Individual Final Project
Initially worked upon three different ideas:
Idea 1 Breathe based diagnositc device
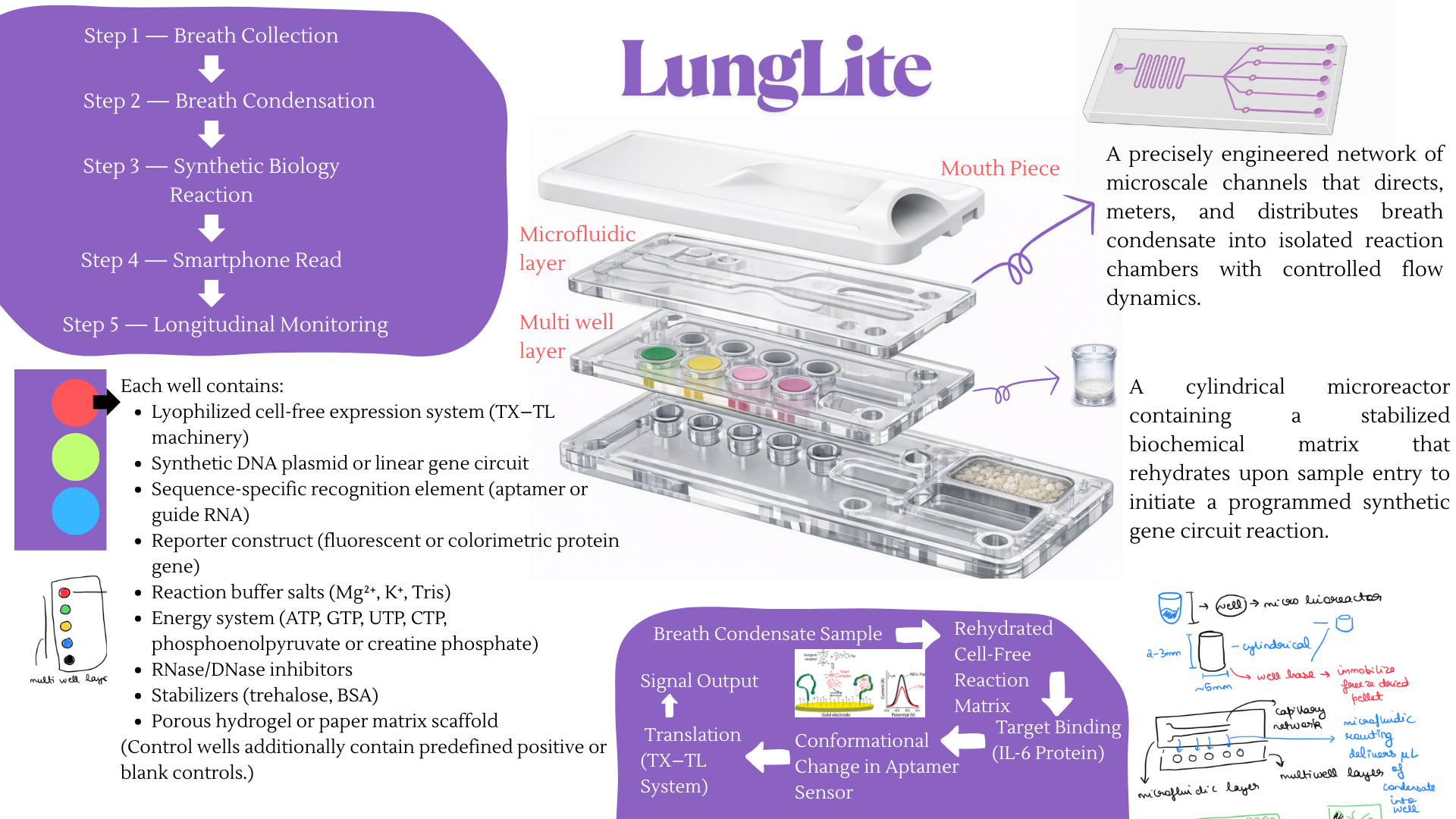

