Week 2 HW: DNA Read Write and Edit
In silico digests
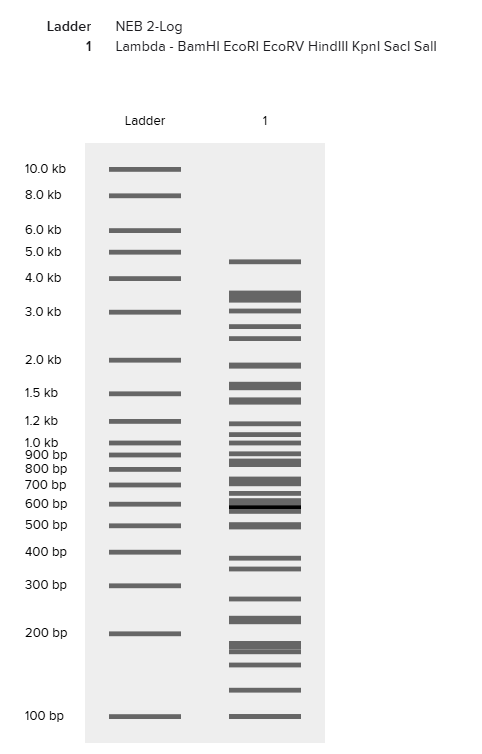
Digestion with restriction enzymes yields DNA fragments of varied size. Running gel electrophoresis with a selection of enzymes can produce artwork.
DNA design challenge
3.1) I have selected YugO, a bacterial potassium channel involved in metabolically mediated electrical signaling in B. subtilis. I am interested in this protein as it relates to my work exploring engineered electrical signaling behaviors in bacteria. Per UniProt, the sequence for this protein is : MKSNRIFISWLRWPLFIRIGVIILCLILLFGQIIYILEPKQFTSVFEGIWWAVVTVSTVGYGDYVPHTPLGQAAGILLILSGASFVTAYFATLSAAAFSRQHRYIEGKVAYKGRDHIILIGWNEKTNRLLKDLQLAAPSKTVVLIDESLTEGPLIENVHFIRGHAADDGTLKRANITEAESVMITADQYKSETDADMLSVLTLLSVKGLNPLAYCIVEILTDRFVTNAERAGANQIIGTSEFISRAMLQHYQVKLRPSKQQNGIKLTLDQHVELLAVPDELKGAAYKTCVLYFLDHNTTIIGIQKKEGPMLSPPLTYKVLETDQFLAI
3.2) The nucleotide sequence for this protein is:
atgaaaagcaaccgcatttttattagctggctgcgctggccgctgtttattcgcattggc
gtgattattctgtgcctgattctgctgtttggccagattatttatattctggaaccgaaa
cagtttaccagcgtgtttgaaggcatttggtgggcggtggtgaccgtgagcaccgtgggc
tatggcgattatgtgccgcataccccgctgggccaggcggcgggcattctgctgattctg
agcggcgcgagctttgtgaccgcgtattttgcgaccctgagcgcggcggcgtttagccgc
cagcatcgctatattgaaggcaaagtggcgtataaaggccgcgatcatattattctgatt
ggctggaacgaaaaaaccaaccgcctgctgaaagatctgcagctggcggcgccgagcaaa
accgtggtgctgattgatgaaagcctgaccgaaggcccgctgattgaaaacgtgcatttt
attcgcggccatgcggcggatgatggcaccctgaaacgcgcgaacattaccgaagcggaa
agcgtgatgattaccgcggatcagtataaaagcgaaaccgatgcggatatgctgagcgtg
ctgaccctgctgagcgtgaaaggcctgaacccgctggcgtattgcattgtggaaattctg
accgatcgctttgtgaccaacgcggaacgcgcgggcgcgaaccagattattggcaccagc
gaatttattagccgcgcgatgctgcagcattatcaggtgaaactgcgcccgagcaaacag
cagaacggcattaaactgaccctggatcagcatgtggaactgctggcggtgccggatgaa
ctgaaaggcgcggcgtataaaacctgcgtgctgtattttctggatcataacaccaccatt
attggcattcagaaaaaagaaggcccgatgctgagcccgccgctgacctataaagtgctg
gaaaccgatcagtttctggcgatt
3.3) I need to optimize codon usage depending on the organism I will be expressing this protein in, because different organisms have different codon preferences. I have selected B. subtilis 168 as this is the organism I am working with in the lab.
