Week 4 HW: Protein Design I
Hi- It’s time to do your homework!


Icosahedronic Molecular Modeling I -RCBeck
Part A
Questions:
How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons) Solve- How many Daltons are in 500 grams? 1g = 6.022173643E+23 Daltons (6.022173643E+23 x 500)/100 = 4.29 x 10E+18
Answer- There are approximately 4.29 x 10E+18 amino acid molecules in 500 grams of meat.
Why do humans eat beef but do not become a cow, eat fish but do not become fish?
Answer- The process of digestion uses acids and enzymes to break tissues down into simpler molecules for use in cellular mechanics. This process helps provide the rescources/energy to power our own cellular processes which reassemble the digested amino acids, sugars and such into the forms required by each cell to perform its function which is dictated by the organisms genetic code.
Why are there only 20 natural amino acids?
Answer- According to Andrew Doig, a chemical biologist at the University of Manchester in the UK, it goes all the way back to LUCA. The twenty natural amino acids found in all living organisms are the building blocks we share in common. It is speculated that these twenty in particular are the most stable do to their structure, and able to maintain their functional forms after being buried, or exposed to the environmental conditions of Earth 3.5–3.8 billion years ago when LUCA first emerged, probably from some archaic form of RNA. They are more hydrophobic than hydrophilic, and that may have something to do with their role in LUCA’s evolution because of the way they are able to fold and work together structuraly.
Can you make other non-natural amino acids? Design some new amino acids.
Answer- Yes, absolutely! The only problem though is when trying to incorporate synthetic amino acids into cell function/DNA-RNA the catalytic function of those new amino acids have to be incorporated into the cell’s chemical function of enzymatic activity.
Where did amino acids come from before enzymes that make them, and before life started?
Answer- Apparently, there are several theories which range from riding in on an astreroid to chemical reactions occuring through electrical activity-i.e., lightening striking in areas rich in oxygen, nitrogen, hydrogen, carbon and sulphur compounds. The astroid theory has been proven viable by the discovery of at least 86 amino acids on the Murichison meteorite, which landed in Australia in 1969. I beleive it was probably a combination of planetary chemical reactions and celestial seeding.
If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
Answer- Left, because the a helix is usually right handed for the most common amino acids becuase the carbons have an L configuration, but in D-amino acids, the carbons have a D configuration, they mirror the L configuation. x
Can you discover additional helices in proteins? In theory, yes. Although detecting structures outside the typical three is difficult, not to mention the limitation of the physical and chemical properties of helical structures. But with synbio, the possibility of discovering something new is certainly possible, maybe even likely.
Why are most molecular helices right-handed?
Answer- It’s a mystery… I think, but it has something to do with chirality. Perhaps polarity, too? Or maybe the ionic charge of particles, laws of attraction- opposites attract- who knows? Certainly, there must be an answer.
Why do β-sheets tend to aggregate? What is the driving force for β-sheet aggregation?
Answer- It appears as though the proteing is programmed to fold in such a way that it wants to form aggregate structures. This behavior is probably caused by the alternating arrangement of charged and hydrophobic amino acids, which makes them inclined to gather through attraction of oppositely charged particles?
Part B
URD53675.1 orange carotenoid-binding protein (plasmid) Chroococcidiopsis sp. CCNUC1
- For the sake of consistency I decided to stick with the first protein from the week two homework. It’s interesting to see how many different organisms this protein exists in, all with there own unique functions using this molecule.
- It is 319 amnino acids long, most common is A with a count of 29. According to the UNIProt Homology, there are 239 homologs, and InterProscan identifies it as belonging to NTF2-like domain superfamily. Belongs to the orange carotenoid-binding protein family.
- There were quite a few hits for OCP in RCSB PDB. The model above is a 3-D rendering of OCP2 Gloeocapsa sp. PCC 7428 from RCSB PDB.
