Week 2 HW: DNA Read, Write, Edit — SOD1 Molecular Journey
🧬 Week 2 Documentation
DNA Read → DNA Write → DNA Edit
A Molecular Design Journey
This week was not just a technical exercise. It was an exploration — from abstract sequence to physical plasmid, from conceptual art to molecular execution. Below is the full documentation of my process, including failures, iterations, and insights gained.
🧪 Part 0: Basics of Gel Electrophoresis
Lectures + Recitation
I attended/watched all required lecture and recitation materials.
Conceptual Understanding
Gel electrophoresis separates DNA fragments based on size using:
- Negatively charged DNA backbone
- Electric field
- Agarose matrix
- Size-dependent migration
Smaller fragments travel further.
🎨 Part 1: Benchling & In-silico Gel Art
Step 1: Benchling Account + Lambda DNA Import
- Created Benchling account
- Imported Lambda DNA reference sequence
Step 2: Simulated Restriction Digestion
Enzymes used:
- EcoRI
- HindIII
- BamHI
- KpnI
- EcoRV
- SacI
- SalI
Initial Failure
My first digestion simulation produced fragmented bands that were too similar in size. The pattern looked visually indistinct.
Iteration Strategy
- Tested different single and double digests
- Compared fragment size outputs
- Adjusted enzyme combinations
Eventually, I selected combinations that produced strong band separation.
Kindly find attach all the simulations carried out for the same task:
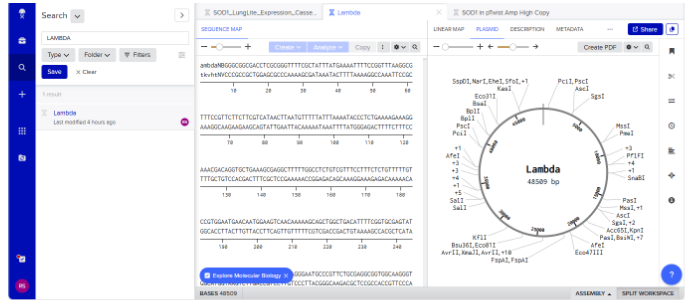
The following image represents setting up the Benchling account and loading lambda sequence, ultimately I was able to visualize as shown here-

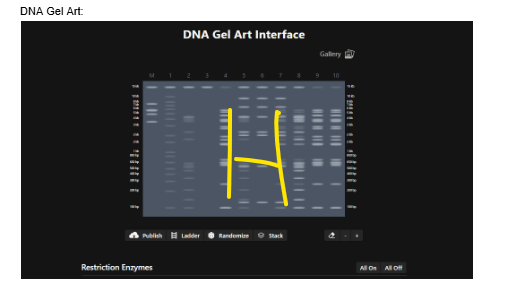
The following image shows the end result after carrying out the digestion process, I worked on a pattern design of “H Letter”, reason being my startup company’s first letter is H! Although, I must say I struggled alot and I intend to re run all of these simulations and tasks at least 5-6 times!


🧪 In-Silico Gel Art
I did try to work out on gel art, but yet again this part of the homework was something I really struggled.

Insight
Never had I imagined that biological mechanisms could generate such striking and beautiful art forms. As someone who once dreamed of becoming an artist but ultimately pursued engineering, I find this intersection deeply exciting. Working with gel patterns and molecular design has rekindled a childhood aspiration I once held close — the dream of opening an art studio.
🧬 Part 3: DNA Design Challenge
3.1 Choose Your Protein
Selected Protein: Human Superoxide Dismutase 1 (SOD1)
UniProt ID: P00441
Why SOD1?
SOD1 converts:
O₂⁻ → O₂ + H₂O₂
It protects against oxidative stress and is implicated in ALS.
It also integrates mechanistically with my LungLite platform — serving as a biochemical actuator.
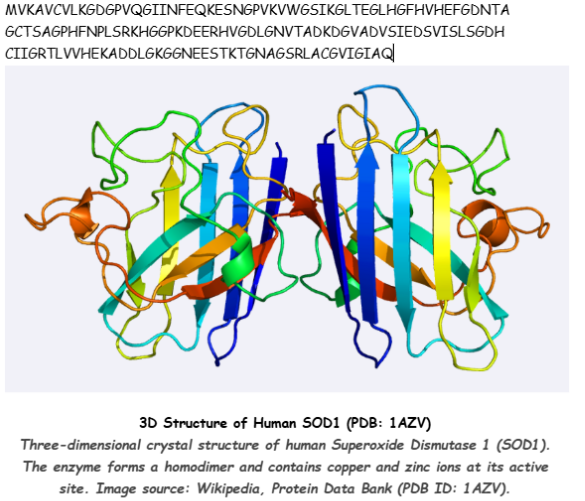

