Ritika Saha — HTGAA Spring 2026


About me
Hello! I’m Ritika Saha, a student in HTGAA (Spring 2026).
My interests include:
- 🧬 Synthetic biology + diagnostics
- 🤖 Responsible AI for health
Contact info
Let’s connect:
HTGAA Committed Listener (CL) Agreement
I am a HTGAA Committed Listener, my responsibilities are:
- Watching class lectures and recitations
- Participating in node reviews Developing and documenting my homework
- Actively communicating with other students and TAs on the forum
- Allowing HTGAA and BioClub to share my work (with attribution) Honestly reporting on my work, and appropriately attributing and citing the work of others (both human and non-human)
- Following locally applicable health and safety guidance
- Promoting a respectful environment free of harassment and discrimination
- Signed by committing this file to my documentation page/repository,
Ritika Saha 9 March 2026
Homework
- Principles, practices, and governance for the LungLite concept. Along with week 2 lecture prep
- A documented journey through gel electrophoresis, SOD1 DNA design, codon optimization, plasmid construction, and DNA read/write/edit technologies.
- Opentrons scripting, lab automation exploration, and final project ideation.
- This week focuses on how sequence, structure, and energetics can be modeled and manipulated to create or optimize proteins with specified functions.
- This week we learn how cutting-edge AI and protein language models are used to design functional proteins and peptides “in silico”.
- This week we learn core molecular biology tools and techniques for processing and assembling DNA, including PCR and Gibson Assembly.
- This week covers neuromorphic genetic circuits, showing how engineered gene networks can implement neural-network “perceptron”-like computation and learning.
- This week introduces synthesis of proteins using cellular machinery outside of a cell.
- This lecture presents a range of advanced technologies to do precision measurement of proteins at atomic scales, characterizing chemical composition, and detecting protein sequence and structure.
- Cloud Labs and Cell free extract
Labs
I will also share how I adapt lab work to a home setup and translate those workflows into scalable lab or office environments.
Projects
- Initially worked upon three different ideas: Idea 1 Breathe based diagnositc device Idea 2 Digital Cell Twin Modeling for Cancer and Oncology Virtual Cell Hypothesis Generation Idea 3 Decoding the genetic circuitry of lung cancer cells Later finalized to go with idea number one i.e Real time diagnostic system for lung health monitoring.
- Group Formed Proposal: https://docs.google.com/document/d/1ENvPHhRbBgtl0ERrfqmomJKxPg68nfvCugrPQrDdM7o/edit?tab=t.0 Documentation: https://pages.htgaa.org/2026a/ritika-saha/homework/week-05-hw-protein-design-part-ii/index.html By: 2026a-nourelden-rihan, 2026a-ritika-saha, 2026a-rahul-yaji, 2026a-keerthana-gunaretnam We decided to focus on the main area of increasing the stability of the MS2 phage lysis protein L, with a possible secondary goal of reducing the dependency on host DnaJ, while still maintaining the lysis action. The tools AlphaFold, Clustal Omega, BLAST, ESM, and ESMFold were discussed. BLAST can pull out homologous lysis proteins from the databases. Clustal Omega can create MSAs to identify essential L48-S49 residues, and the pore-forming regions that must not be mutated. ESM can create mutation heatmaps, which can guide the use of ESMFold to obtain highest score foldings in mutatable regions. AlphaFold Multimer predicts whether the subunits of our protein can successfully create a pore in the host membrane, and also to check whether N-terminus can break the interaction with DnaJ. We also identified a few pitfalls, with majors ones dealing with limited training datasets, that may not be properly aligned towards creating a transmembrane lysis protein. Some other pitfalls include the lack of proper annotations for amurins; the possibility of an over-stable protein to form non-functional aggregates; and the vulnerability of modified protein to host proteases.