Idea 2 Digital Cell Twin Modeling for Cancer and Oncology Virtual Cell Hypothesis Generation
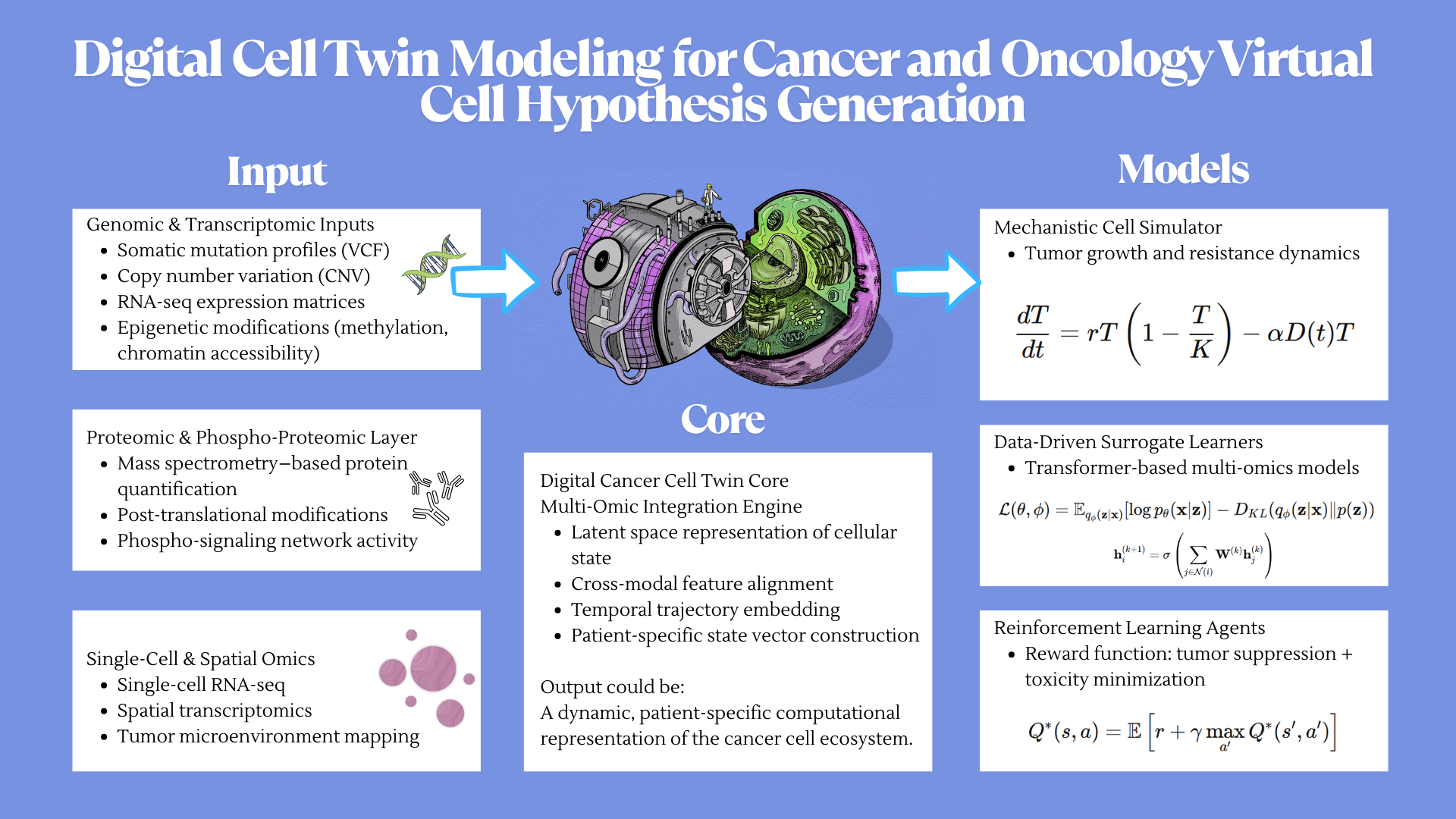