Amino Acid Sequence
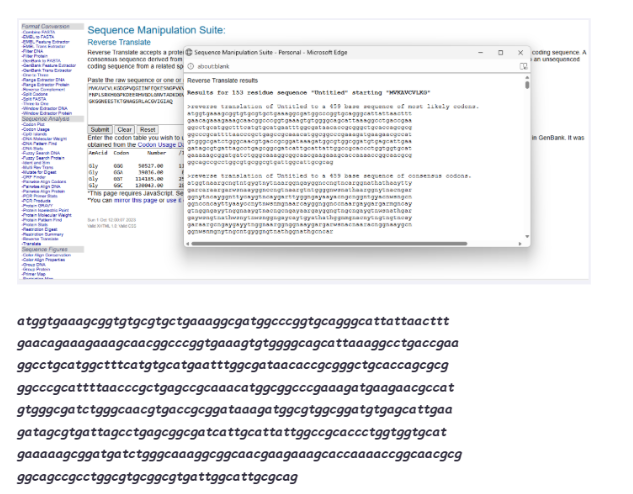
3.2 Reverse Translation
Using online reverse translation tools, I generated a nucleotide sequence.
Failure
Reverse translation produced multiple valid sequences due to codon degeneracy.
There is no single “correct” DNA sequence for a protein.
Resolution
I selected one biologically valid version as a starting template.
Pre-optimization DNA:


3.3 Codon Optimization
Why Optimize?
Different organisms prefer specific codons due to tRNA abundance.
Without optimization:
- Ribosome stalling
- Low yield
- Translation inefficiency
Host Chosen: Escherichia coli
Reasons:
- Fast growth
- High recombinant yield
- Standard lab organism
Final Codon Optimized Sequence

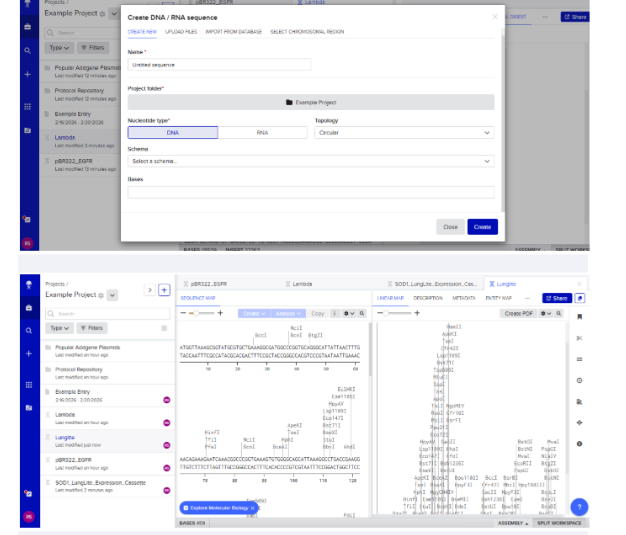

3.4 From DNA to Protein
Expression Methods:
Cell-Dependent
- Transform plasmid into E. coli
- Antibiotic selection
- Transcription
- Translation
- His-tag purification
Cell-Free Option
- TX-TL system
- Direct protein production without cells