Codon optimized sequence:
ATG AAA TCA AAC AGA ATC TTC ATC TCT TGG TTG CGT TGG CCG CTG TTT ATT CGG ATT GGA GTG ATC ATC CTG TGC CTG ATA TTG CTC TTT GGA CAA ATT ATC TAT ATC TTG GAG CCG AAA CAG TTT ACA AGC GTT TTC GAG GGG ATT TGG TGG GCC GTT GTC ACA GTG TCA ACA GTC GGG TAC GGG GAC TAT GTC CCT CAT ACC CCT TTA GGA CAG GCG GCC GGC ATT TTA CTT ATA CTT AGC GGG GCG TCA TTT GTG ACA GCA TAT TTC GCG ACG CTG TCT GCG GCT GCT TTC AGT CGG CAG CAT CGC TAC ATT GAA GGA AAA GTC GCA TAT AAG GGC AGA GAT CAT ATC ATC CTC ATA GGA TGG AAT GAG AAA ACC AAT CGG TTA CTT AAA GAC CTT CAA TTA GCG GCT CCT AGT AAA ACA GTA GTG CTT ATC GAT GAA TCA CTG ACG GAA GGC CCA CTG ATC GAG AAC GTG CAT TTC ATC CGG GGC CAT GCG GCG GAT GAT GGC ACG CTC AAA CGG GCG AAC ATT ACG GAA GCG GAG AGC GTT ATG ATT ACA GCT GAT CAG TAC AAA TCA GAA ACA GAT GCC GAT ATG CTT AGC GTA CTG ACA TTA CTC TCA GTG AAA GGC CTT AAT CCG TTA GCA TAC TGT ATT GTC GAG ATT CTC ACC GAT AGA TTT GTG ACG AAC GCC GAG AGA GCG GGC GCC AAC CAG ATT ATC GGC ACA TCT GAA TTT ATT TCC CGC GCT ATG CTT CAG CAT TAC CAA GTT AAA CTG CGT CCG AGC AAA CAA CAG AAC GGC ATT AAG TTG ACG CTG GAC CAA CAC GTT GAA TTA TTA GCG GTT CCG GAT GAA TTA AAA GGG GCG GCG TAT AAG ACG TGT GTC CTG TAT TTC CTC GAT CAT AAT ACA ACT ATT ATT GGG ATT CAA AAA AAA GAG GGT CCT ATG TTG AGC CCG CCA TTA ACC TAT AAA GTG TTG GAA ACA GAT CAA TTT CTT GCT ATT
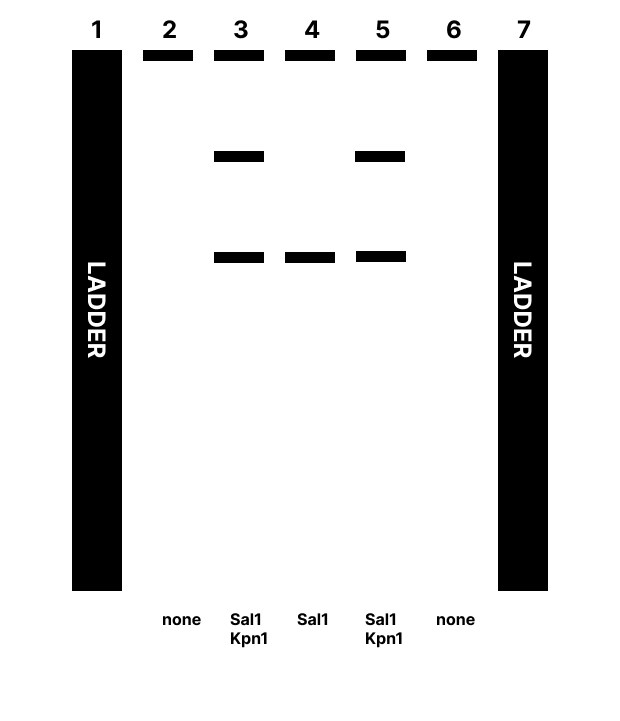
3.4) In this case I am intending to express this as a functional protein in a living community (this particular sequence is already native to B. subtilis 168, I am intending to include chimeric components), thus a cell dependent approach will be taken. This sequence can be synthesized, cloned into a plasmid using enzyme digestion and Gibson assembly, and transformed in competent B. subtilis 168.
Twist DNA
5.1 DNA Read
i)
ii)
5.2 DNA Write
i)
ii)
5.3 DNA Edit
i)
ii)