- PyMol Renderings: Stick and Ribbon (below)
Pymol - Orange Carotenoid-binding Protein
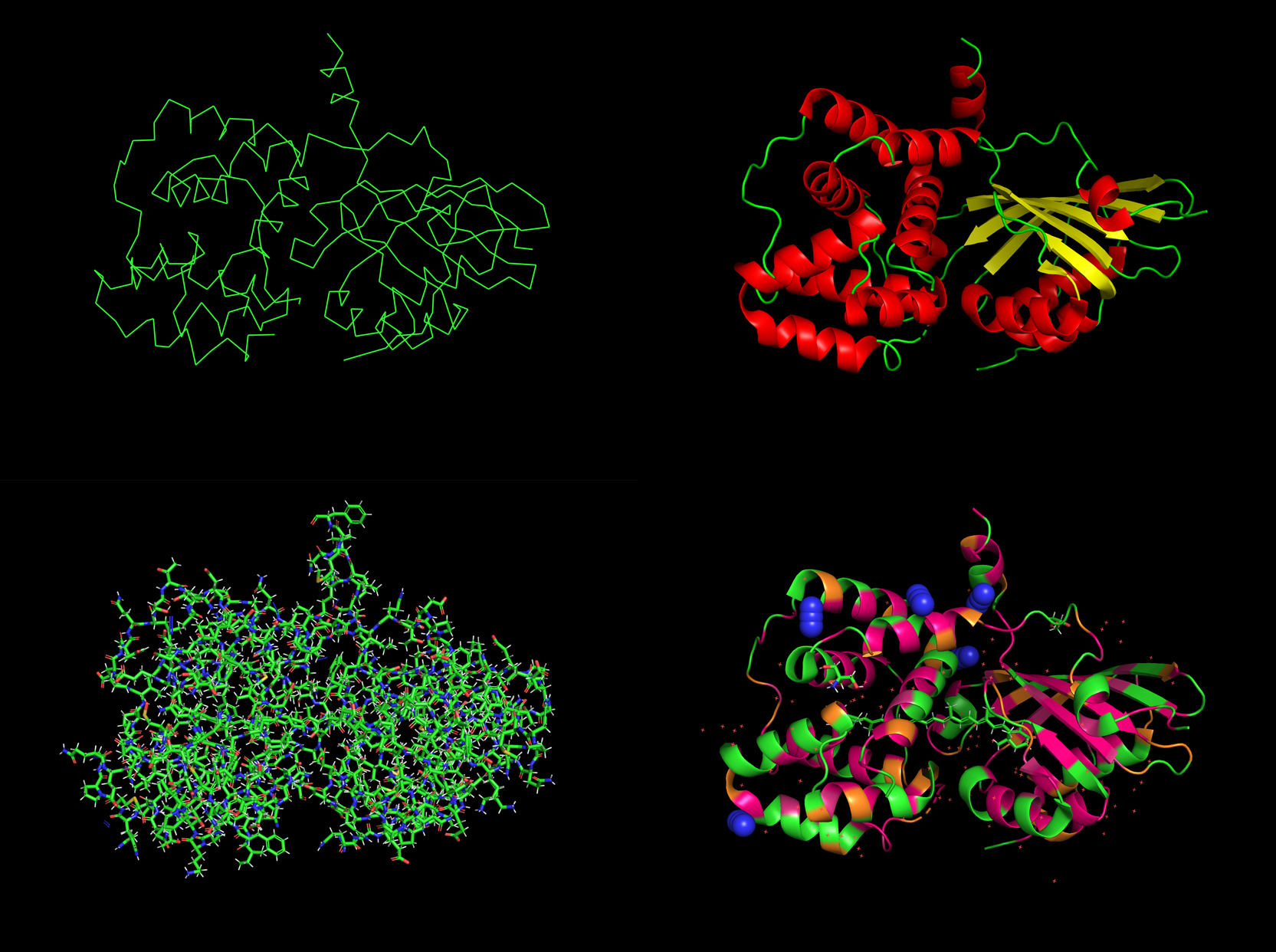
 Clockwise from topleft: ribbon, secondary, stick, hyrdro
Clockwise from topleft: ribbon, secondary, stick, hyrdro
Number of helices: 10
Number of sheets: 6
Hyrophobic with 2,363 atoms in hydrophobic residues vs. Hydrophilic residues with a count of 782 atoms
Binding Pocket Yes, definately several pockets and even some holes leading to core.
