Week 11 — Bioproduction & Cloud Labs
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork


- For this project, my contribution was small—I added a dot to the artwork.
- What I liked most about the project was seeing how everyone’s individual contributions came together to form a larger, more complex design. It showed how even small inputs can matter when working as a community, and it was interesting to see the diversity of ideas and styles in one shared piece.
- For next year, the project could be improved by giving clearer guidance or structure so participants can better understand how their contributions will fit into the final design. It might also be helpful to have a more interactive element or planning stage so people can collaborate more directly rather than working in isolation.
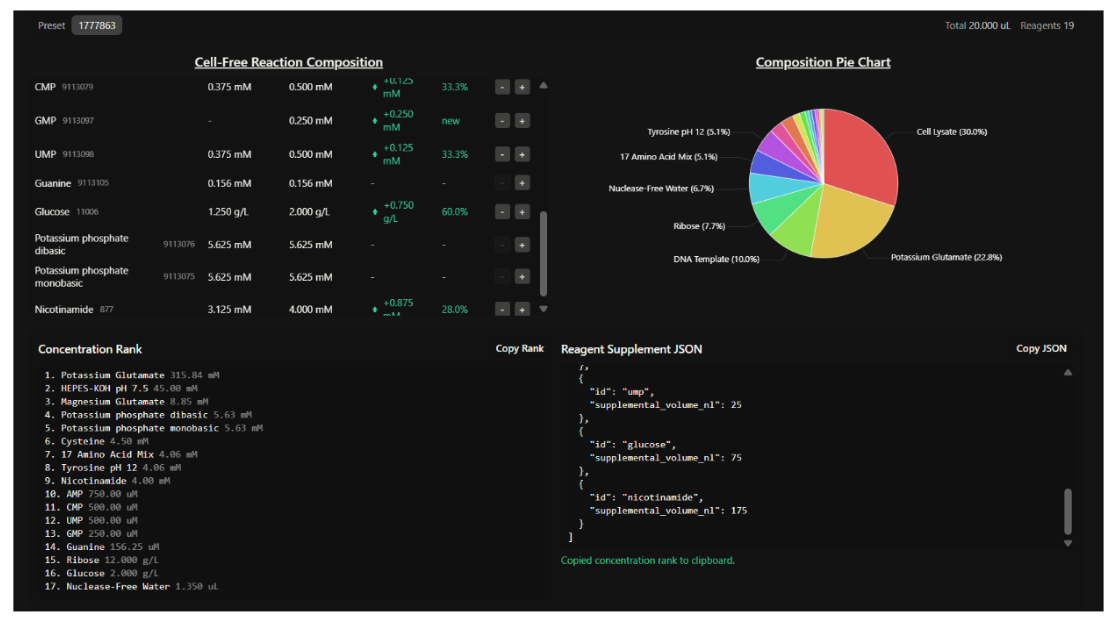
Part B: Cell-Free Protein Synthesis | Reagent Roles
E. coli Lysate: BL21 (DE3) Star Lysate (includes T7 RNA Polymerase) This lysate provides the core molecular machinery for transcription and translation, including ribosomes, tRNAs, aminoacyl-tRNA synthetases, and metabolic enzymes. The built-in T7 RNA polymerase enables strong transcription from T7 promoters on the DNA template.
Salts / Buffer
Potassium Glutamate Maintains intracellular-like ionic strength and stabilizes ribosomes and enzymes, improving protein synthesis efficiency. HEPES-KOH pH 7.5 Acts as a buffering agent to maintain stable pH, which is critical for enzyme activity and protein folding. Magnesium Glutamate Provides Mg²⁺ ions, which are essential cofactors for ribosome structure, ATP utilization, and RNA polymerase activity.
Potassium Phosphate Monobasic / Dibasic Together form a phosphate buffer system that helps maintain pH and provides phosphate for metabolic and nucleotide-related reactions.
