Week 11: Bioproduction and Cloud Labs
OMG! Last One!

 Drawing Air (2026) Detail, Graphite on Paper -RCBeck
Drawing Air (2026) Detail, Graphite on Paper -RCBeck
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
My contribution to the artwork was limited to one pixel, which was initially made to test interaction with the interface. I noticed how pixel contribution was rather competitive, where scripting was automatically writing over contributions. Unfortunately, this discouraged me from contributing any further, which lead me to focus on my final project alternatively.
For the optimization part of the assignment I decided to use AI since my overall understanding of CFPS is still fairly limited. This decision lead me to reserve my interaction with the optimization protocal. Additionaly, I switched gears and chose to work with AI for the sake of time as well as the complexity of the 16 or so variables. Through my interaction with the AI I noticed how certain variables within the protocol were more or less fixed, as certain starting concentrations were more or less set targets.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
- E. coli - BL21 (DE3) Star Lysate: The cell lysate approach is advantageous as it releases cellular contents directly into bioreactor where product of interest may be synthesized. The reactions can be optimized with addition of protease to eliminate specific pathways by breaking proteins down into peptides or even smaller amino acids.
- Salts:
- Potassium Glutamate- Key component in E. coli cytoplasm important to protein folding. In CFPS it’s important as a stablizing agent which promotes efficient protein production.
- HEPES-KOH pH 7.5- The primary buffering agent used to keep pH stable in CFPS, which is important to maintaining optimal conditions for protein production.
- Magnesium Glutamate- Improtant cofactor in enzyme activity and energy systems, such as ATP, required for reactions to take place.
- Potassium phosphate monobasic- Important buffering agent neede to maintain pH as well as phosphate nutrient supply supporting biochemical reactions in protein production.
- Potassium phosphate dibasic- Another important buffering agent, which in contrast to potassium phosphate monobasic, provides potassium and phosphate ions required for enzymatic activity and metabolic functions.
- Energy / Nucleotide System:
- Ribose- Essential component of RNA backbone which translates DNA to protein. It’s also a component of ATP, a neccessary reagent which provides energy in protein synthesis.
- Glucose- Required for the synthesis of ATP, which as previously stated, a neccessary energy source to drive chemical reactions in protein construction. Glucose also helps sustain reactions in CFPS.
- AMP, CMP, GMP, UMP- are nucleotides important to CFPS during RNA transcription.
- Guanine- One of the main nucleotide bases required for production of GTP, which provides energy in the translation process.
- Translation Mix (Amino Acids)
- 17 Amino Acid Mix- Amino acids are the building blocks of protein; without them the translation process will not have product to build proteins.
- Tyrosine- Amino acid with important post transcriptional properties which are an integral part of protein desgin.
- Cysteine- Key amino acid involved in disulfied bonds required for proper protein formation/folding. It’s also important for proper cell function as many cellular proteins require Cysteine for their construction.
- Additive:
- Nicotinamide- Also known as NAD, or adenine dinucelotide, is a coenzyme involved in redox reactions making it an important factor in metobolic processes. It’s also important for DNA repair, cell senescence and imune function required for healthy cell aging.
- Backfill:
- Nuclease free water- nuetral additive important for acheiving desired concentrations of reagents, which helps prevent nucletide degredation and improves efficiency of CFPS.
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
- Identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems.
- sfGFP- SF stands for super folder which is one of this green fluorescent protein’s main strengths, and why it’s extensively used in research. Because of its molecular stability, it is one of the most reliable systems used for successful gene delivery verification.
- mRFP1- Monomeric Red Fluorescent Protein, optimal tempertature for maturation within one hour is 37°C. Overnight incubation improves can improve its brightness. -mKO2- mKusabira-Orange2, recombinant monomer derived from Fungia concinna, it is oxygen-dependent for maturation, and has a higher phototoxicity compared to other FPs.
- mTurquoise2- Has a fast muturation rate < one hour under optimal conditions with long fluorescence lifetime and resistance to photobleaching.
- mScarlet_I- Red fluorescent protein characterised by rapid maturatation (~35m) and superior brightness. Optimal temperature 32-37C.
- Electra2- Monomer, optimal tempreature range is 30°C or 37°C, and can reach maturation within several hours.
- Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation.
- mTurquoise2- Given it’s fast muturation rate (< one hour under optimal conditions), lowering incubation temperature to ~30 will effect maturation rate and slow down the reaction. The long incubation period should be carried out with a sealed plate to avoid evaporation. Adjusting magnesium glutamate for ideal concentration is important in the transcription/translation processes in CFPS. Having too much can cause DNA/RNA build up, whereas too little can slow down cellular machinery thereby inhibiting protein production. Correct nucleotide and amino acid concentrations are also important as they drive both transcription and translation processes.
- The second phase of this lab will be to define the precise reagent concentrations for cell-free experiment: Considering complexity of designing a cell free master mix containing over 16 reagents with conditional variables I experimented with the following AI prompts to generate optmized mixtures.