Part C - Using ML-Based Protein Design Tools
C1. Protein Language Modeling
Deep Mutational Scans- pdb_00008qx5 Helical Carotenoid Protein 4 (HCP4) from Anabaena with bound Canthaxanthin
.png)
.png) • Heat map visuals indicate binding probabilities for:
• Heat map visuals indicate binding probabilities for:- Sites 43 and 112 exhibit high probability for following bases: A, D, E, G, K, N, Q, R, S, and T.
- C has low probability across all sites except 43 and 112, where it is medium.
- Y and X are fairly low across entire sequence.
Latent Space Analysis
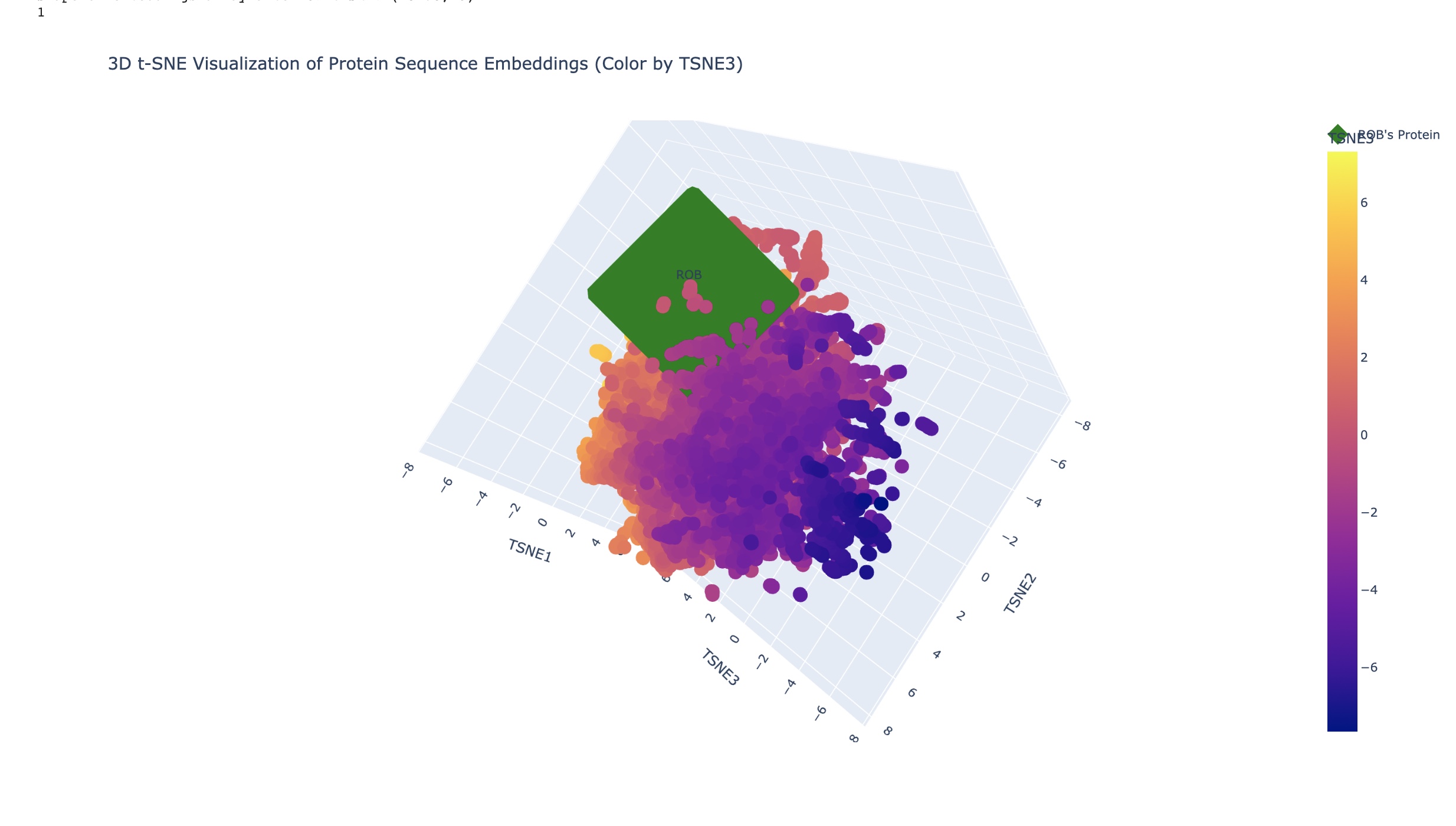

