Individual Final Project
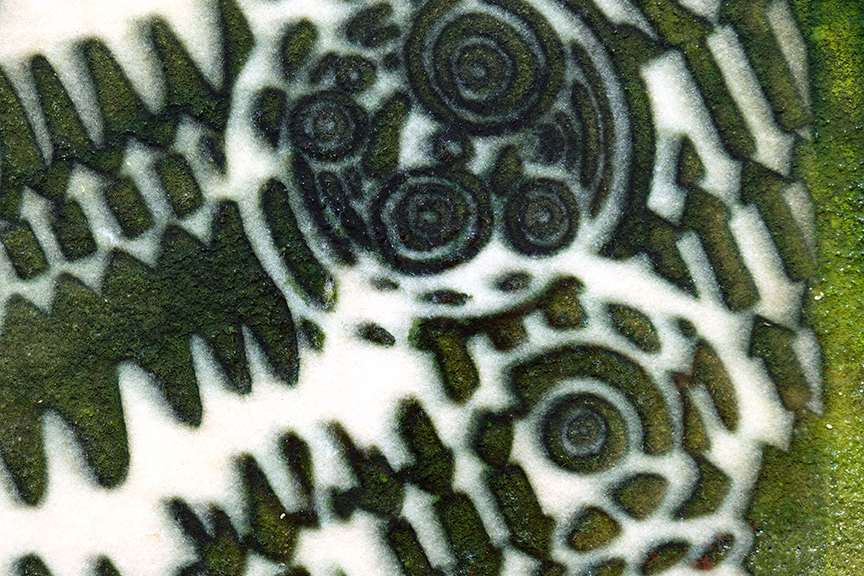


WTF: Wave,Temperature, and Frequency, Cyanobacterial Image on synthetic felt (2020) -RCBeck
PHOTOSYNTHETIC IMAGE GENERATION: Co-Cultured Autotrophic-Heterotrophic Color Controls for Bacteria Based Imagery
Abstract
When Nicéphore Niépce fitted his camera obscura with a special pewter plate in 1822 [1], he created the process of photography and changed the world forever. Since then, photography has evolved to produce striking reflections of real time in space by combining a wide range of materials and processes. Beginning with Niépce’s chemical preparation of a surface to receive reflected light imagery, photography has evolved through chemical means until electronic imaging arrived in the early 1960’s [2] marking the dawn of digital photography.
Both analog and electronic processes rely on photoreceptive elements, which is similar to the photosensitivity of chloroplasts found within the cells of photoautotrophic plants and bacteria. By exploring metabolic pathways in cyanobacteria, this project develops a biochemical photographic process which brings the science of photography full circle, back to its origins within analog media. Considering the critical role biology plays in technology, this potential development would provide a platform for discussion related to biotechnology in visual media arts.
Photosynthetic Image Generation (PIG) harnesses atmospheric carbon to produce cellular energy in the form of sugar which can support the formation of “photographic” products. Further development is promising with the introduction of a co-culture system, which allows cyanobacteria to be engineered for the secretion of sugars and concurrently support the growth of E. Coli [x]. In theory, E. Coli grown together with cyanobacteria could express chromo-proteins, which would introduce the control of color modulation within co-cultured imagery.
Project Aims
- Design and construct plasmids for homologous recombination of cyanobacterial strain, Synechococcus elongatus PCC7942 with the Sucrose permease cscB gene for secretion of sucrose.