Energy / Nucleotide System
Ribose Serves as a precursor for nucleotide synthesis, enabling regeneration of nucleotides over long reactions.
Glucose Acts as a slow-release energy source via glycolysis-like pathways in the lysate, sustaining ATP production.
AMP, CMP, GMP, UMP These nucleotide monophosphates are precursors that can be converted into triphosphates (ATP, CTP, GTP, UTP) required for transcription and energy transfer.
Guanine A nucleobase that can be salvaged into GMP and eventually GTP, supporting transcription even if GMP is limited.
Translation Mix (Amino Acids)
17 Amino Acid Mix Provides most amino acids required for protein synthesis, ensuring ribosomes can elongate polypeptides.
Tyrosine & Cysteine Added separately because they are prone to degradation or oxidation; cysteine is especially sensitive and important for disulfide bond formation.
Additives
Nicotinamide Supports redox balance by contributing to NAD⁺/NADH metabolism, which is important for sustaining metabolic activity in long reactions.
Backfill
Nuclease-Free Water Used to adjust final reaction volume without introducing nucleases that could degrade DNA or RNA.
Differences Between Master Mixes The 1-hour PEP-NTP system uses phosphoenolpyruvate (PEP) as a high-energy phosphate donor and directly supplies nucleotide triphosphates (NTPs), enabling rapid and high initial protein production but with quick energy depletion.
The 20-hour NMP-Ribose-Glucose system relies on slower metabolic regeneration of energy and nucleotides from nucleoside monophosphates, ribose, and glucose.
This leads to lower initial rates but much longer-lasting protein synthesis.
- Why Transcription Works Without GMP Even without added GMP, transcription can proceed because guanine can be salvaged into GMP through enzymatic pathways in the lysate. This GMP is then phosphorylated into GTP, which RNA polymerase uses for RNA synthesis.
Fluorescent Proteins Properties
sfGFP (superfolder GFP): sfGFP folds very efficiently and rapidly, even under suboptimal conditions, making it highly robust in cell-free systems. Its fast maturation leads to strong early fluorescence signals.
mRFP1: mRFP1 has slower chromophore maturation and less efficient folding compared to GFP variants, which can delay fluorescence onset in cell-free reactions.
mKO2: mKO2 matures relatively quickly but is somewhat sensitive to environmental conditions like pH, which can affect fluorescence intensity.
mTurquoise2: This cyan fluorescent protein has very high quantum yield but requires precise folding and is sensitive to oxidative conditions, impacting brightness.
mScarlet_I: mScarlet-I is a bright red protein with improved maturation compared to older RFPs, but still slower than GFP variants and dependent on proper oxygen availability.
Electra2: Electra2 (a newer engineered protein) is optimized for brightness but may require specific folding or redox conditions, making its performance sensitive to reaction composition.
Hypothesis for Optimization
- Protein: mScarlet-I
- Reagents to adjust: Increase oxygen availability (e.g., reduce reaction volume or increase surface area) and optimize magnesium concentration.
- Expected Effect: Improved chromophore maturation (which is oxygen-dependent) and enhanced ribosome activity will increase correctly folded protein, leading to higher fluorescence over 36 hours.
- Protein: mTurquoise2
- Reagents: Add nicotinamide and optimize redox balance
- Effect: Improved folding environment and reduced oxidative stress will enhance fluorescence intensity.
- To maximize fluorescence over 36 hours:
- Use glucose + ribose system for sustained energy
- Optimize Mg²⁺ concentration for translation efficiency
- Adjust amino acid balance, especially cysteine
- Maintain stable pH buffering
Part C: Final Cell-Free Master Mix Design (sfGFP)
- Reaction (20 μL total)
- 6 μL Cell Lysate
- 10 μL 2X Optimized Master Mix (sfGFP preset)
- 2 μL DNA Template (sfGFP)
- 2 μL Custom Reagent Supplement
- This composition supports long-duration (20–36 hr) expression using a ribose–glucose energy system.