Idea 3 Decoding the genetic circuitry of lung cancer cells
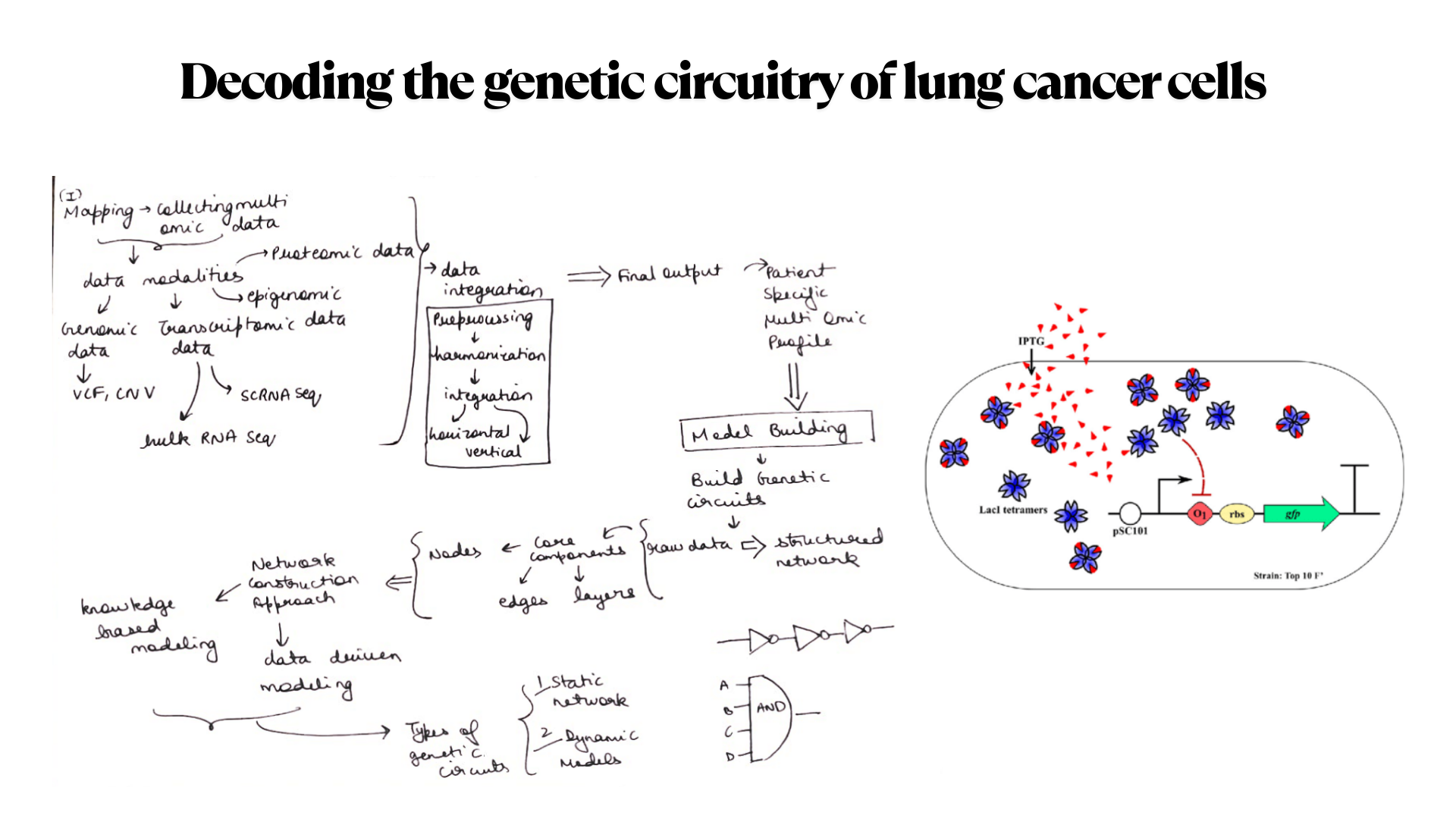

