Week 2 HW: Reading and Writing DNA Sequences
Start Your Homework!
Part 1: Benchling & In-silico Gel Art
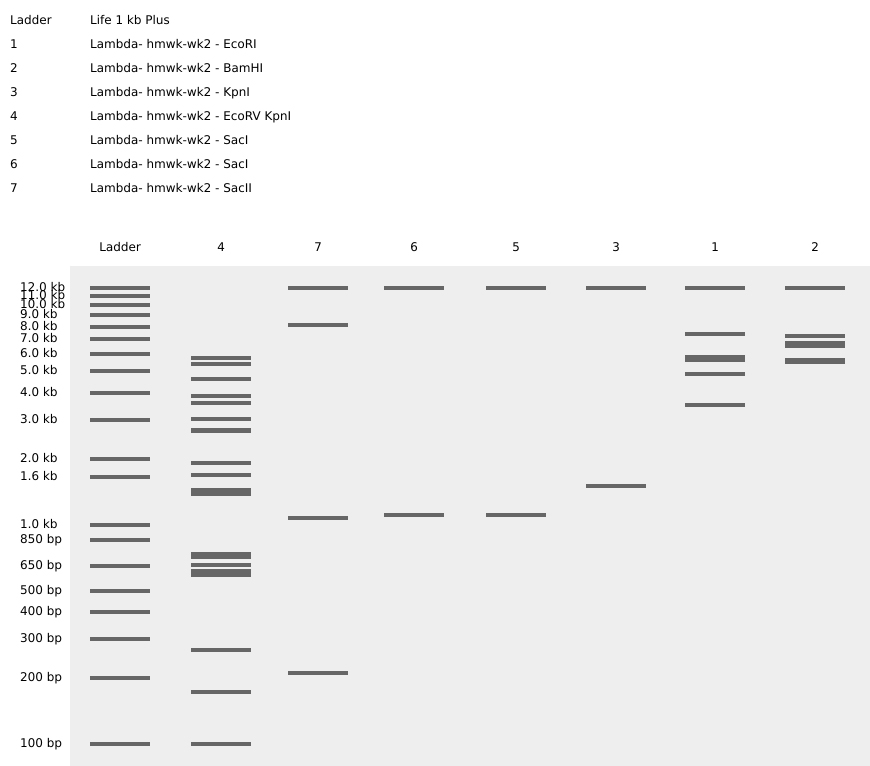
Johnny Got His Gun
-The sequence has been rearranged into the shape of a pistol. I wanted to keep the number of columns restricted to the original seven enzymes for this composition, but I’m sure the image could be improved with additional columns. Question? How many columns can be introduced to the gel matrix?

Part 3: DNA Design: Pick a Protein… Any Protein?
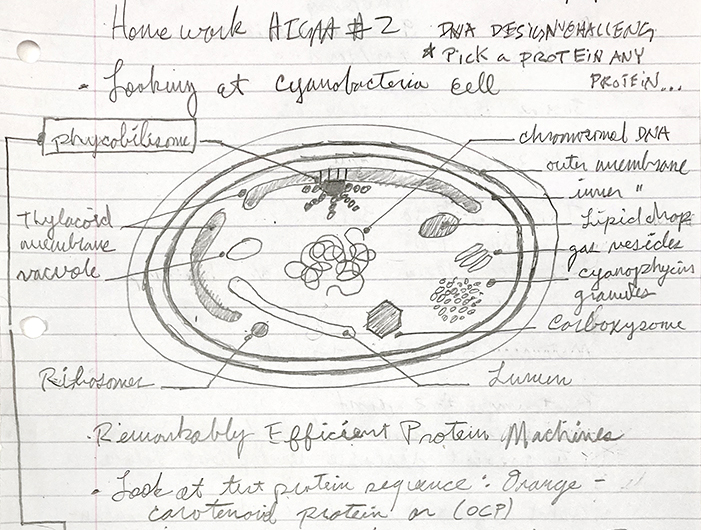
The organism - Chroococcidiopsis - contains the Orange Carotenoid Protein (OCP)
Microscopic detail of Chroococcidiopsis
As seen in the page from my notes above, I’m interested in Chroococcidiopsis, a type of cyanobacteria, which is the focal point of my research with PIG. A protein, OCP, is found in the Phycobilisome (PBS), which is a large light harvesting complex attached to the Thylacoid Membranes within the cyanobacteria cell body. It has 6483 genes, and I have no idea what any of this means, and I’m not sure if this protein - which was one of over 41,000 results to my search for “proteins commonly found in cyanobacteria” - will be of any importance to my project. Apparently, it acts as a sunscreen with its orange coloration for the thylacoid membrane, which is important for regulating photosynthesis within the cell. I chose this protein in particular for primarily two reasons, it’s a necessary part of a functional cyanobacterial cell, and it has been isolated as a plasmid, which seems to be important in context to the work we will be doing within HTGAA.