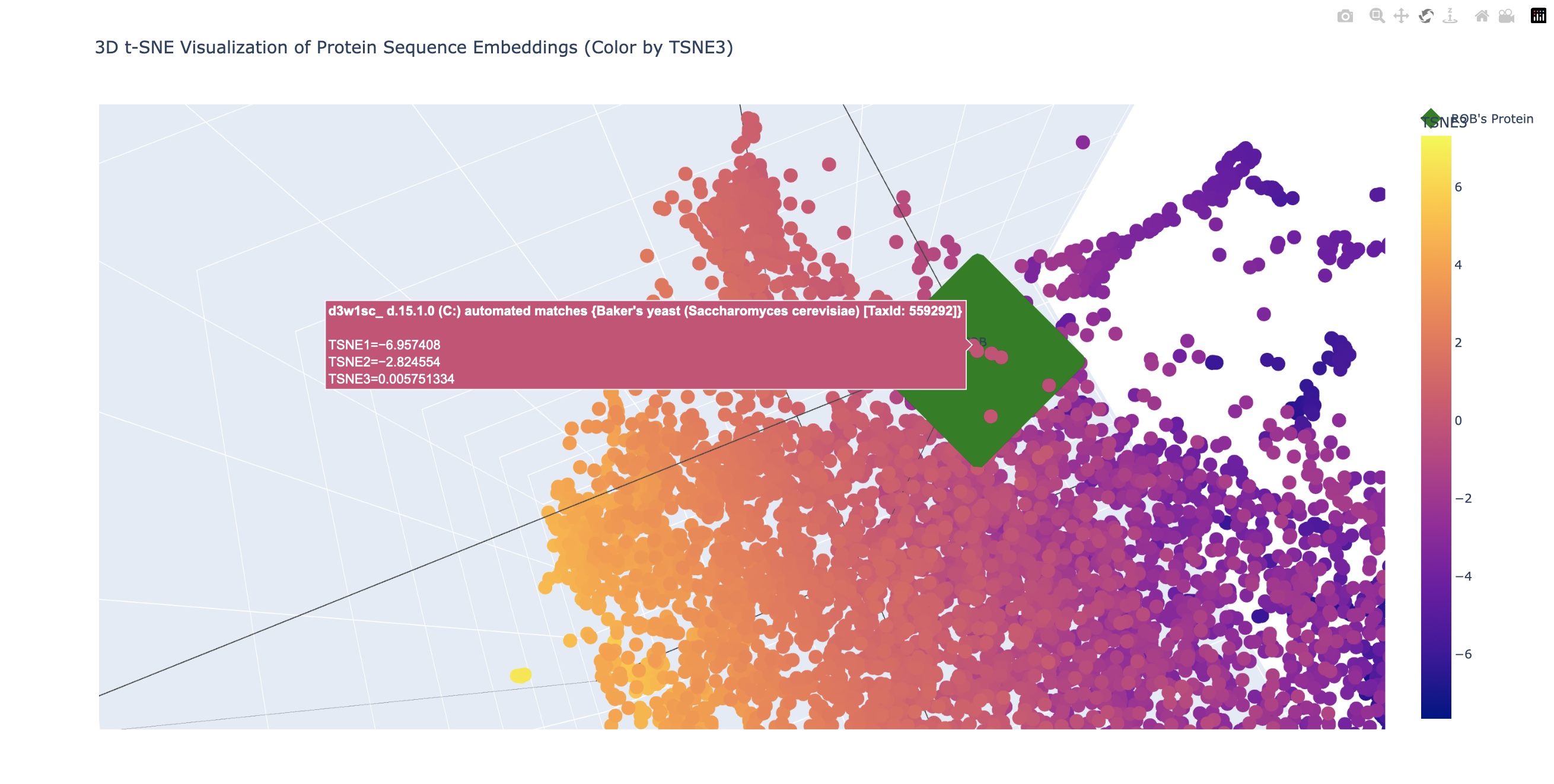
 In the examples above the closest related matches were:
In the examples above the closest related matches were:- Baker’s yeast (Saccharomyses cerevisiae) TaxId: 559292
- Eukaryotic peptide chain release factor ERF2 C-terminal domain (fission yeast, Schizosaccharomyces pombe) TaxId: 48961
- Cytochrome f subunit of the cytochrome b6f complex, transmembrane anchor (Green Alga, Chlamydomonas reinhardtii) TaxId: 3055
C2. Protein Folding
Amino Acid Probability - PDB 8qx5
.png)
.png) Folding a protein:
Folding a protein:
Fold your protein with ESMFold.
