Week 2 HW: DNA Read, Write, & Edit
3.1 mCherry
I was thinking about observation under UV illumination — red fluorescence seems easier to detect compared to blue or green light. It also has stronger visual impact.
Protein Sequence
Source: https://www.fpbase.org/protein/mcherry/
3.2 Original DNA Sequence
3.3 Codon Optimization
Why Optimize?
The same amino acid can be encoded by multiple codons, but different organisms have different codon usage preferences (tRNA abundance, translation efficiency, mRNA structure, etc.). To allow the host to express the protein more efficiently, codon optimization is necessary.
Organism Selected
E. coli K-12.
It is one of the most commonly used host strains. The technical maturity and widespread adoption of this system make it highly suitable for experimental work.
Optimized DNA Sequence
Source: https://www.kazusa.or.jp/codon/cgi-bin/showcodon.cgi?aa=1&species=83333&style=N&utm
3.4 Cell-Dependent Expression
Goal
To have E. coli transcribe this DNA into mRNA and translate it into a fluorescent protein.
Method
Insert the optimized CDS into an expression plasmid including:
- Promoter
- RBS
- Terminator
Transform the plasmid into E. coli.
In E. coli:
- RNA polymerase recognizes the promoter
- DNA is locally unwound
- Complementary mRNA is synthesized
The ribosome binds via the Shine–Dalgarno sequence and begins translation at the start codon.
- One codon (3 nucleotides) is read at a time
- tRNAs deliver corresponding amino acids
- Amino acids are linked into a polypeptide chain
- Final product: fluorescent protein
4.1
/
4.2
https://benchling.com/s/seq-7VHbcY2Zp8vixGlBz1td?m=slm-7HxrEb6QjOZ58RtT8p96
(There’s unknown error for the optimization version,,,)


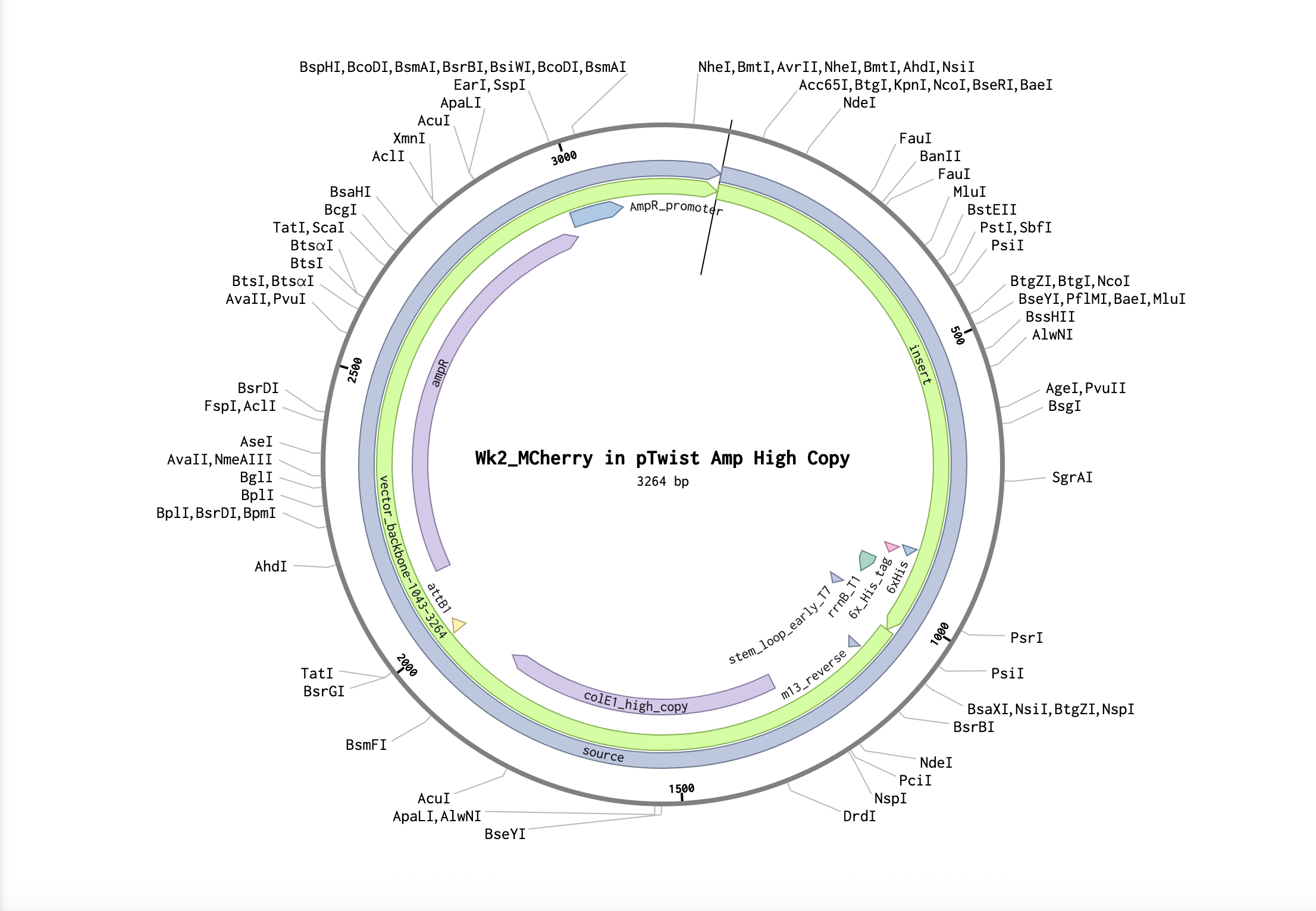

5.1 DNA Read
5.1(i) What DNA Would I Sequence?
A synthetic DNA library used for digital DNA data storage.
Artificially designed DNA fragments encoding digital information (text or images).
Why?
DNA here serves as an information storage medium rather than biological genetic material.
Sequencing verifies:
- Whether the written digital information is preserved