- Subaims
- Subaims
- Subaims
- The co-culture system…
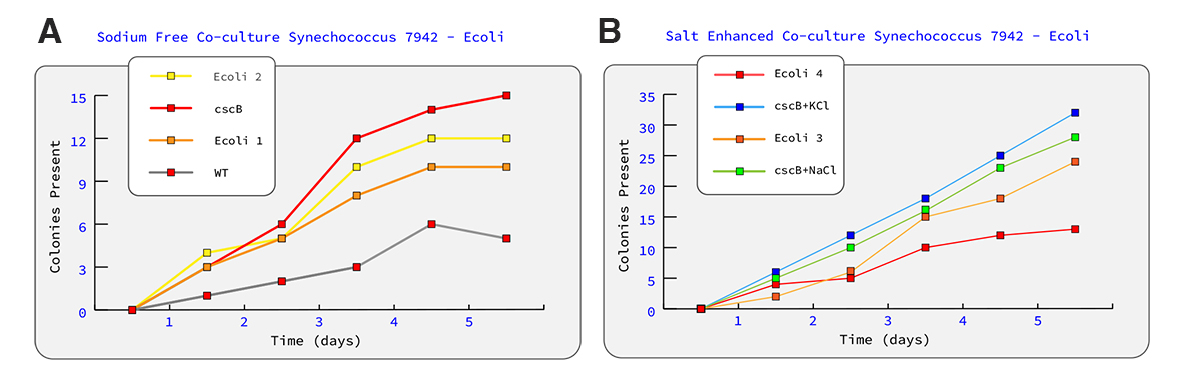
 Figure 1 (Hypothetical Data). Graph A represents growth curve of sodium free cultures of PCC 7942 and Ecoli expressing chromoprotein. Graph B represents two cultures testing the difference between NaCl salt induced stress and KCl as a potential salt stress.
Figure 1 (Hypothetical Data). Graph A represents growth curve of sodium free cultures of PCC 7942 and Ecoli expressing chromoprotein. Graph B represents two cultures testing the difference between NaCl salt induced stress and KCl as a potential salt stress.
Theory Background and Development
Phase 1. Based on research of co-culturing Cyanobacteria Synechococcus elongateus 7942 cscB (S. cscB 7942) with Escherichia coli for bio-chemical synthesis, this project will explore the adaptation of the co-culturing process to produce visual color modifications within an image by generating muticolored bacteria based pixels in a continuous reaction. The origin of this theoretical approach to image cultivation began through a series of visual experiments in collaboration with my partner, Sarah Max Beck, entitled studioHydrostatic. The project was set ontop of a Brooklyn rooftop where we were experimenting with aqauponic technology to create a recirculating, human based, organic food production system. Two large flood tables grew a variety of edible plants and herbs in nitrogen rich water, where I discovered blue-green algae growing in a polka-dot like pattern on a sheet of bubble-foil insulation. Recognizing the growing pattern lead to the developent of a glass plate negative for growing wild caught cyanobacteria in visual patterns. Documentation from 2020 -2021 proved the viability of the concept when the first full size plate was tested in a prototype bioractor producing WTF as seen above.
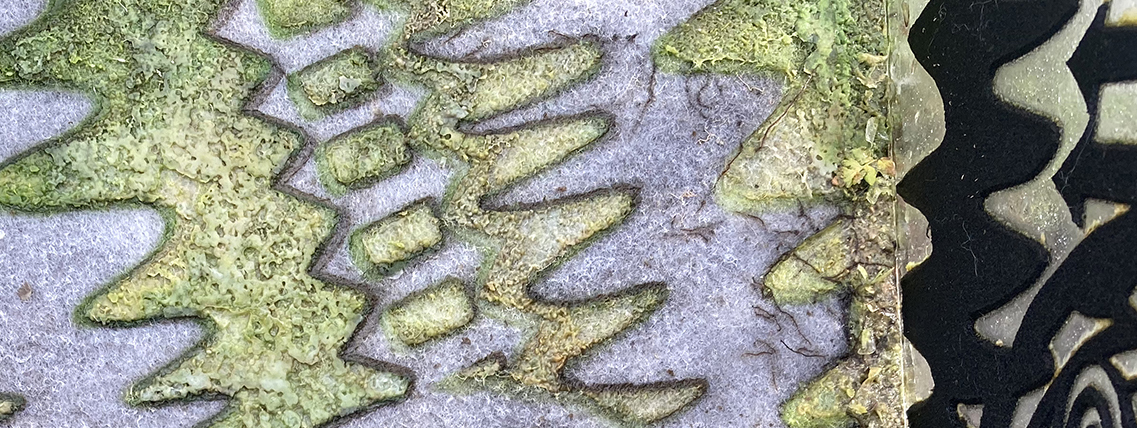


Figure 2. Detail of WTF after one year of growth.