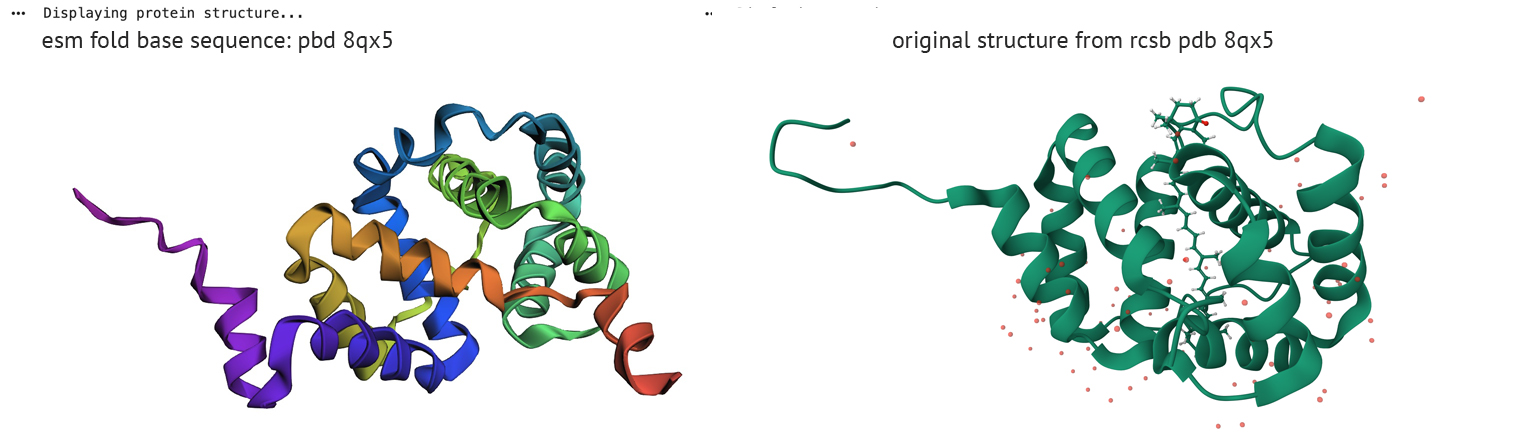

Do the predicted coordinates match your original structure?
