often produces unwanted indels
- low precision for large-scale combinatorial editing
Recombineering solution
- Uses phage proteins (Redβ, Exo, Gam)
- Introduces ssDNA or dsDNA with homology
- DNA integrates at the replication fork
- No DSB required
- Editing is highly precise and “scarless”
MAGE (Multiplex Automated Genome Engineering)
- Introduces many ssDNA oligos at once
- Creates combinatorial diversity across many genomic sites
- Enables genome-scale reverse genetics
4) CRISPR Technology: A Decade of Genome Editing is Only the Beginning, Wang, Doudna, et al., https://www.science.org/doi/10.1126/science.add8643
Focus area
CRISPR made genome editing programmable, accessible, and fast — dramatically lowering the barrier to entry.
Main points
- Cas9 + guide RNA enables targeting by base pairing
- Enabled:
- knockouts
- pooled genetic screens
- animal models
- crop editing
- emerging human therapies
Newer CRISPR-derived tools
- Base editing: A→G or C→T without DSBs
- Prime editing: templated edits with higher precision
Remaining challenges
- off-target effects
- delivery into cells/tissues
- limited multiplexing at large scale
- HDR inefficiency in many systems
Summary
Biotechnology has made DNA reading extremely scalable (sequencing), but DNA writing (synthesis) and DNA rewriting (editing) are still constrained by cost, accuracy, delivery, and scalability.
Sequencing is now a general-purpose measurement tool, while synthesis and editing are rapidly improving — raising both exciting capabilities and new governance needs.
I used artificial intelligence tools, including ChatGPT-5.0, for language refinement, structural organization, and clarity of expression in this documentation. All scientific concepts, design decisions, sequence selections, experimental reasoning, and technical interpretations reflect my own understanding and work. The AI tool was used solely to improve readability, coherence, and presentation quality.