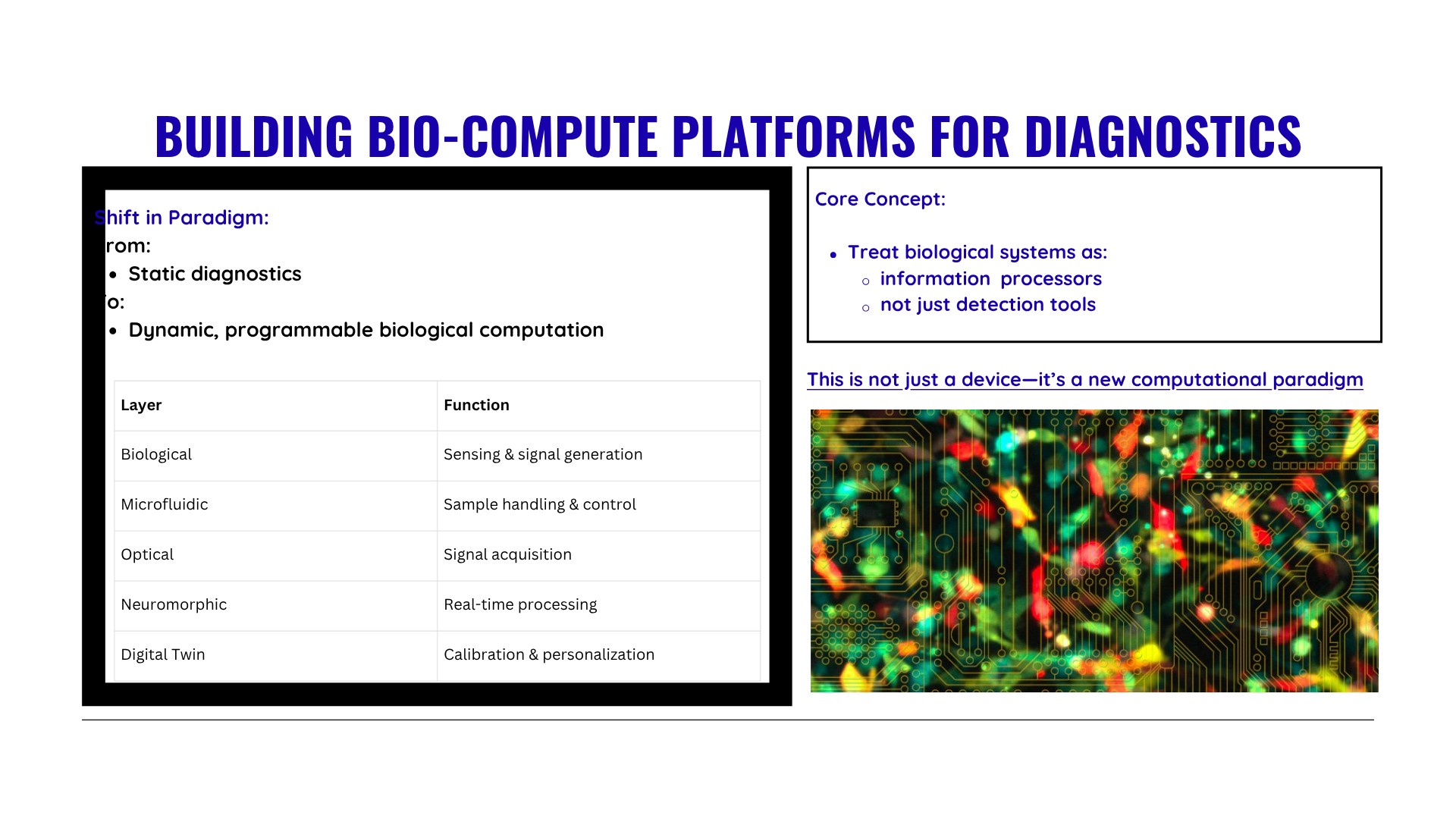
Later finalized to go with idea number one i.e Real time diagnostic system for lung health monitoring.