Reverse Translation of OCP Protein sequence to DNA sequence.
The OCP sequence: 1 mpytiesars ifpdtqvasa vptivesfeq lsaedrlall wfaytemgvt itpaamqvan 61 mmfaektlaq ieqipaaeqt qvmcdlinht dtpicrtysy fgmnvklgfw yqlgewmkqg 121 ivapipegyk lsakasnvlq tirqleggqq ltvlrdivvn mghspttatq kveepvvppk 181 dlaprtkivi eginnstvls ymenmnafdf eaavalfaed galqppfeep ivgqesilaf 241 mreecyglkl ipergisepg ergftqikvm gkvqtpwagd svginlawrf linrqgkiff 301 vaidvlaspq ellnlglvk
Reverse translation (most likely codons) of URD53675.1 orange carotenoid-binding protein (plasmid) [Chroococcidiopsis sp. CCNUC1] to a 957 base sequence:
atgccgtataccattgaaagcgcgcgcagcatttttccggatacccaggtggcgagcgcg gtgccgaccattgtggaaagctttgaacagctgagcgcggaagatcgcctggcgctgctg tggtttgcgtataccgaaatgggcgtgaccattaccccggcggcgatgcaggtggcgaac atgatgtttgcggaaaaaaccctggcgcagattgaacagattccggcggcggaacagacc caggtgatgtgcgatctgattaaccataccgataccccgatttgccgcacctatagctat tttggcatgaacgtgaaactgggcttttggtatcagctgggcgaatggatgaaacagggc attgtggcgccgattccggaaggctataaactgagcgcgaaagcgagcaacgtgctgcag accattcgccagctggaaggcggccagcagctgaccgtgctgcgcgatattgtggtgaac atgggccatagcccgaccaccgcgacccagaaagtggaagaaccggtggtgccgccgaaa gatctggcgccgcgcaccaaaattgtgattgaaggcattaacaacagcaccgtgctgagc tatatggaaaacatgaacgcgtttgattttgaagcggcggtggcgctgtttgcggaagat ggcgcgctgcagccgccgtttgaagaaccgattgtgggccaggaaagcattctggcgttt atgcgcgaagaatgctatggcctgaaactgattccggaacgcggcattagcgaaccgggc gaacgcggctttacccagattaaagtgatgggcaaagtgcagaccccgtgggcgggcgat agcgtgggcattaacctggcgtggcgctttctgattaaccgccagggcaaaatttttttt gtggcgattgatgtgctggcgagcccgcaggaactgctgaacctgggcctggtgaaaReverse translation (consensus codons) of URD53675.1 orange carotenoid-binding protein (plasmid) “"[Chroococcidiopsis sp. CCNUC1]” to a 957 base sequence:
atgccntayacnathgarwsngcnmgnwsnathttyccngayacncargtngcnwsngcn gtnccnacnathgtngarwsnttygarcarytnwsngcngargaymgnytngcnytnytn tggttygcntayacngaratgggngtnacnathacnccngcngcnatgcargtngcnaay atgatgttygcngaraaracnytngcncarathgarcarathccngcngcngarcaracn cargtnatgtgygayytnathaaycayacngayacnccnathtgymgnacntaywsntay ttyggnatgaaygtnaarytnggnttytggtaycarytnggngartggatgaarcarggn athgtngcnccnathccngarggntayaarytnwsngcnaargcnwsnaaygtnytncar acnathmgncarytngarggnggncarcarytnacngtnytnmgngayathgtngtnaay atgggncaywsnccnacnacngcnacncaraargtngargarccngtngtnccnccnaar gayytngcnccnmgnacnaarathgtnathgarggnathaayaaywsnacngtnytnwsn tayatggaraayatgaaygcnttygayttygargcngcngtngcnytnttygcngargay ggngcnytncarccnccnttygargarccnathgtnggncargarwsnathytngcntty atgmgngargartgytayggnytnaarytnathccngarmgnggnathwsngarccnggn garmgnggnttyacncarathaargtnatgggnaargtncaracnccntgggcnggngay wsngtnggnathaayytngcntggmgnttyytnathaaymgncarggnaarathttytty gtngcnathgaygtnytngcnwsnccncargarytnytnaayytnggnytngtnaar
Question 1. The translation results were produced using bioinformatics’ Sequence Analysis - Reverse Translate which produced the results above. In addtion it produced code for graphing the base probabilities. How is this code converted in a graph? Python?