3.5 Central Dogma Alignment
DNA:
RNA:
Protein:
Each 3 nucleotides = 1 amino acid
T → U during transcription
🧬 Part 4: Prepare a Twist DNA Synthesis Order
4.1 Accounts
- Created Twist account
- Created Benchling account
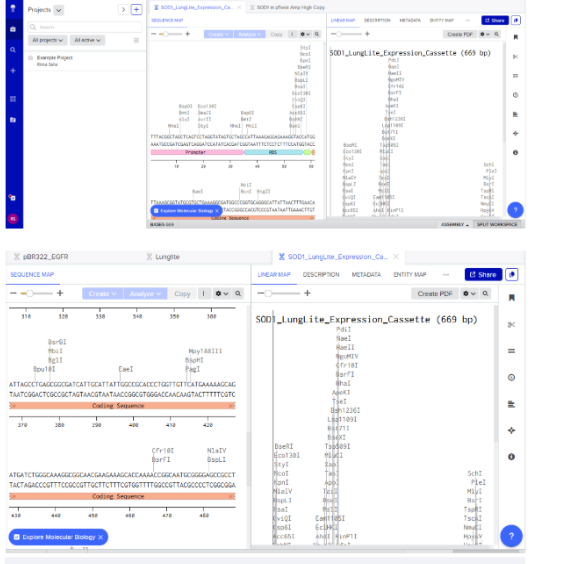
4.2 Build Expression Cassette
Structure:
Failure
Initially forgot to annotate regions in Benchling.
Fix
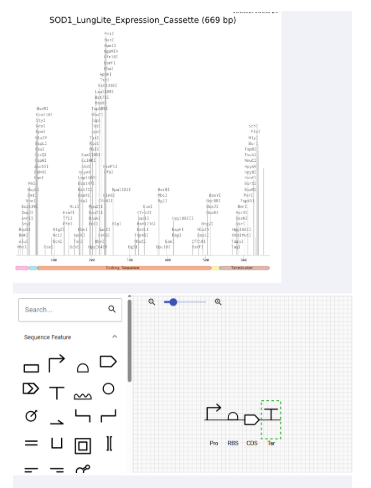
Annotated:
- Promoter
- RBS
- CDS
- His Tag
- Terminator
Verified via Linear Map view.
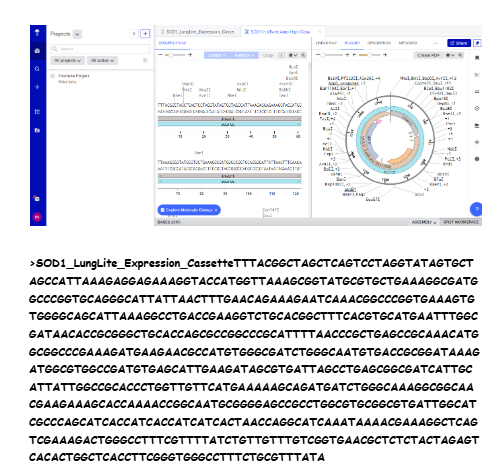
Final Insert Sequence
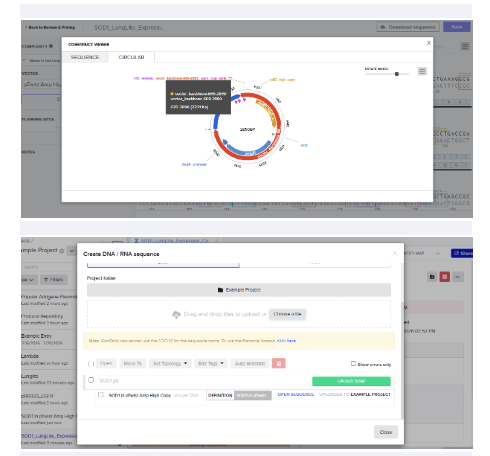
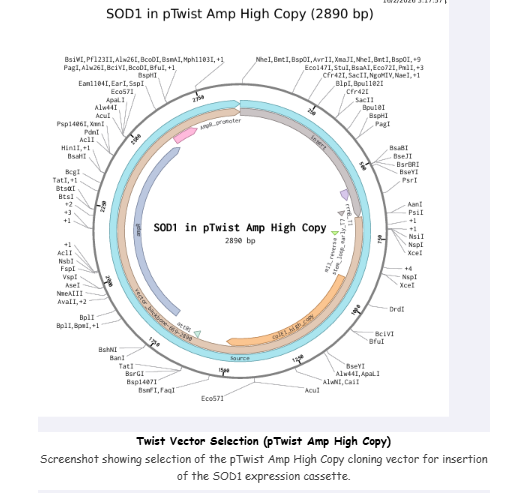
4.3–4.6 Twist Order
Selected:
- Genes → Clonal Genes
- Vector: pTwist Amp High Copy
Imported GenBank file back into Benchling to confirm construct.
I built my first plasmid.



