Week 2: DNA Design and Characterization
This week, I focused on the design, simulation, and cloning workflow for CCT4, the core component of my project, ICN (Intercellular Care Network).
1. Part 1: Benchling and In Silico Gel Art
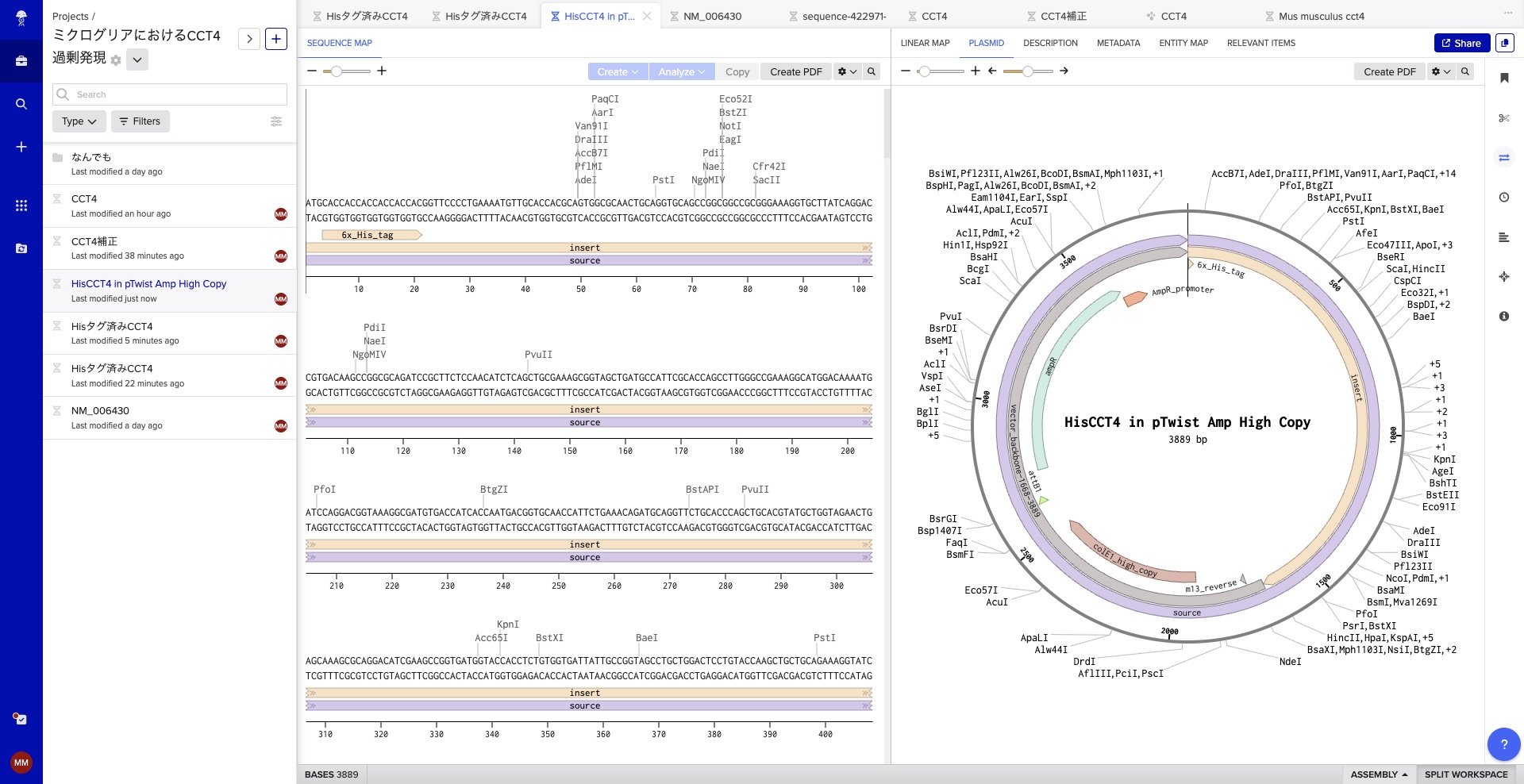
Benchling Simulation
Having long been interested in Benchling, I used this assignment to dive deep into the tool. Instead of the suggested Lambda DNA, I worked with the CCT4 protein sequence relevant to my project. I simulated restriction digests using the following enzymes:
- Enzymes used:
EcoRI,HindIII,BamHI,KpnI,EcoRV,SacI,SalI


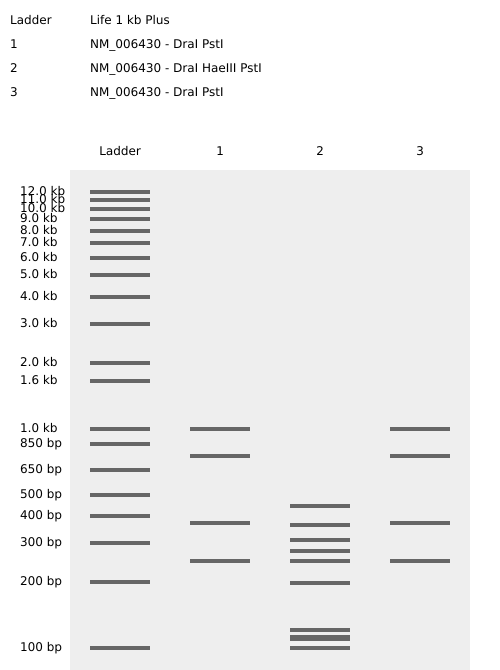
Gel Art: The Tree🌲
I used the virtual electrophoresis simulator to create “Gel Art” representing a tree. Initially, I struggled with the manual placement of bands, but once I understood the physical mechanism of fragment migration, I was able to dial in the design more effectively.

2. Part 2: Wet Lab