Question 2. What is the difference between most likely codons and consensus codons?
Question 3. In addition to nitrogenous bases ACTG, there are also: n,y,h,r,w,s and m. These represent codon combos which form amino acids? - yes
Codon Optimization
Clearly there are advantages as well as disadvantages to Codon Optimization, and after reviewing the basic concept it seems there are two divergent camps, pro optimization vs. the purist. Let’s compare.
Some Advantages:
- Optimization can increase protein production by making more efficient use of codons by using a simplified version of synonymous amino acid combinations. This could help improve the efficiency of producing therapeutic drugs, and potentially reduce production cost.
- Certain codon combinations can potentially increase the expression of a gene thereby making more affective treatments.
- Optimization helps to align a target gene’s codon usage with the preferred codons of the host organism.
Some Disadvantages:
- Modification of protein structure can affect protein function or performance.
- Potentially, there are deeper levels of code in specific codon combinations which could be related to rythms associated with protein folding patterns during elongation, or functional aspects which have not yet been discovered.
- Optimization can also affect wobble of tRNAand potentially cause unwanted mutations or render proteins disfunctional.
Based off this surface research, I can see reasons for and against codon optimization - and as with any experimental research - we must carefully weigh the potential benefits of optimization relative to the application. For instance, if the optimization is for use in animal research, which could potentially impact the wellbeing of an organism with unintended “side effects”, then comprehensive multi level testing should be conducted to exhaust or reduce error within the application. If the optimization is purely lab based and not able to modify DNA replication, and will improve the speed of data and research assimilation then I can see this as a definite advantage to employ the technology.
Codon Optimization- novoprolabs.com, I used this because the Twist link was broken. The sequence type was: DNA/RNA with E.coli as expression host which produced the following result from most likely codon group:
ATGCCGTATACCATCGAGTCTGCACGTAGCATCTTCCCAGACACCCAGGTTGCGTCTGCGGTTCCGACCATTGTGGAGTCTTTCGAGCAGCTGTCCGCGGAAGATCGCCTGGCGCTGCTGTGGTTCGCTTACACCGAAATGGGCGTCACCATCACCCCGGCGGCGATGCAGGTAGCAAACATGATGTTCGCGGAAAAGACCCTGGCTCAAATCGAACAGATTCCGGCAGCAGAACAGACTCAGGTAATGTGCGATCTGATCAACCATACCGATACGCCGATTTGCCGTACTTACTCCTACTTTGGCATGAACGTCAAACTGGGTTTCTGGTATCAACTGGGTGAATGGATGAAGCAGGGTATCGTTGCGCCGATCCCGGAAGGCTACAAACTGTCCGCCAAGGCCTCCAACGTGCTGCAGACTATCCGCCAGCTGGAGGGTGGCCAGCAGCTGACCGTTCTGCGTGACATCGTCGTTAACATGGGTCACTCCCCAACGACCGCTACTCAGAAAGTTGAAGAGCCGGTCGTGCCGCCGAAAGATCTGGCGCCGCGCACCAAAATCGTGATTGAAGGTATCAACAACAGCACCGTTCTGAGCTATATGGAGAACATGAACGCGTTCGATTTCGAGGCCGCAGTTGCCCTGTTCGCAGAAGACGGCGCTCTGCAGCCGCCGTTCGAAGAACCGATCGTTGGCCAGGAGTCCATCCTGGCCTTCATGCGTGAAGAATGCTATGGCCTGAAACTGATCCCGGAACGTGGCATTAGCGAACCGGGTGAGCGTGGTTTCACTCAGATCAAAGTGATGGGCAAAGTTCAAACCCCGTGGGCGGGCGACAGCGTAGGCATCAACCTGGCATGGCGTTTCCTGATCAACCGTCAAGGCAAGATTTTCTTCGTTGCCATCGACGTACTGGCGAGCCCGCAGGAGCTGCTGAATCTGGGTCTGGTAAAA
Results from consensus codons, sequence type- protein, expression host- Ecoli:
1 GCAACCGGCT GCTGCAACAC CGCGTACGCG TGTAACGCTA CCCATGGTGC GCGTTGGTCC AACGGCTGTA
71 ACATGGGCAA CTGGTCCAAC GCTACGCATA CCACTTACTG CTGCAACGGC GCTTACGCGT GTAACTGCGC
141 ACGCGGCACC AACGGTTGCA ACTGGTCCAA CGGTTGTAAC GGTACTAATT GTTGCAACGC ATGCAACGCG
211 ACGCACGGTA CCAACGGTGC GCGTTGGTCT AACACCACTT ACGGCGCTCG TTGCGCGCGC TACACCAACT
281 GGTCTAATGG CTGCAACGGC GCGCGTGGCG CATACATGGG TAACTACACT AATGGCTGTA ACTACACCAA
351 CTACACCAAC ACCGGCGGTA CCACGTACGG TTGTAACACT GCATATGCAT GTAACGGCGC GCGTGCAACT
421 GGCGGTGGTA ATGGTACGAA CGCCTGTAAC GCGACTCACG CGTGCAACTG TTGCAACGGC TGCAACGGTT
491 GCAACGCAAC CGGCTGTGCT CGTGGCACTA ATGGCTGCAA CGCAGCTTAT GCAACCGGCG CTACCGGTAC
561 CACCTACGGT TGTAACGGTG CGCGTGCCGC TCGTGCCTGC AACTACACCA ATGGTTGTAA TTGCGCTCGC
631 GCGACCCACG GTGCCCGCTG CGCTCGTGCA ACCCATTGCT GCAACGGCTG TAATGGCTGT AATGGCGCGC
701 GCTGCGCGCG TGCATGTAAT TGCGCGCGCG GTACCAACGC GACTGGCACC GGTTACGGCG CCTATTACAC
771 CAACGCGACC CATGCAGCAT ACTGTGCCTA TGCGTGCAAC GGCGCATATG CATGCAATTG CTGCAACGCG
841 ACCCACACCG GTTATATGGG TAATGCGTGC AACACGGCTT ATTGGAGCAA TACGGCTTAC ACCACTTACG
911 GTGGTAACGC CACCGGTGCT GCGTATGGCA CGAACGCTGC GCGTTATACG AACGGTGGCA ACACCACCTA
981 CACTGGCGGC ACCGCGTATT GTGCGCGTTA CACCAACGGT GGTAACGGCG CTCGCACCGG TGGTGCCACG
1051 GGCGCGGCAC GTTGCGCACG TGGCGGTAAC GCTACCCACG GCACTAACGG TTGCAATTGT TGCAACGCGA
1121 CTCACTGCTG CAACGGTGCA CGTGGCGGTA ACACCGCATA TGCTGCGCGT TACACCAACT GGTCTAACGG
1191 CTGCAATGCG GCTCGCGGTT GCAACTGGAG CAACGCAGCG TACGGCACCA ACTACACTAA CTGCGCACGC
1261 GCTTGCAACG CAACCCATAT GGGCAACTGT GCGCGTTATA CGAACGGCGC GCGTGGTGGT AACGGCGGTA
1331 ACTGCGCTCG TTGTGCTCGT TACACCAACG CGTGTAATGG TACTAACTAC ACCAACATGG GTAACGGTGC
1401 ATACGCGACT CACGGTACCA ACGGTACGAA CGCTGCGTAC GCGACTGGCG GCGGCAACTG CGCTTACTGG
1471 TCTAATTGTT GCAACGCCTG CAATGCATGT AACGGCTGTA ACGCCTGCAA CTGCGCTCGT GCGGCACGTG
1541 GTACCAACGG CGCGCGTGGT GCTCGCTGCT GCAACGGTAC CAACGGTACC AACTGTTGCA ACTGTTGCAA
1611 CGCTGCGCGC GGTGCGTACT ACACCAACGG CTGCAATTGC TGTAACATGG GCAACGCTTG TAACGCAGCG
1681 CGTGCAACTC ACGGCACTAA CGCGACTCAC GGCGCTCGTG GTGGTAACGC GACCCACGCT GCTTACGCGG