Well M14
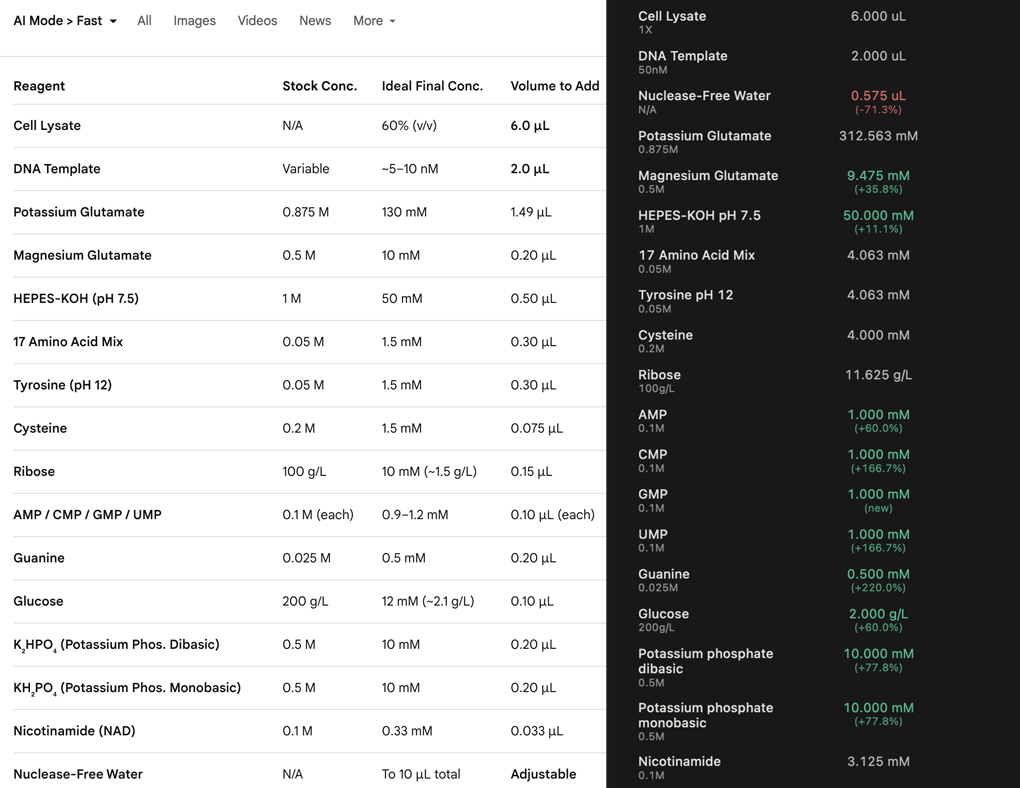
AI prompt: Design cell free protein synthesis protocol for fluorescent protein mTurquoise2 starting with: 6.0 microLiter cell lysate, 2.0 microLiter DNA template, Potassium Glutamate- 0.875M stock with 312.5 millimolar concentration. Adjust amount of Nuclease-Free water in mixture to keep total volume of mixture equal to 10 microliters. Calculate ideal millimolar concentrations for following reagents in mixture:
- Magnesium Glutamate- 0.5M stock
- HEPES-KOS pH 7.5- 1M stock
- 17 Amino Acid Mix- 0.05M stock
- Tyrosine pH 12- 0.05M stock
- Cysteine- 0.2M stock
- Ribose- 100g/L stock
- AMP- 0.1M stock
- CMP- 0.1M stock
- GMP- 0.1M stock
- UMP- 0.1M stock
- Guanine- 0.025M stock
- Glucose- 200g/L stock
- Potassium Phosphate dibasic- 0.5M stock
- Potassium Phosphate monobasic- 0.5M stock
- Nicotinamide- 0.1M stock

Well M-13