As I do not currently have access to a wet lab facility, I focused on digital simulations and perfecting the genetic design for future implementation.
3. Part 3: DNA Design Challenge - CCT4
Why CCT4? (Hypothesis)
CCT4 is a chaperonin that, when overexpressed, not only promotes the formation of TNTs (Tunneling Nanotubes) but also accelerates the speed of cargo transport through them. My hypothesis is that by overexpressing CCT4 in microglia, we can significantly speed up the clearance of toxic proteins like Tau, which are central to Alzheimer’s disease progression. (See my Week 1 HW for more details).
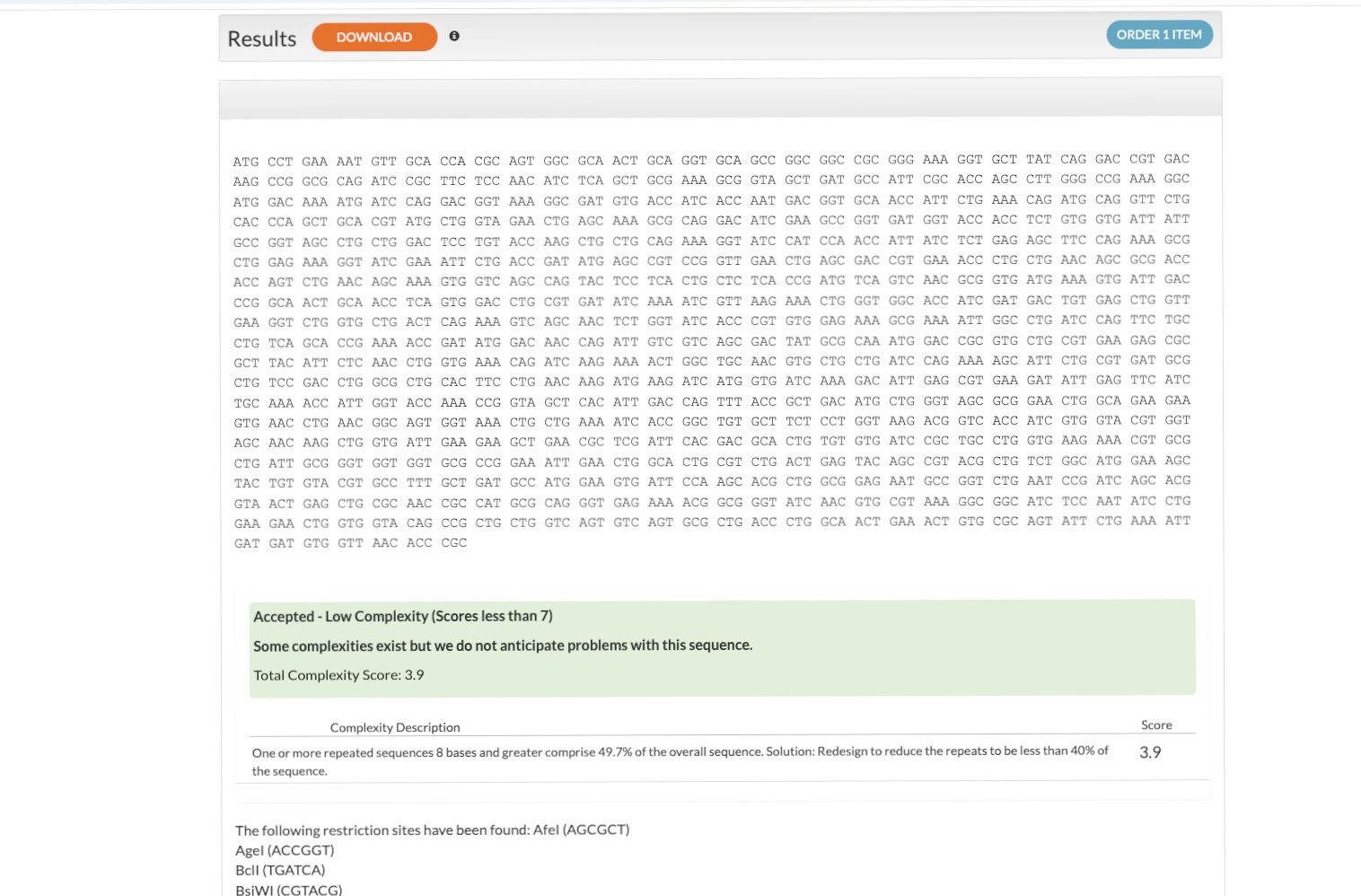
Sequence & Codon Optimization
Based on UniProt P50991 (Isoform 2), I used the IDT Codon Optimization Tool to design a DNA sequence optimized for expression in Escherichia coli (E. coli).
Amino Acid Sequence:
MPENVAPRSGATAGAAGGRGKGAYQDRDKPAQIRFSNISAAKAVADAIRTSLGPKGMDKM IQDGKGDVTITNDGATILKQMQVLHPAARMLVELSKAQDIEAGDGTTSVVIIAGSLLDSC TKLLQKGIHPTIISESFQKALEKGIEILTDMSRPVELSDRETLLNSATTSLNSKVVSQYS