1751 CATACTGGTC TAACGCATGC AACGGCACCA ATTACACCAA CTGGTCTAAC ACCGCCTACG CCACTGGCGG
1821 TGCGCGTGCG GCGTATGCAA CCGGCGCCGC GTATGGTTGT AACACTACCT ACGGTGCTTA CACCACGTAC
1891 GGCGCGCGCG GCTGCAACGG CTGTAACGGC ACCAATGGTT GCAACTACAC CAACACTACC TACGGTTGTA
1961 ATGGTGCTCG TGGCGCTTAC GGTGGCAACG GCTGTAACTA TACCAACTGC GCTCGCTGCT GCAACTGTTG
2031 TAACACCACT TACGGTGCAC GCGGTGCACG TTGTTGCAAC GCTACCCACG GTACCAATGG CGGTAATTGC
2101 GCTCGTGGTG CTCGTTGGTC TAACGCAACC CACTATACTA ACGGTTGCAA TACCACCTAC GCAACCGGTA
2171 TGGGCAATGG CGCCCGTGGT GCGCGTACCG GTTATACCGC CTACGGTGGT AACTACACCA ACGCAGCTCG
2241 TTACACCAAC GCCACCCACT GCTGTAACGG CGCTCGTATG GGTAATGGTG GCAATGCTAC CCACTGGTCC
2311 AACGGCGCAC GCTGCTGCAA TGGTGGTAAC GGTGCGCGTA TGGGTAACGG CGGCAACACC ACCTATGCCT
2381 GCAACTGCGC ACGTGCAACT CACGCCGCGC GTGGTACTAA CGCGACCGGC GGCGGTAATG CTGCTCGCGG
2451 CACGAACTGT GCACGTGCTT GCAACTGCTG CAATACTGGC GGTGGCTGTA ACGGTGGCAA CGGTGCATAC
2521 TGGAGCAACG GCACCAACGG CGGTAACGCG ACCCATGCAG CTTATTACAC TAACGGTTGT AACACCGGCG
2591 GTATGGGCAA CACCACCTAT TACACCAACG CGACCCACGC AGCTTACATG GGTAATTGTG CACGTGGCGG
2661 CAACGCGGCG CGCGCAACGC ACACGACCTA CACCACCTAC GGCACGAACG GTTGTAACGC GACTCACGGT
2731 GCCTACGGCA CCAACTATAC CAACGGCTGT AATTGGTCTA ACTGTTGTAA CTGCGCTCGT GGCGCCCGTT
2801 ACACGAACTA TACCAACGCG GCCTACTACA CCAATGGCGG CAACTACACT AACGGCACCA ACGCCGCACG
2871 T
Question 4. In the optimized sequence above the 25th character is T of TAC which would indicate RNA start codon AUG?
question 5. Then in the next line below I found the sequence ACTTAC, which would indicate a stop codon with another start directly after in the sequence. Everything in between the start and stop is a particular gene in the chain? Is that the promoter of the sequence?
Question 6. If AUG is start codon, which contains uracile, then start and stop codons only exist in RNA?
You have a sequence! Now what?
I don’t know, this is where it falls apart for me… WAH Wah wah… GAME OVER… INSERT ANOTHER TOKEN!!! I’m failing to grasp the connection between the Orange Caratenoid Protein OCP, which I chose to explore, and how it relates to the host organism, Ecoli, in the codon optimization stage of homework.
Question 7. Am I trying to cut and paste the OCP into the ecoli genome using the benchling software? Or am I looking for the expression genes within the OCP sequence by identifying the starts and stops?
Question 8. How do I determine where start and stop codons are within the sequence? I know Methionine (AUG) is start and there are three stop codons, UAA, UAG, UGA. Do I start reading in sets of three from the begining of optimized sequence looking for TAC, which would be on the DNA side of AUG?
Annotation of Sequence (I think I need a tutor!)