- Whether errors occurred during storage or amplification
- The error rate (substitutions, insertions, deletions)
This is effectively a biotechnology-based data integrity check.
5.1(ii) Sequencing Technology
First-generation sequencing: Sanger sequencing.
Why?
- High accuracy
- Suitable for validating single fragments
Characteristics
- Reads one DNA template at a time
- Read length ~700–900 bp
- Very high accuracy
Inputs
- Template DNA
- Primer
- DNA polymerase
- dNTPs
- Fluorescently labeled ddNTPs
Core Principle
DNA synthesis is terminated at random positions using ddNTPs.
Process:
- Polymerase copies the template
- ddNTP incorporation stops elongation
- Fragments of different lengths are produced
- Fragments separated by size
- Fluorescent signal read to reconstruct sequence
Output
- DNA sequence
- Chromatogram
Limitations
- Cannot sequence thousands of fragments simultaneously
5.2 DNA Write
5.2(i) What DNA Would I Synthesize?
An expression cassette expressing mCherry in E. coli.
Includes:
- Promoter
- RBS
- Codon-optimized mCherry CDS
- His tag
- Terminator
Reason:
- Produces visible red fluorescence
- Strong contrast under blue light
5.2(ii) Synthesis Technology
Solid-phase chemical DNA synthesis (phosphoramidite method) + gene assembly.
Steps
- Design sequence computationally
- Split into short oligos
- Chemically synthesize oligos
- Assemble via PCR or Gibson Assembly
- Clone into vector
- Sequence verify
Limitations
- Error rate increases with length
- Assembly required
- Sequencing verification required
- Cost scales with length
5.3 DNA Edit
5.3(i) What DNA Would I Edit?
A single-base mutation (e.g., disease-causing point mutation).
Reason:
- Represents the most precise editing scenario
- Relevant for therapeutic research
5.3(ii) Editing Technology
CRISPR-Cas9 + HDR repair template.
Principle
- Design gRNA
- Cas9 creates double-strand break
- Provide donor DNA
- HDR replaces base precisely
Required Inputs
- gRNA
- Cas9 protein or plasmid
- Donor DNA
- Target cells
Limitations
- Low HDR efficiency
- Possible off-target effects
- Complex delivery
- Cell-type dependent precision