- Key Features of sfGFP Master Mix
- High potassium glutamate (~313 mM) → mimics intracellular conditions, stabilizes ribosomes
- Balanced Mg²⁺ (~7 mM) → supports translation and proper folding
- Ribose + glucose system → enables sustained ATP regeneration over long incubation
- Complete amino acid mix + cysteine/tyrosine supplementation → prevents bottlenecks in translation
- Nicotinamide (3.125 mM) → supports redox balance for long reactions
This is ideal for sfGFP, which benefits from:
- fast folding
- high robustness
- efficient translation
- sfGFP-Specific Biophysical Considerations
- sfGFP properties affecting expression:
- Extremely fast folding (superfolder variant)
- High tolerance to ionic and environmental variation
- Oxygen-independent chromophore formation (mostly robust)
- Implication:
- sfGFP is translation-limited, not folding-limited, so improving:
- ribosome efficiency
- energy availability → increases fluorescence output.
Reaction Setup (unchanged)
- Cell Lysate → 6.000 μL
- DNA Template → 2.000 μL
- Master Mix → 10.000 μL
- Custom Supplement → 2.000 μL
MASTER MIX FINAL TARGET CONCENTRATIONS Set reagents at:
- Core Ions & Buffer
- Potassium Glutamate → 315 mM ⬆ (increase slightly from 312.56)
- Magnesium Glutamate → 8.5 mM ⬆
- HEPES-KOH (pH 7.5) → 45 mM (kept same)
- Potassium phosphate (mono + dibasic) → 5.6 mM each (kept same)
Amino Acids
- 17 AA Mix → 4.1 mM (kept same)
- Tyrosine → 4.1 mM (kept same)
- Cysteine → 4.5 mM ⬆ (slight increase improves stability over time)
- Energy System (KEY FOR 36h)
- Ribose → 12 g/L ⬆ (small boost for nucleotide regeneration)
- Glucose → 2.0 g/L ⬆⬆ (VERY IMPORTANT for long reactions)
Nucleotides
- AMP → 0.75 mM ⬆
- CMP → 0.5 mM ⬆
- UMP → 0.5 mM ⬆
- Guanine → 0.2 mM ⬆
- GMP → leave OUT
Additives
- Nicotinamide → 4.0 mM ⬆ (improves long-term metabolic stability)
More Details about the master mix-
- Magnesium Increase
- Boosts ribosome activity
- Increases translation rate
- sfGFP tolerates higher Mg²⁺ well
This alone can significantly increase yield
- Glucose Increase
- Extends ATP production beyond 20 hours
- Prevents early reaction collapse
- Critical for 36-hour fluorescence
- Slight Potassium Increase
- Improves ribosome stability
- Enhances protein synthesis efficiency
- Cysteine + Nicotinamide Boost
- Protects against oxidation
- Maintains enzyme activity long-term
- Nucleotide Increase
- Prevents transcription bottlenecks over time
- Increasing magnesium glutamate and glucose concentrations will enhance ribosomal activity and extend energy availability, respectively. Because sfGFP folds efficiently, improving translation rate and reaction longevity will directly increase total protein production, resulting in higher fluorescence over a 36-hour incubation.
EXPECTED RESULT
- Faster fluorescence onset
- Higher peak fluorescence
- Longer sustained signal
- Better total yield
- Potassium Glutamate 315.84 mM
- HEPES-KOH pH 7.5 45.00 mM
- Magnesium Glutamate 8.85 mM
- Potassium phosphate dibasic 5.63 mM
- Potassium phosphate monobasic 5.63 mM
- Cysteine 4.50 mM
- 17 Amino Acid Mix 4.06 mM
- Tyrosine pH 12 4.06 mM
- Nicotinamide 4.00 mM
- AMP 750.00 uM
- CMP 500.00 uM
- UMP 500.00 uM
- GMP 250.00 uM
- Guanine 156.25 uM
- Ribose 12.000 g/L
- Glucose 2.000 g/L
- Nuclease-Free Water 1.350 uL