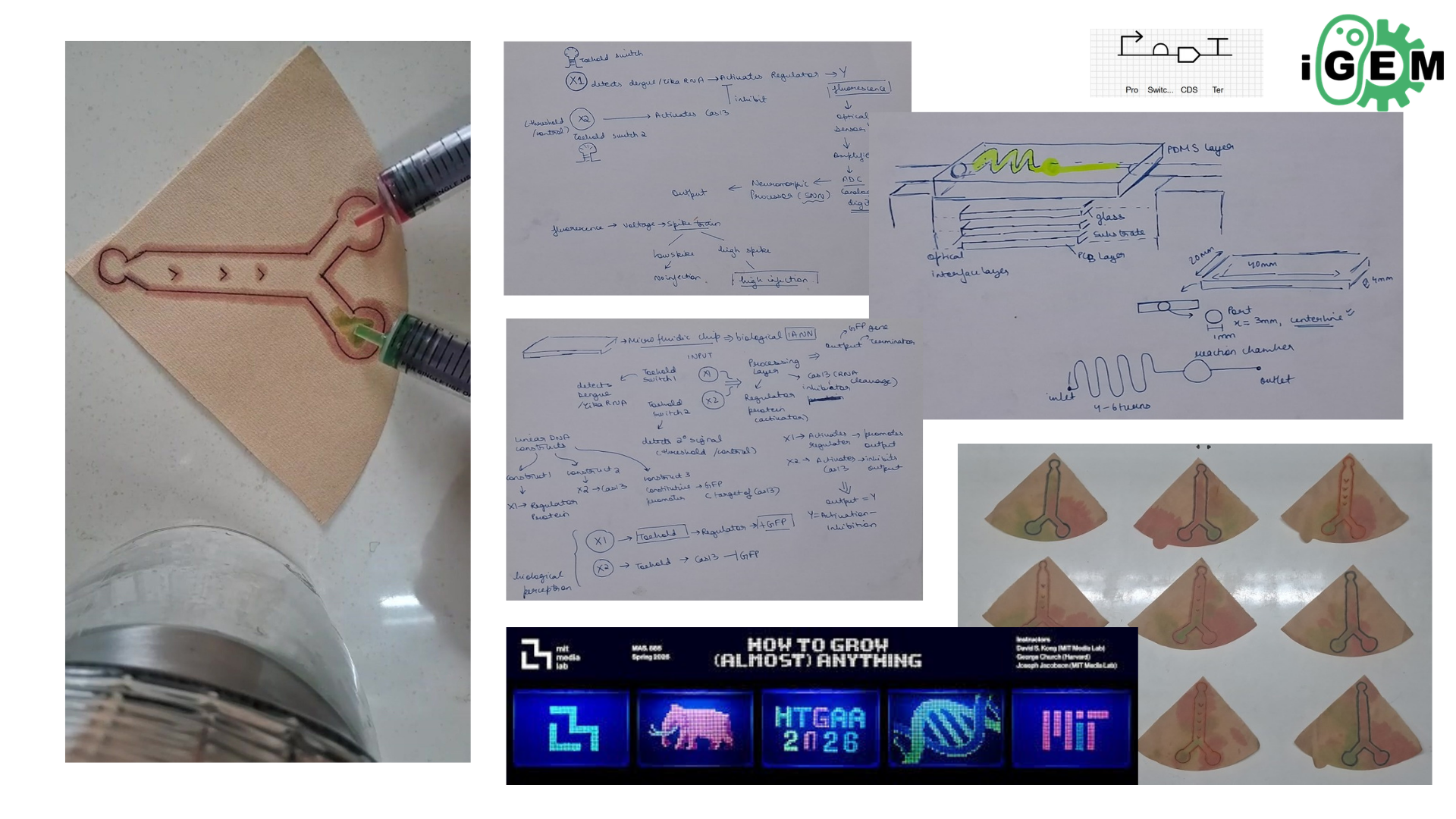



Benchling Link for twist order: https://benchling.com/reet123/f_/DvufGAFHIG-final-project-construct/
SECTION 1: ABSTRACT
Respiratory diseases and systemic inflammation are often diagnosed only after symptoms become severe, limiting opportunities for early intervention. This project addresses the need for a real-time, non-invasive diagnostic platform capable of continuously monitoring key biomarkers in breath condensate or saliva. The overall goal is to develop a microfluidic, cell-free biosensing system that integrates synthetic biology with computational signal processing to enable early disease detection.