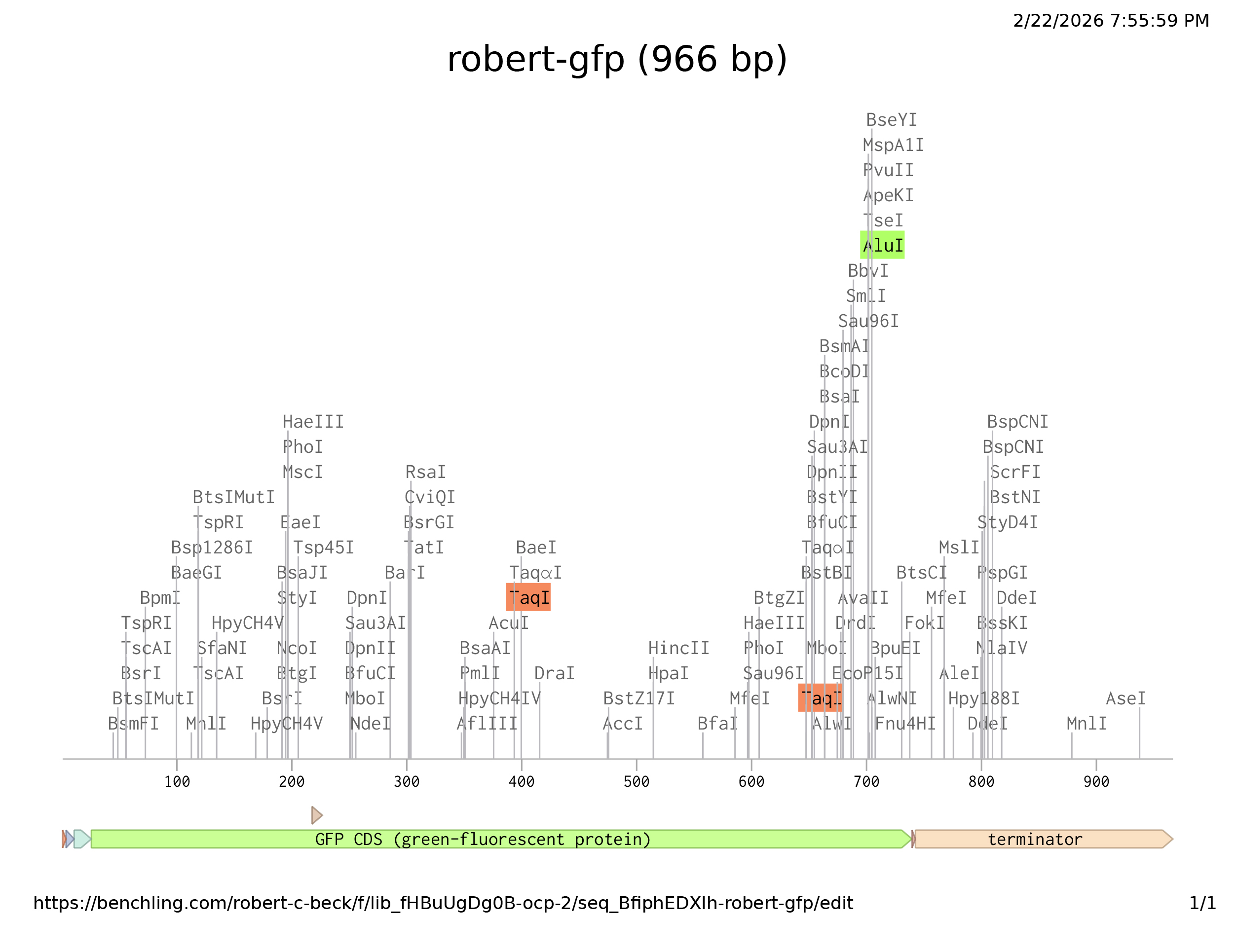

Prepare a Twist DNA Synthesis Order
I was able to create both Benchling and Twist accounts no problem. I imported the optimization sequence above as a new sequence, which marked all the restriction enzyme sites, but was not really able to distinguish promoter region etc. for annotation. Although, after following through with predicted regions and annotation, I arrived at the graphic above. When I imported the file into Twist Twist could not complile the sequence. I’m not sure where I went wrong.
Question 9. How do I determine promoter, RBS, etc. ?
-RBS stands for Ribosomal Biding Site- It is a sequence of of nucleotides (codons) upstream of the start codon (AUG Methionine) in mRNA and has 5-7 nucleotides rich in A and G.
-Start codon in DNA is TAC, and it should be followed by a stop codon (ATT,ATC,ACT).
Question 10. Should I have fed the optimization results from the consensus codons into Benchling?