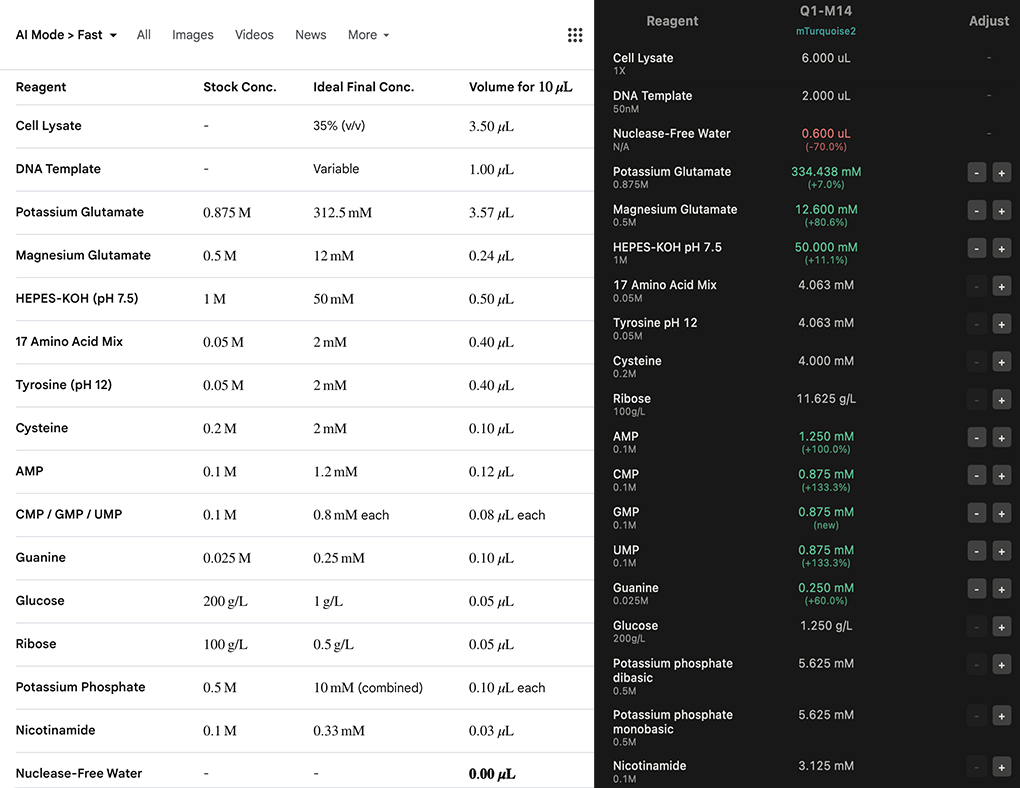
AI prompt: Design cell free protein synthesis protocol for fluorescent protein mTurquoise2 starting with: 6.0 microLiter cell lysate, 2.0 microLiter DNA template. Adjust amount of Nuclease-Free water in mixture to keep total volume of mixture equal to 10 microliters. Calculate ideal millimolar concentrations for following reagents in mixture:
- Potassium Glutamate- 0.875M stock
- Magnesium Glutamate- 0.5M stock
- HEPES-KOS pH 7.5- 1M stock
- 17 Amino Acid Mix- 0.05M stock
- Tyrosine pH 12- 0.05M stock
- Cysteine- 0.2M stock
- Ribose- 100g/L stock
- AMP- 0.1M stock
- CMP- 0.1M stock
- GMP- 0.1M stock
- UMP- 0.1M stock
- Guanine- 0.025M stock
- Glucose- 200g/L stock
- Potassium Phosphate dibasic- 0.5M stock
- Potassium Phosphate monobasic- 0.5M stock
- Nicotinamide- 0.1M stock
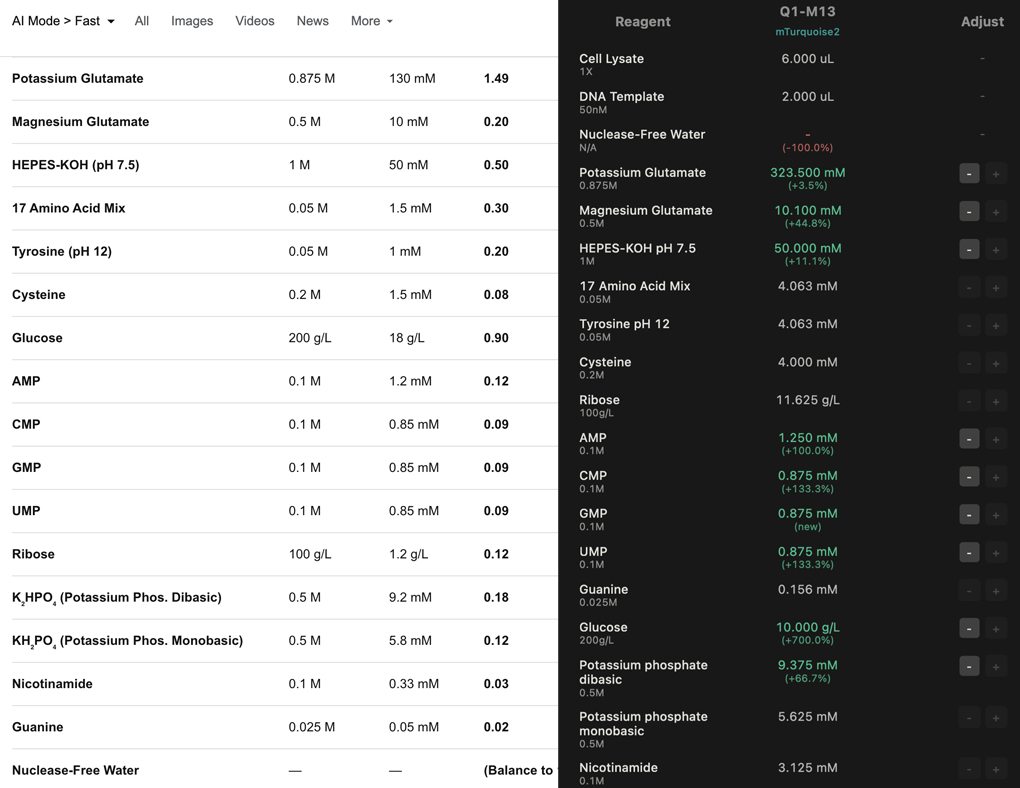

Well M-12