The central hypothesis is that combining optimized cell-free gene expression systems with biomarker-specific genetic circuits and computational signal interpretation will enable sensitive, real-time detection of disease-relevant molecules. The project focuses on three biomarkers: IL-6 (inflammation), viral/host RNA (infection), and hydrogen peroxide (oxidative stress). Specific aims include designing a microfluidic device with independent reaction chambers, optimizing cell-free reactions to maximize fluorescence output, and developing a neuromorphic signal processing framework calibrated with a digital twin model.
Methods include DNA construct design (T7-driven sfGFP reporter with aptamer regulation), cell-free transcription-translation (TX-TL) optimization, microfluidic integration, and fluorescence-to-signal conversion. The expected outcome is a scalable, portable diagnostic system capable of continuous health monitoring, with potential applications in early disease detection, personalized medicine, and low-resource healthcare settings.
SECTION 2: PROJECT AIMS
- Aim 1: Experimental Aim
The first aim of my final project is to design and validate a microfluidic device that enables controlled entry of breath condensate or saliva samples into three independent reaction wells, each containing a freeze-dried cell-free gene expression system engineered with specific genetic circuits to detect IL-6, viral/host RNA, and hydrogen peroxide, producing distinct fluorescence outputs measurable via an integrated optical sensing layer.
- Aim 2: Development Aim
The second aim is to develop an integrated signal processing framework that converts fluorescence-derived optical signals into neuromorphic spike trains and calibrates them using a digital twin model based on virtual cell simulations, improving sensitivity, specificity, and robustness to sampling variability.
- Aim 3: Visionary Aim
The third aim is to establish a fully integrated, non-invasive diagnostic platform that combines synthetic biology, microfluidics, and neuromorphic computing to classify individuals into health risk categories in real time, enabling continuous and personalized monitoring of respiratory and systemic health.
SECTION 3: BACKGROUND
Literature Context Cell-free systems have become powerful tools for diagnostics due to their programmability and portability. Several studies have demonstrated that freeze-dried TX-TL systems can detect viral RNA and environmental signals outside of laboratory settings. Additionally, certain studies have shown that optimizing energy systems (e.g., glucose and ribose) significantly improves protein yield and reaction duration in cell-free systems. Despite these advances, current systems often lack long-term stability, multiplexing capability, and integration with computational frameworks. This project addresses these limitations by combining multi-biomarker detection, optimized reaction chemistry, and real-time signal processing.
Innovation
This project is innovative because it integrates:
- Microfluidics + cell-free biosensing + neuromorphic computing
- Multiplexed detection of multiple biomarkers in parallel
- Biochemical optimization (Mg²⁺, glucose, nucleotides) for long-duration expression
- Additionally, the use of a digital twin model to interpret biological signals introduces a novel interface between synthetic biology and computational modeling.
- Impact
This project targets the major challenge of early detection of respiratory and systemic diseases. Current diagnostics are often invasive and episodic, missing dynamic changes in patient health. By enabling continuous monitoring, this system could transform healthcare toward preventive and personalized medicine.