SLLSPMSVNAVMKVIDPATATSVDLRDIKIVKKLGGTIDDCELVEGLVLTQKVSNSGITR VEKAKIGLIQFCLSAPKTDMDNQIVVSDYAQMDRVLREERAYILNLVKQIKKTGCNVLLI QKSILRDALSDLALHFLNKMKIMVIKDIEREDIEFICKTIGTKPVAHIDQFTADMLGSAE LAEEVNLNGSGKLLKITGCASPGKTVTIVVRGSNKLVIEEAERSIHDALCVIRCLVKKRA LIAGGGAPEIELALRLTEYSRTLSGMESYCVRAFADAMEVIPSTLAENAGLNPISTVTEL RNRHAQGEKTAGINVRKGGISNILEELVVQPLLVSVSALTLATETVRSILKIDDVVNTR

3.4. Specific Procedures in Cell-Based Expression (E. coli Expression System)
For this project, I am assuming the process using E. coli, which is the most common and reliable cell-based expression system.
Transformation Introduce the designed DNA (plasmid) into the E. coli cells. This is the process of having the E. coli read the CCT4 blueprint by applying stimuli such as heat shock.
Culturing and Expansion Efficiently grow only the E. coli that harbor the DNA in a medium containing antibiotics. We maximize the population of E. coli in an incubator, providing optimal temperature and agitation.