🧬 Part 5: DNA Read / Write / Edit
5.1 DNA Read
What Would I Sequence?
The SOD1 gene sequence to understand its structure, variants, and oxidative stress relevance in lung epithelial biology.
Why This Matters
Superoxide Dismutase 1 (SOD1) is a cytosolic antioxidant enzyme that catalyzes the conversion of superoxide radicals (O₂⁻) into oxygen and hydrogen peroxide. Because oxidative stress is central to airway inflammation, SOD1 represents the molecular boundary between resilience and pathology in lung tissue. Mutations in SOD1 are linked to Amyotrophic Lateral Sclerosis (ALS), and its structure and function are well-characterized, making it ideal for recombinant engineering and diagnostic integration.
Technology Chosen: Oxford Nanopore
Generation: Third-generation sequencing
Input:
- Extracted DNA containing SOD1
- Adapter ligation
Mechanism:
- DNA passes through nanopores
- Ionic current changes → base calling
Output:
- FASTQ long reads of SOD1 sequence
Why Nanopore?
- Long reads allow full-length SOD1 sequencing
- Detects structural variants and potential regulatory regions
- Portable and scalable
Limitations:
- Higher error rate than Illumina
- Correctable with sequencing depth and consensus alignment
5.2 DNA Write
What Would I Synthesize?
A codon-optimized SOD1 expression cassette and ROS-responsive genetic circuits for LungLite.
Rationale
To integrate SOD1 into LungLite, the gene must be optimized for expression in bacterial or cell-free systems. This enables recombinant production and functional embedding into oxidative stress detection circuits.
Technology
- Phosphoramidite oligo synthesis
- PCR assembly
- Clonal gene insertion into expression vector
- 7×His tag for purification
Application in LungLite
Biological Amplifier Strategy
- ROS activates redox-sensitive promoter
- Induces SOD1 expression in freeze-dried TX–TL system
- SOD1 converts superoxide → H₂O₂
- Coupled colorimetric/fluorescent reaction produces smartphone-readable signal
Calibration Standard Strategy
- Purified recombinant SOD1 embedded in microfluidic wells
- Known concentrations normalize ROS dye response
- Enables quantitative oxidative stress scoring
Limitations
- Length constraints in synthesis
- Synthesis errors
- Cost scaling for large constructs
5.3 DNA Edit
What Would I Edit?
Upregulate antioxidant pathways — including SOD1 expression — in lung epithelial cells.
Technology: CRISPR-Cas9
Steps
- gRNA design targeting regulatory region
- Cas9-induced double-strand break
- HDR-mediated repair with enhanced promoter template
Input:
- gRNA plasmid
- Cas9
- Donor DNA template
- Target lung epithelial cells
Goal
Increase endogenous SOD1 buffering capacity to restore redox balance in oxidative stress conditions.
Limitations
- Off-target effects
- Variable editing efficiency
- Delivery challenges in airway epithelium
🌬 Final Reflection
What began as:
Lambda DNA
→ Restriction digest
→ Gel electrophoresis
Evolved into:
DNA Read → Sequencing SOD1
DNA Write → Engineering ROS-responsive SOD1 circuits
DNA Express → Recombinant protein production
DNA Integrate → Embedding SOD1 into LungLite microfluidic diagnostics
SOD1 is not merely a recombinant protein in this project. It becomes a functional biochemical actuator — translating environmental oxidative exposure into measurable signal output.
Growing up in Delhi, where severe air pollution makes oxidative stress a daily lived experience, reframes SOD1 from an abstract enzyme to a molecular proxy for environmental exposure. LungLite transforms this molecular logic into a portable, AI-integrated, noninvasive public health device.
The DNA Design Challenge is no longer just molecular cloning — it becomes the foundation for a programmable redox-sensing health platform.
I acknowledge that I used artificial intelligence tools, including ChatGPT-5.0, for language refinement, structural organization, and improvement of clarity in this documentation.
All scientific concepts, experimental designs, sequence selections, analytical reasoning, and technical interpretations presented in this work reflect my own understanding and independent effort. The AI tool was used solely to enhance readability, coherence, grammar, and overall presentation quality.
The prompts primarily included instructions such as: “Rewrite the text and correct grammatical errors.”