The platform could be deployed in low-resource settings due to its portability and low cost, improving global health equity. It also reduces reliance on centralized laboratories and enables rapid response to infectious disease outbreaks. Scientifically, this work advances synthetic biology by demonstrating how biochemical tuning and computational integration can enhance system performance.
- Ethical Implications
This project raises ethical considerations related to data privacy, accessibility, and responsible deployment of diagnostic technologies. The principle of beneficence applies, as the system aims to improve early detection and health outcomes. However, justice must be ensured so that such technologies are accessible across socioeconomic groups and do not exacerbate healthcare disparities. To ensure ethical implementation, safeguards must be established for data security and informed consent, especially when continuous monitoring is involved. Potential unintended consequences include overdiagnosis or anxiety due to continuous health tracking. To mitigate this, the system should be used as a decision-support tool rather than a standalone diagnostic, and results should be interpreted alongside clinical expertise. Regulatory oversight and transparent validation are essential to ensure safety and reliability.
SECTION 4: EXPERIMENTAL DESIGN
- DNA Construct (Benchling Design)
- T7 Promoter
- 5′ UTR with aptamer-based regulatory element
- Ribosome Binding Site (RBS)
- sfGFP reporter gene
- Transcription terminator
This design enables biomarker-responsive translation control, where the aptamer regulates expression based on target molecules.
- Cell-Free Reaction Design (Optimized)
Final Reaction Composition (20 μL)
- 6 μL Lysate
- 10 μL 2X Master Mix
- 2 μL DNA template
- 2 μL Custom supplement
Final Optimized Concentrations
- Potassium glutamate: 315.84 mM
- Magnesium glutamate: 8.85 mM
- HEPES: 45 mM
- Cysteine: 4.5 mM
- Nicotinamide: 4.0 mM
- AMP: 0.75 mM
- CMP/UMP: 0.5 mM
- GMP: 0.25 mM
- Ribose: 12 g/L
- Glucose: 2 g/L
Step-by-Step Experimental Plan
- Design DNA constructs for sfGFP and biomarker-responsive circuits
- Order DNA via Twist Bioscience
- Prepare or obtain BL21 cell lysate
- Prepare 2X master mix
- Add optimized supplement reagents
- Assemble 20 μL reactions
- Load into microfluidic device wells
- Introduce simulated breath/saliva samples
- Incubate at 30°C
- Capture fluorescence using optical sensor
- Record time-course data (0–36 hrs)
- Convert fluorescence to digital signals
- Apply neuromorphic encoding
- Compare outputs across biomarkers
- Validate reproducibility
Expected Results
- Increased Mg²⁺ → higher protein expression
- Increased glucose → longer reaction duration
- Multiplex detection → distinct fluorescence outputs
- Signal processing → improved classification accuracy
Techniques Used ✔ Cell-Free Systems ✔ DNA Construct Design ✔ Microfluidics ✔ Lab Automation ✔ Data Analysis ✔ Bioethical Considerations
Technique Expansion
Cell-Free Systems Used to express reporter proteins in a controlled environment. Enables rapid testing and optimization without living cells.
DNA Construct Design Used to engineer biomarker-responsive circuits using aptamers and regulatory elements controlling sfGFP expression.
SECTION 5: RESULTS & VALIDATION
Validation
I validated my project by designing and optimizing a cell-free sfGFP expression system with enhanced reagent composition to maximize fluorescence output.
Protocol
Prepare optimized master mix Add lysate, DNA, supplement Incubate at 30°C Measure fluorescence over 36 hours
Techniques Used
Cell-free reactions enabled rapid testing of protein expression. DNA design ensured efficient transcription and translation. Optimization of Mg²⁺ and glucose improved yield. Fluorescence measurement provided quantitative validation.
Data & Analysis
Challenges
SECTION 6: ADDITIONAL INFORMATION
References
Budget
DNA synthesis (Twist): ~$120 Cell-free lysate: ~$200 Reagents: ~$150 Consumables: ~$50 Instrumentation: ~$100 Total: ~$620