Induction of Expression Once the E. coli has sufficiently proliferated, reagents like IPTG are added to flip the CCT4 protein production switch. Protein synthesis within the cells begins immediately from this point.
Recovery and Purification Once a sufficient amount of protein has been produced, the E. coli cell walls are disrupted (lysis) to retrieve the contents. Subsequently, tags such as His-tags are used to extract only the pure CCT4 protein from the mixture.
3. Reasons for Choosing This Method
While cell-based methods take more time compared to cell-free methods, their greatest advantage lies in the ability to mass-produce large quantities of protein at a low cost. In order to broadly deploy the ICN network and care for many cells in the future, this E. coli-based mass production system will be an indispensable infrastructure.
4. Part 4: DNA Synthesis & Cloning
I constructed a CCT4 expression cassette for E. coli transformation.
Design Refinement: I resolved initial issues by adding a Stop Codon and a His-tag for purification. I excluded promoters/RBS as they are provided by the destination vector.
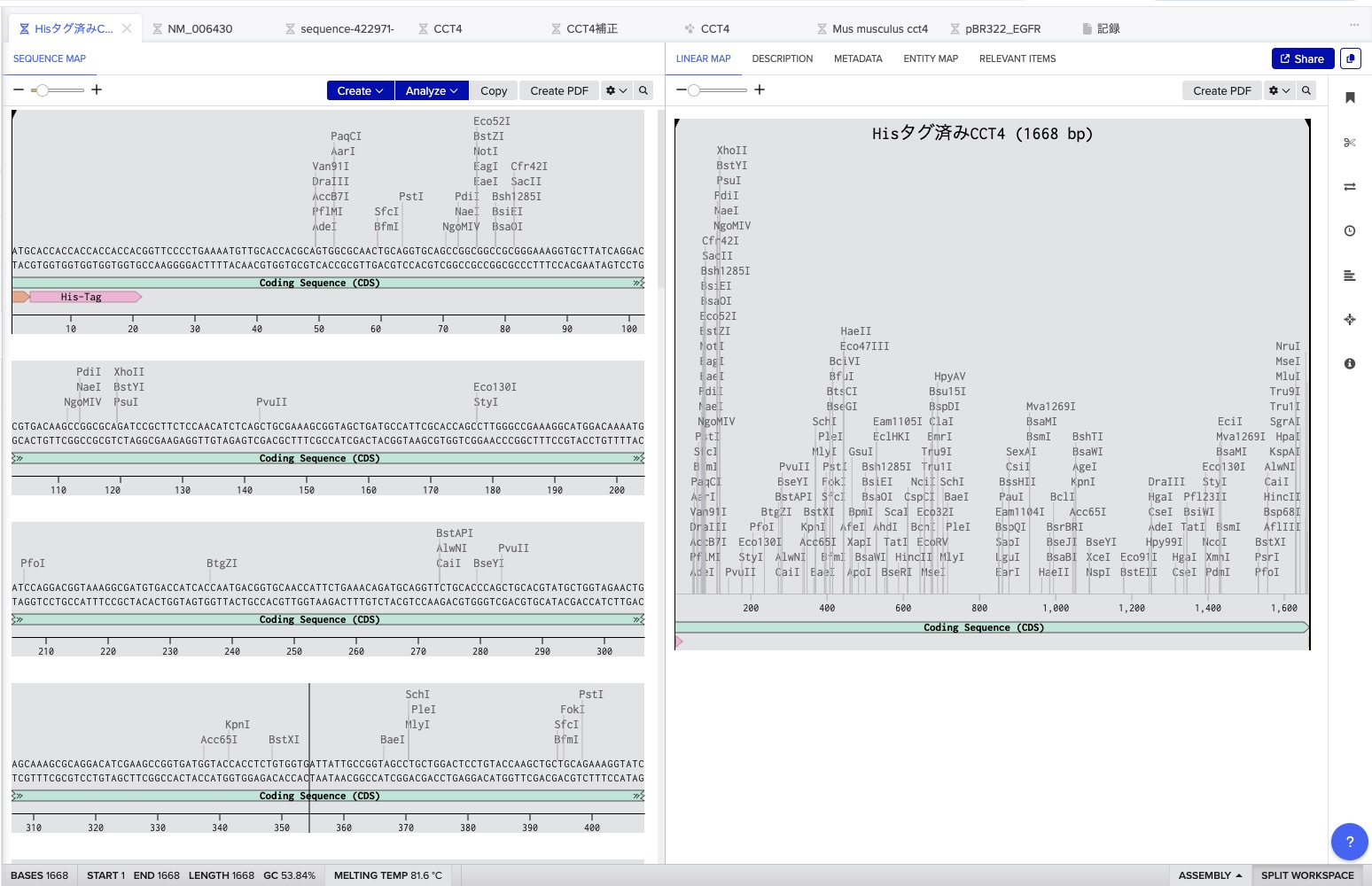

Execution: I exported the FASTA from Benchling and uploaded it to Twist Bioscience. After cloning into the
pTwist Ampvector, I imported the Genbank file back into Benchling to finalize the plasmid map. This was my first-ever digital cloning experiment!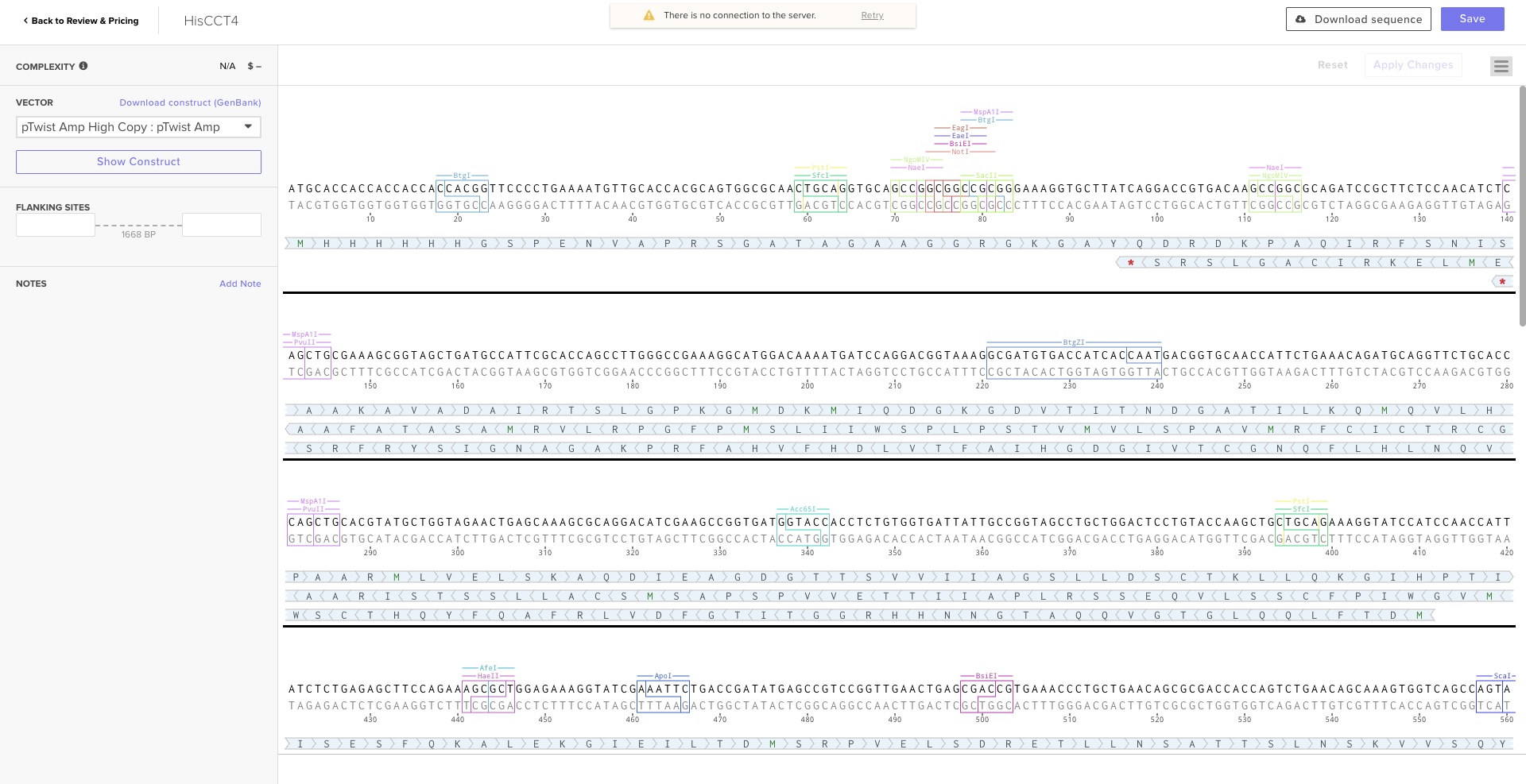

5. Part 5: DNA Reading, Writing, and Editing
5.1 DNA Reading
Target DNA and Reason: I will target the CCT4 gene region within human microglial cells and its corresponding mRNA to reflect its expression status. The objective is to quantitatively understand how chaperonin function within microglia changes during the progression of Alzheimer’s disease and to verify if the engineered genes are functioning as intended.
