Week 6 Genetic Circuits Part I: Assembly Technologies
Assignment: DNA Assembly & Chromophore Engineering
1. Phusion High-Fidelity PCR Master Mix Analysis
Question:
What are some components in the Phusion Master Mix and what is their purpose?
Calculation:
The master mix is formulated for high-speed, high-accuracy amplification.
Result:
Phusion DNA Polymerase: A pyrococcus-like enzyme with a processivity-enhancing domain. Its $3’ \to 5’$ exonuclease activity provides proofreading for “high-fidelity” results.
2. Factors Determining Primer Annealing Temperature ($T_a$)
Question:
What factors determine $T_a$ during PCR?
Calculation:
$T_a$ is calculated based on the melting temperature ($T_m$) of the primer-template hybrid.
Result:
GC Content: Higher GC content increases $T_m$ due to triple hydrogen bonds.
Primer Length: Longer primers generally have higher $T_m$.
Salt Concentration: Monovalent and divalent cations ($Na+, Mg{2+}$) stabilize the DNA backbone, increasing $T_m$.
Mismatches: Intentional mismatches (like those used for amilCP mutation) lower the effective $T_m$.
3. DNA Linearization: PCR vs. Restriction Enzyme Digest
Question:
Compare and contrast these two methods.
Calculation:
Comparison of protocol flexibility and accuracy.
Result:
PCR: Amplifies a specific region from a small amount of template. It is preferable when you need to add Gibson overlaps to the ends of the fragment via primer overhangs.
Restriction Digest: Cuts existing plasmids at specific sites. It is preferable for preparing a backbone from a known vector where high sequence fidelity (no PCR errors) is required.
Contrast: PCR creates “new” DNA copies; Digest simply cuts existing DNA.
4. Pre-requisites for Gibson Assembly
Question:
How to ensure sequences are appropriate for Gibson cloning?
Calculation:
Design based on homology and purity.
Result:
Overlap Identity: Fragments must share 20-40 bp of identical sequence at their junctions.
Sequence Purity: PCR products must be treated with DpnI to remove the methylated template plasmid and then purified (e.g., Zymo kit) to remove primers and salts.
Verification: Run a diagnostic gel to confirm that the backbone and insert fragments show bands at the correct predicted sizes.
5. Mechanism of Bacterial Transformation
Question:
How does plasmid DNA enter $E. coli$ cells?
Calculation:
Analyzing the impact of the temperature gradient.
Result:
During Heat Shock ($42^\circ\text{C}$ for 45s), the rapid temperature change creates a thermal imbalance across the cell membrane. This induces the formation of transient pores in the chemically competent cell wall, allowing the plasmid DNA to enter the cytoplasm via diffusion.
6. Alternative Assembly: Golden Gate Assembly (GGA)
1.Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
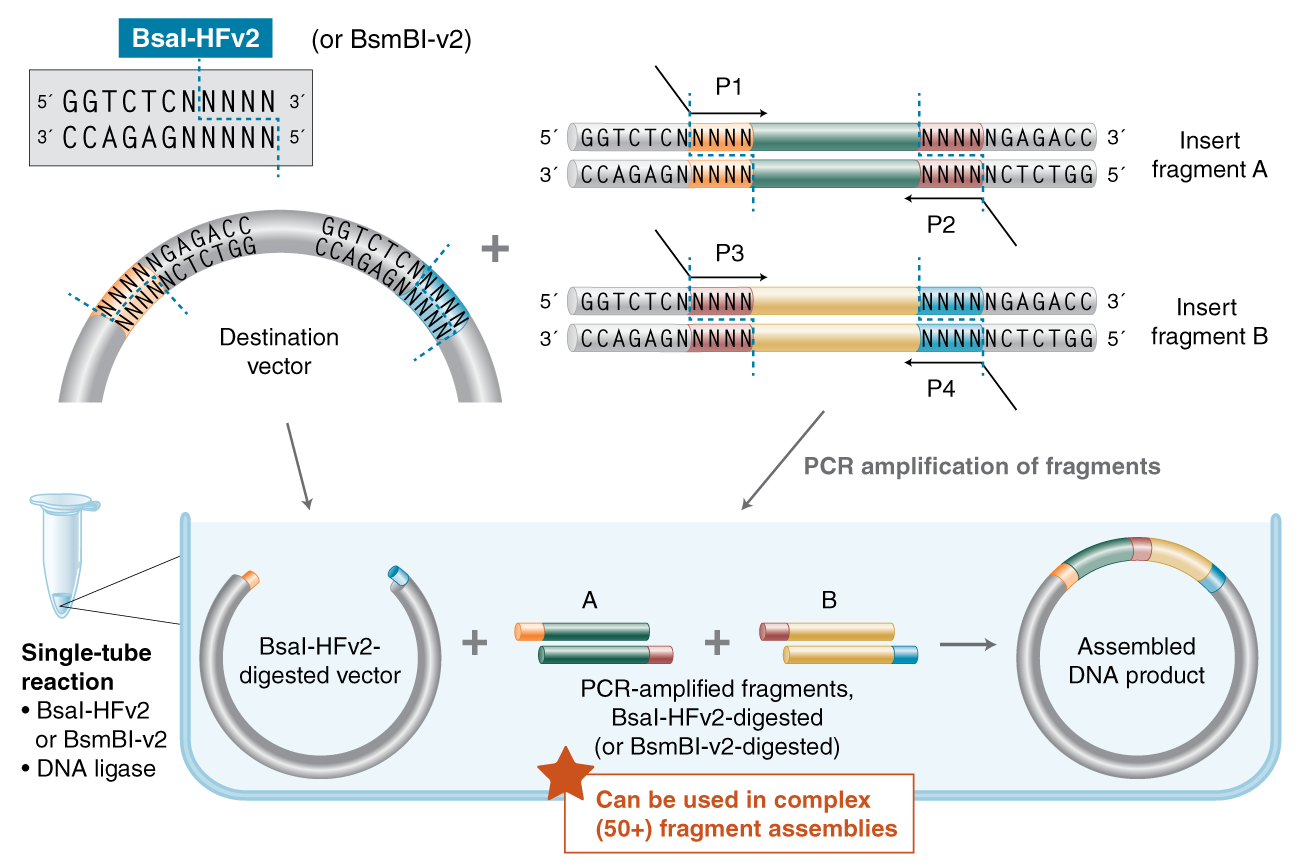
Golden Gate Assembly (GGA) is a highly efficient molecular cloning method that utilizes Type IIS restriction enzymes and T4 DNA ligase in a single-pot reaction. Unlike traditional enzymes, Type IIS enzymes like BsaI or BsmBI cut DNA at a specific distance away from their non-palindromic recognition sites, creating unique 4-nucleotide overhangs. These overhangs can be custom-designed to be complementary, allowing multiple DNA fragments to be assembled in a precise, pre-defined order. Since the recognition sites are oriented to be removed during the digestion process, the final assembled product lacks the original restriction sites, making the reaction irreversible and “scarless.” This allows the digestion and ligation steps to occur simultaneously through repeated temperature cycling in a thermocycler. GGA is particularly powerful for creating combinatorial libraries or complex multi-part genetic circuits where high speed and accuracy are required. By carefully choosing the 4-bp overhangs, researchers can ensure that only the correctly assembled circular plasmid is stable, drastically reducing background colonies.