Proposed Idea
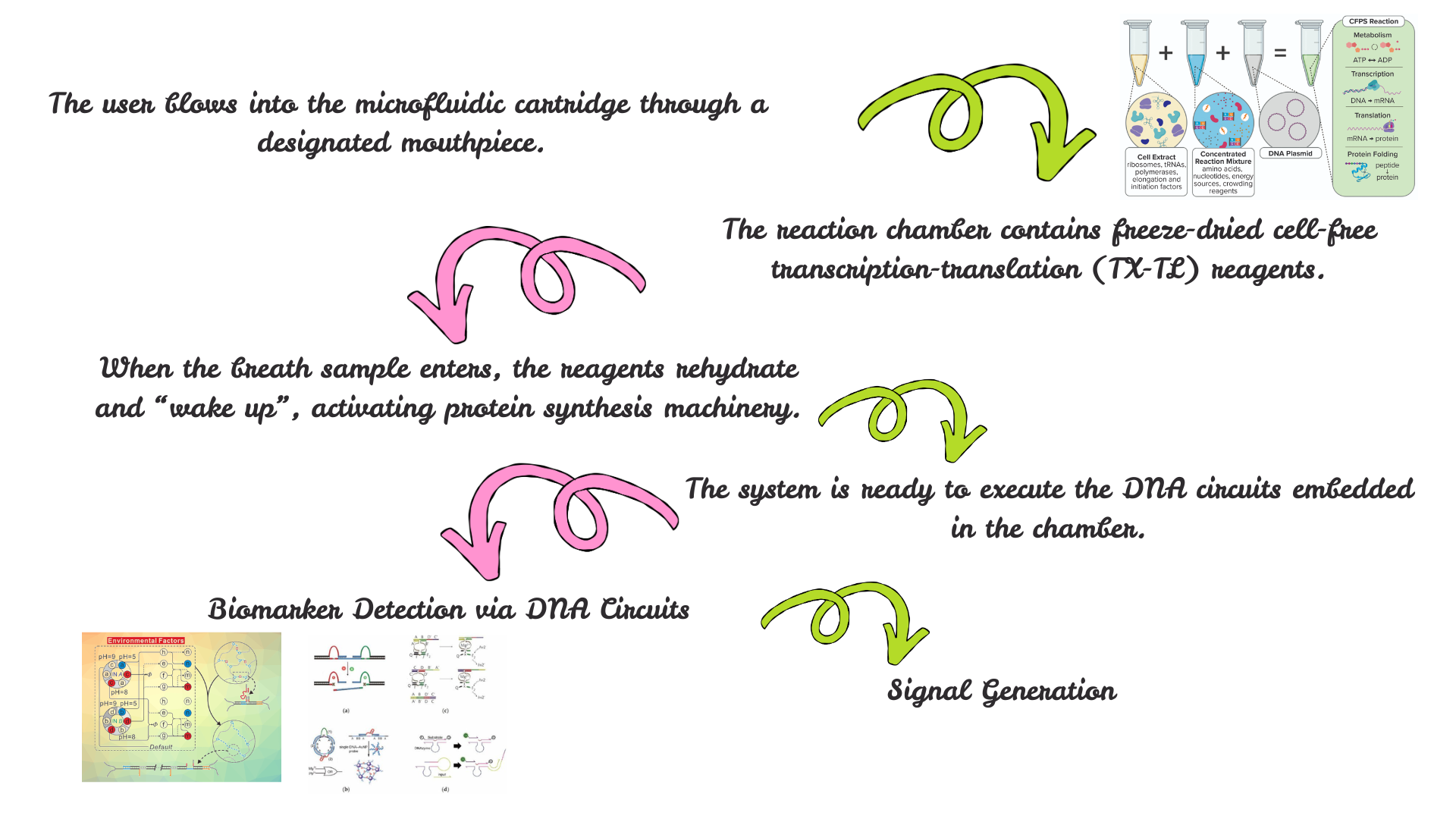

I am exploring a project at the intersection of synthetic biology, diagnostics, and responsible AI.
The goal is to design systems that:
- Enable low-cost, rapid biological diagnostics
- Integrate AI responsibly into healthcare workflows
- Improve accessibility of advanced diagnostics in resource-limited settings
This section will evolve as the idea matures through the course.
Follow My Journey
I document my learning, experiments, and reflections here:
- ✍️ Substack: Substack Link Would Be Added Here!