Technology and Reason: I will utilize Nanopore Sequencing, which is classified as a 3rd-generation technology. Unlike conventional methods, it can read long-read DNA or RNA in real-time without PCR amplification. This provides a significant advantage by allowing analysis without losing structural complexity or epigenetic information related to repair networks.
Detailed Process:
- Input: High-purity genomic DNA extracted from microglia.
- Preparation: The extracted DNA is purified to an appropriate length, and adapters containing motor proteins are ligated to both ends to guide the DNA into the nanopores.
- Decoding: As the DNA passes through the nanopore, changes in ionic current are detected as waveforms, which are then converted into base sequences using neural networks (basecalling).
- Output: Data in FASTQ format, containing base sequences and quality scores.
5.2 DNA Writing
Synthetic DNA and Reason: I will synthesize an optimized sequence of the human CCT4 gene designed for maximum production efficiency in E. coli. This physical DNA fragment is required to create a prototype aimed at enhancing microglial intercellular care functions and suppressing the aggregation of abnormal proteins.
Technology and Reason: I will use the Silicon-based Phosphoramidite method. This technology, utilized by companies like Twist Bioscience, enables the simultaneous synthesis of tens of thousands of DNA fragments on a microscopic silicon plate, achieving overwhelming scalability and cost reduction for individual gene synthesis.
Detailed Process:
- Key Steps: Each cycle consists of four chemical reactions—deprotection, coupling, oxidation, and capping—to accurately assemble the sequence base-by-base.
- Limitations: Due to the nature of the chemical synthesis process, a length of approximately 1.6 kb requires a lead time of about two weeks. Additionally, since error rates accumulate as the sequence lengthens, post-synthesis verification via sequencing is mandatory.
5.3 DNA Editing
Target DNA and Reason: I will target safe harbor loci, such as the AAVS1 region within the microglial cell genome. This allows for the stable integration of the exogenous CCT4 gene into the genome to maintain long-term therapeutic effects while avoiding the risk of disrupting other vital genes.
Technology and Reason: I will use CRISPR/Cas9. This technology allows for extremely high-precision genome editing, as designing guide RNAs (gRNA) to target specific base sequences is straightforward.
Detailed Process:
- Editing Mechanism: A complex consisting of gRNA, which binds to the target genomic sequence, and the Cas9 enzyme, which acts as molecular scissors, is introduced into the cell. After the DNA is cleaved at the target site, accurate gene insertion is performed via Homology-Directed Repair (HDR) using the provided CCT4 template.
- Preparation and Input: This requires selecting the target sequence, designing the gRNA, and preparing the Cas9 enzyme along with the plasmid containing the CCT4 gene to be integrated.
- Limitations: There are risks of off-target effects, where unintended locations are edited, and physical constraints such as the tendency for gene integration efficiency to decrease in non-dividing or slowly dividing cells like microglia.