ESM fold/reverse fold comparison
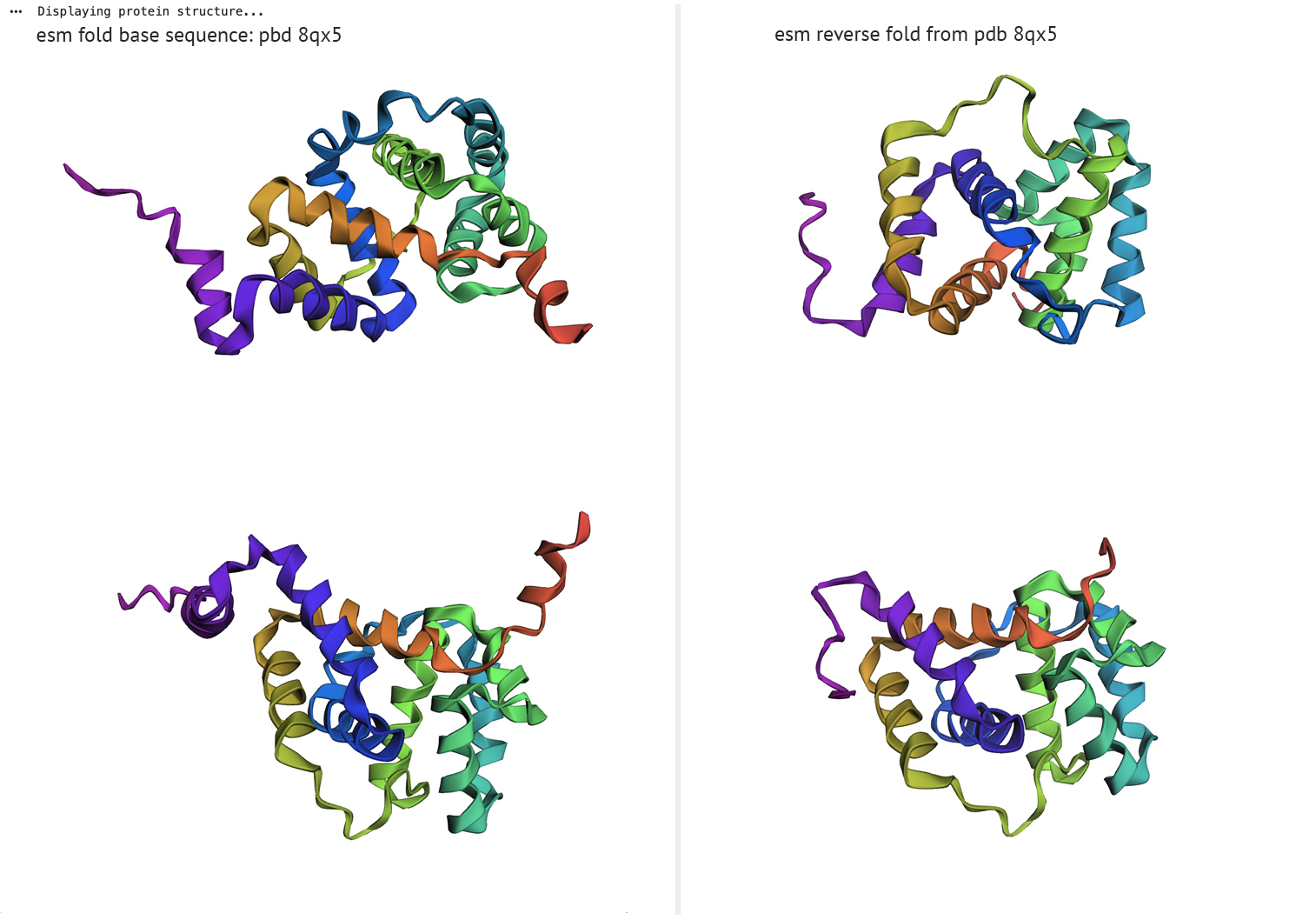
 The left column represents ESM fold from PDB sequence-8qx5, right column displays reverse fold results
The left column represents ESM fold from PDB sequence-8qx5, right column displays reverse fold results
Part D: Group Brainstorm on Bacteriophage Engineering
Proposal by: Robert C Beck, Sameen Nasar
Concept
This is not an easy task - to approach Bacteriaphage theory with a brief introduction, and then cook up a conceptual modification of an extremely complex micro machine by altering its genetic code. In the spirit of HTGAA, here we go.
The first step is to consider what the phage does, what’s its purpose? It’s a virus which carries genetic information, and as a virus its job is to find a host which it can use to replicate itself and spread genetic information. As a machine it consists of different parts - a vessel which contains the genetic information, an attachment mechanism allowing it to connect itself to a host - and a system for transferring its genetic payload to the host. It’s pretty simple in theory, but how these different parts interact with one another to perform their function is very, very, very complex.