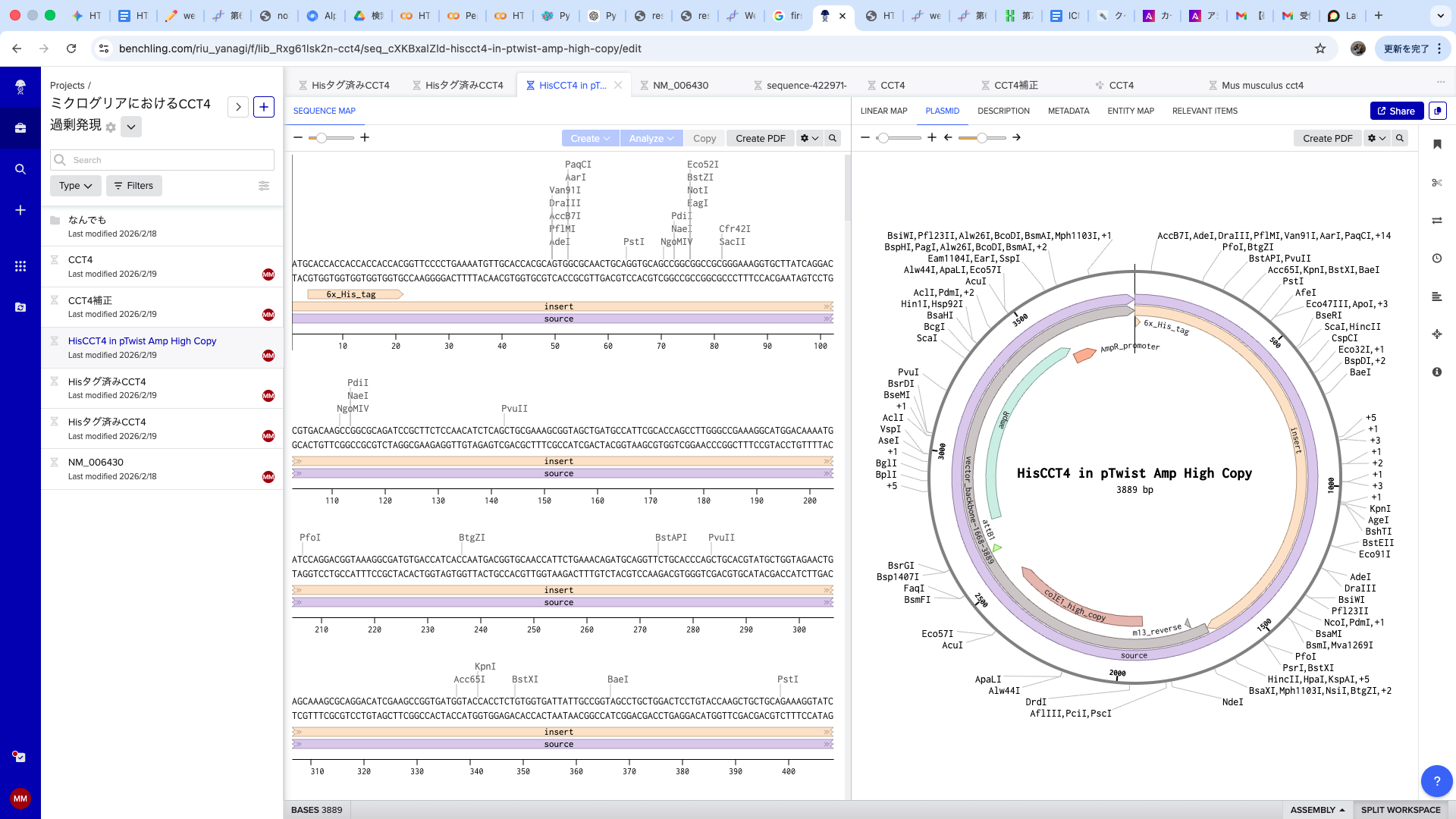
2.Model this assembly method with Benchling or Asimov Kernel