DNA Read
- My Primary goal with HTGAA is to build my understanding cyanobacteria as the building block of not only my artwork, but also as a cellular organism which forms the foundational base of the ecological food chain. I cannot even begin to think or dream of how, what, or why I would want to alter the cellular systems of cyanobacteria at this point, but as my knowledge of these organisms develops I know my perspective will change.
DNA Write
What DNA would you want to synthesize (e.g., write) and why?
When I started this homework assignement, over a month ago, I had no idea what I could or would want do with this opportunity, and to be honest, I’m not sure I feel much differently. Obviously, this technology has amazing potential to make changes which could, and have had a profound impact on people’s lives. As it relates to my research within visual art and its relationship with biological materials, I can see working with microorganisms to express complex concepts, which can enrich subject matter by creating dynamic relationships with other organisms. It’s really profound to think about it from this perspective. But, I’m at the begining of a new journey, and I would like to begin by writing an expression of color into a cyanobacterial cell. With this accomplishment I will have established enough understanding to begin exploring other methods to work with bacteria.What technology or technologies would you use to perform this DNA synthesis and why?
“if I have seen further [than others], it is by standing on the shoulders of giants.”
Isaac NewtonRefering to Newton, the research that’s been done in this field by others is really quite humbling. Especially when trying to work through some of these exercises. I follow the giant’s instructions, but I’m blinded by the light of perspective. Working with a known cynobacterial genome/sequence seems like the best place to start. Studying it in silico by working with Benchling will help me to understand how genes, and thier functional coding process produces protein for cellular operations. I hope this will lead to a basic comprehension of the systems by making simple adjustments one step at a time. I have to walk before I can run. Once I have taken my first step, and actually attempted to produce some form of DNA, probably by using restriction enzymes to cut and knock in a new sequence, I will need to test my work by sequencing it. If I am successful, then I would think using PCR to amplify the synthesized product to test in vivo would be the next step. As for scalability, I am not concerned with this yet. I think it would be amazing to have the ability to produce enough plasmid to modify a cyanobacterial image which is at least 30cm square. If I could do that, then I’m sure I will be able to work out larger scales.
DNA Edit
- What DNA would you want to edit and why?
I pretty much feel the same way about this as writing, and if I’m writing anything it will be something I’ve copied. In a certain sense I guess that’s what editing really is, little tweaks to something which already exists, and I would approach this one step at a time with the most appropriate technique. Since I would like to work with cyanobacteria, plasmids seem to be the most approachable form of modification. The goal is to shift the blue/green color spectrum of cyanobacteria from yellow to yellow-green, green to blue-green, and blue-green to blue. I would probably start by identifying a protein which is strong within the yellow spectrum. Then design a plasmid coded for the chosen protein. Order the plasmid, and test the design using ecoli, since it won’t produce any background color competition, to see if/how well it works. If I have positive results, I would then try it in cyanobacteria, by using either a heat shock or electroporation to introduce the plasmid. Culture cells on agar and compare results against an unmodified color control.
References
- Image of Chroococcidiopsis -Villanueva, Chelsea & Hasler, Petr & Dvorak, Petr & Poulíčková, Aloisie & Casamatta, Dale. (2018). Brasilonema lichenoides sp. nov. and Chroococcidiopsis lichenoides sp. nov. (Cyanobacteria): Two novel cyanobacterial constituents isolated from a tripartite lichen of headstones. Journal of Phycology. 54. 10.1111/jpy.12621.
- Sound JK, Bellamy-Carter J, Leney AC. The increasing role of structural proteomics in cyanobacteria. Essays Biochem. 2023 Mar 29;67(2):269-282. doi: 10.1042/EBC20220095. PMID: 36503929; PMCID: PMC10070481.
- Shang,J. Chroococcidiopsis_sp._CCNUC1 genome. (2022). College of Life Sciences, Central China Normal University, Central China Normal University, Wuhan, Hubei, 430079, China. Available at: https://www.ncbi.nlm.nih.gov/nuccore/CP097482.1 (Accessed: 23-3-2026)
- Paremskaia AI, Kogan AA, Murashkina A, Naumova DA, Satish A, Abramov IS, Feoktistova SG, Mityaeva ON, Deviatkin AA, Volchkov PY. Codon-optimization in gene therapy: promises, prospects and challenges. Front Bioeng Biotechnol. 2024 Mar 28;12:1371596. doi: 10.3389/fbioe.2024.1371596. PMID: 38605988; PMCID: PMC11007035.