The problem with the phage mechanism is bacterial recognition of its presence, which leads to resistance. This raises a question about how a phage attaches to its host, and when it does, how the bacteria responds to the phage. This is what leads to resistance in the first place, bacterial recognition or memory of what the phage’s handshake is like. It reminds me of meeting people during and after covid, and how awkward it was. It was never really clear what to do. Do you bump fists or elbows, nod your head, or should we just wave at each other since you can’t see a polite smile from behind a mask? In short, it was a level of strangeness which comparatively could exist between bacterial cells and phages. Although this is rather tricky indeed, being that bacteria and viruses alike don’t have any feelings per say, but they do have parts which have to interact with one another in order to function.
So, what would it be like if a phage had hands to shake with its bacterial host? Obviously it doesn’t have hands, but it does have a protein which binds to a receptor site on the bacteria’s cell wall. Once it’s attached, it bridges the cell’s wall to inject its genetic payload. If the bacteria gets to know the phage it will no longer shake hands with it, or lean in for a hug and kiss.
What if a genetic circuit was designed and planted into the phage which would alter its handshake? What if that circuit was turned on and off through the action of attaching to bacteria or unloading its payload? The result then might be some form of an alternating handshake, a secret handshake, which would shift from generation to generation.
Group Project Goal:
Engineering a chaperone-independent efficient MS2 lysis protein
Project Rationale
The efficacy of bacteriophage MS2 as an antibacterial agent is currently limited by the host’s ability to evolve resistance. Specifically, E. coli can mutate the molecular chaperone DnaJ (e.g., at position P330), disrupting the essential interaction required for the MS2 lysis (L) protein to fold and function [1.] This interaction is required for proper function of the lysis protein, as DnaJ binds to the N-terminal domain of MS2 lysis protein and alleviates its inhibitory effect on lytic activity.
We propose engineering a self-activating L protein by replacing its inhibitory, chaperone-dependent N-terminal region with a computationally designed, thermodynamically stable scaffold. As this original domain is dispensable for actual lysis but creates the DnaJ dependency [2], our redesign conceptually eliminates the need for the molecular “handshake” between host and phage, allowing MS2 to fold independently and bypass bacterial control mechanisms entirely.
Chamakura KR, Tran JS, Young R. MS2 lysis of Escherichia coli depends on host chaperone DnaJ. J Bacteriol. 2017;199(9):e00058-17. doi:10.1128/JB.00058-17.
Chamakura KR, Edwards GB, Young R. Mutational analysis of the MS2 lysis protein L. Microbiology (Reading). 2017;163(7):961–969. doi:10.1099/mic.0.000485.
Homework Citation:
- Rachel Brazil (2018). The Alphabet Soup of Life. Available at: https://www.chemistryworld.com/features/why-are-there-20-amino-acids/3009378.article (Acessed: 28 February 2026)
- Kirschning A. On the Evolutionary History of the Twenty Encoded Amino Acids. Chemistry. 2022 Oct 4;28(55):e202201419. doi: 10.1002/chem.202201419. Epub 2022 Jul 28. PMID: 35726786; PMCID: PMC9796705.
- Huang, W., Wang, S., Wei, Y. et al. Design and evolution of artificial enzyme with in-situ biosynthesized non-canonical amino acid. Nat Commun 16, 8698 (2025). https://doi.org/10.1038/s41467-025-63733-3
- Medeiros-Silva J, Dregni AJ, Hong M. Distinguishing Different Hydrogen-Bonded Helices in Proteins by Efficient 1H-Detected Three-Dimensional Solid-State NMR. Biochemistry. 2024 Jan 2;63(1):181-190. doi: 10.1021/acs.biochem.3c00589. Epub 2023 Dec 21. PMID: 38127783; PMCID: PMC10880114.
- Greg Huber et al, Entropy and chirality in sphinx tilings, Physical Review Research (2024). DOI: 10.1103/PhysRevResearch.6.013227
- The PROSITE database Sigrist CJA, Cuche BA, de Castro E, Coudert E, Redaschi N, Bridge A. The PROSITE database for protein families, domains, and sites. Nucleic Acids Res. 2026; doi: 10.1093/nar/gkaf1188 [In press] PubMed:41263099 [Full text] [PDF version]