Phase 2. The HTGAA program brings this projects’ theoretical development to the surface.
- As the cyanobacteria growth continued to progress in the WTF image, the density of bacteria progressively darkened the image. It then started to build on itself creating a slightly elevated surface of growth; it appeared to be different from the cyanobacteria. As it progressed the biomass reached a thickness of aproximately 2-3 mm after one year of growth. Eventually, it took over the image, and significantly suppressed the cyanobacteria’s chlorophyll pigmentation (see figure 1). While I can only speculate, it looks as though the image was overtaken by some form of ecoli judging from the color and texture of the biomass.
- My speculation is based on lab grown E. Coli cultures seen during HTGAA labs at Lifefabs indicating there’s a natural relationship which exists between these organisms. Searching for evidence to support the connection, the paper entitled, Cross-feeding between cyanobacterium Synechococcus and Escherichia coli in an artificial autotrophic-heterotrphic cocuture system revealed by integrated omic analysis, revealed that S. 7942 could be engineered to export intracellular sucrose and provide energy for E. Coli, which in turn stabilizes the basifying nature of S. 7942 cultures [x].
In aquatic environments, the ecological interaction between photoautotrophic and heterotrophic species is based on cross-feeding and metabolite exchange. To date, multiprotein complexes that cross cell membrane(s) and extracellular vesicles have been evaluated in photoautotrophs for transporting materials from the interior to the exterior of the cell, which facilitate the secretion of various chemicals ranging from targeted photosynthetic intermediates, such as glycolate, osmolytes and fatty acids, and extracellular polymeric substances, to the products of cell lysis, including sugars, proteins, lipids and nucleic acids. These organic compounds could support the cell growth of the heterotrophic species [x].
- Ma J, Guo T, Ren M, Chen L, Song X, Zhang W. Cross-feeding between cyanobacterium Synechococcus and Escherichia coli in an artificial autotrophic-heterotrophic coculture system revealed by integrated omics analysis. Biotechnol Biofuels Bioprod. 2022 Jun 22;15(1):69. doi: 10.1186/s13068-022-02163-5. PMID: 35733176; PMCID: PMC9219151.
SECTION 1: ABSTRACT
- Provide a concise, self-contained summary of your project (minimum 150 words). The abstract should allow a reader to understand the purpose, approach, and expected outcomes of the work without referring to other sections.
- Your abstract should briefly address the following elements: Significance: What problem or question does the project address, and why is it important?
Broad Objective: What is the overall goal of the project? Hypothesis: What prediction or principle is the project testing or demonstrating? Specific Aims: What key steps or milestones will be completed to achieve the objective? Methods: What experimental or technical approaches will be used?
Project in Developmental Stages
Based on feedback from week seven, this project will take shape into one of two forms.
- Working with synthetically designed E. Coli chromoproteins and studio glass technology, this project will culture hybrid images by combining E. Coli with cyanobacteria.
- Project will theoretically combine cultures within a competitive environment where they will coexist in a dynamic interaction.
- The outcome is completely unknown and experimental where observation of interaction is the focus of research.
- Project will incorporate Opentron technology to seed negative shapes with E. Coli in a precultured cyanobacter designed matrix.
- Similarly, this project works with E.Coli to generate advanced imagery using topographical glass imaging plates to create hybrid compositions in concert with bacteria to generate new forms of interaction between visual materials, and the dynamic play of light between them.
- Project will employ use of Opentron technology to generate base layers of E. Coli onto glass plates prepared with Agrose media.
- The multi level stratification of image layers will be enhanced by illumination from multiple sources and interpreted using Reflecitive Transmisive Imaging.