Week 3 pentrons Bio-Art & Lab Automation
Assignment: Telecaster Guitar Bio-Art
This week’s task was to create a Python script for the Opentrons OT-2 robot to draw an artistic design on an agar plate using colored bacteria.
1. Design Concept: The Telecaster
My design is based on the Fender Telecaster, an iconic electric guitar. I used different fluorescent proteins to represent the components:
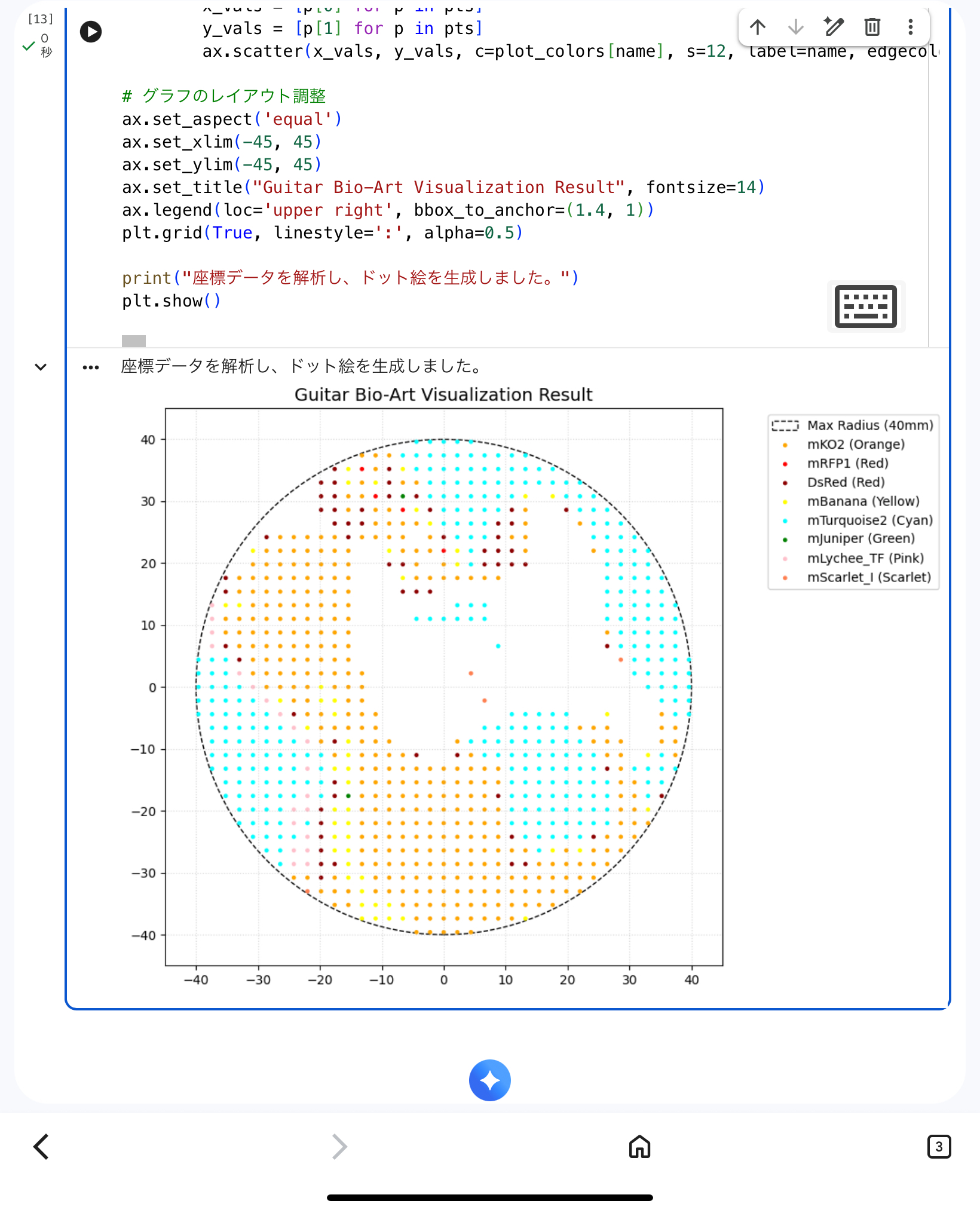
- Body Contour: mKO2 (Orange)
- Neck & Headstock: mTurquoise2 (Cyan)
- Pickguard: mBanana (Yellow)
- Pickups & Hardware: mRFP1 / DsRed (Red)
2. The Challenge: Recovering a “Lost” Design
While I successfully generated the coordinates using the Automation Art Interface, I made the mistake of closing the browser before taking a screenshot.
How I Solved It:
Instead of re-clicking hundreds of points, I took the coordinate list from the exported Python script and used Google Colab with Matplotlib to recreate the design visually. This allowed me to:
- Verify that all points were within the 40mm safety radius.
- Ensure the aesthetic balance of the guitar was preserved.
- Generate a high-fidelity simulation image to replace the lost screenshot.
3. Coding & Troubleshooting (Technical Journey)
I refined the script in Google Colab. Several technical hurdles were overcome:

- Environment Setup: Standard Colab environments do not have the
opentronslibrary. I resolved this by running!pip install opentronsand restarting the runtime. - Simulator Errors:
- Labware Missing: The simulator didn’t recognize the custom
htgaa_agar_plate. I swapped it for a standardcorning_96_wellplate_360ul_flatfor simulation purposes. - Slot Occupied Error: An error occurred because Slot 6 was already occupied by the system. I updated the script to load the source plate into Slot 1 to avoid hardware conflicts.
- Labware Missing: The simulator didn’t recognize the custom
- Droplet Precision: To prevent smearing or “stringing” between dots, I implemented a
dispense_and_detachfunction that moves the pipette 2mm upward after each drop.
4. AI Documentation
I collaborated with Gemini 3 Flash for this assignment:
- Logic Support: Reorganizing raw coordinate data into a clean, iterable dictionary.
- Debugging: Interpreting “LocationIsOccupiedError” and suggesting slot reassignment.
- Visualization: Writing the Matplotlib script to plot the coordinates for verification.
Post-Lab Questions
Q1: Automation in Research
Paper: “An open-source, high-throughput liquid handling system for automated DNA assembly.” (Example: [Kanigowski et al., 2021]) This paper utilizes the Opentrons OT-2 to automate Golden Gate Assembly. By using robots, they reduced human error in complex pipetting steps and scaled up the creation of genetic constructs, achieving a 90% reduction in cost compared to proprietary systems.
Q2: Final Project Automation Plan





