Week 4 HW: Protine Design
Protein & Amino Acid Questions
1) How many molecules of amino acids are in 500 g of meat?
(Assume average amino acid ≈ 100 Da ≈ 100 g/mol)
Upper-bound estimate: treat the 500 g as entirely amino acids.
- Moles of amino acids ≈ 500 g / (100 g/mol) = 5 mol
- Number of molecules ≈ 5 mol × 6.02 × 10²³ molecules/mol
- ≈ 3.0 × 10²⁴ amino acid molecules
2) Why do humans eat beef but do not become a cow?
Dietary proteins are not incorporated intact into our bodies.
During digestion:
- Proteins are broken down into amino acids (and small peptides).
- Cells use those amino acids as generic building blocks.
- Human proteins are synthesized according to human DNA instructions.
Food provides raw materials.
The blueprint comes from our genome and physiology.
3) Why are there only 20 natural amino acids?
Evolution selected 20 genetically encoded amino acids because they balance:
- Chemical diversity
- System simplicity
They provide:
- Hydrophobic residues → core formation
- Polar/charged residues → solubility & catalysis
- Special residues:
- Glycine → flexibility
- Proline → rigidity
- Cysteine → disulfide bonds
Adding more amino acids would require:
- More complex tRNAs
- More aminoacyl-tRNA synthetases
- Increased ribosomal complexity
- Higher error rates
- Greater biological cost
4) Can you make non-natural amino acids?
Yes.
Non-natural amino acids (nnAAs) can be incorporated via:
- Engineered tRNA/synthetase systems
- Chemical synthesis + feeding strategies
They expand protein functionality beyond the natural 20.
Example conceptual designs:
Fluorinated hydrophobic amino acid
- Leucine-like residue with fluorination
- Tunes hydrophobicity & packing
- Can increase stability
- Can modify binding interfaces
5) If you build an α-helix using D-amino acids, what handedness results?
Natural proteins use L-amino acids → right-handed α-helices.
Using D-amino acids:
- Produces the mirror image
- Expected structure: left-handed α-helix
Reason: Switching chirality flips backbone angle preferences and hydrogen-bond geometry.
6) Why are most molecular helices right-handed?
Because biology overwhelmingly uses L-amino acids.
Given L-amino acids:
- Steric constraints favor right-handed helices
- Hydrogen-bond geometry stabilizes this orientation
This reflects:
- Stereochemical constraints
- Evolutionary fixation of chirality
7) Why do β-sheets aggregate?
β-sheet hydrogen bonding is inherently extendable.
Driving forces:
- Backbone hydrogen bonding
- Hydrophobic effect
- Shape complementarity / stacking
If strand edges are exposed:
- Additional strands dock
- Sheets extend
- Fibrils grow
8) Why do amyloid diseases involve β-sheets?
Misfolded proteins expose backbone segments that form β-strands.
These assemble into:
- Extended β-sheet-rich fibrils
- Highly stable, ordered aggregates
Consequences:
- Cellular toxicity
- Sequestration of functional proteins
- Stress on quality-control systems
9) Can amyloid β-sheets be used as materials?
Yes.
Amyloid assemblies are:
- Mechanically robust
- Highly ordered
- Self-assembling
Applications:
- Nanofiber scaffolds
- Hydrogels
- Templates for nanomaterials
- Functional biomaterials
Selected Protein Analysis
1️⃣ Protein Selection
Protein: NanoLuc luciferase
Derived from: Oplophorus gracilirostris
Function:
- Catalyzes chemiluminescent reaction
- Produces bright blue light
Why selected:
- Core engine for “living / glowing ice cream” concept
- Converts chemical energy → light
- Small, bright, structurally characterized
- Ideal computational design test case
2️⃣ Amino Acid Sequence
(Chain A, PDB 5IBO)
- SDNMVFTLEDFVGDWRQTAGYNLDQVLEQGQQNLWDLNTEKQQLQKSLQDLKNEEVDMVLNNKSSN GQWFDVVKQKGGFVDGRTKFVGTNQGSLLGYYKDSDQLKTTNIKQVVSTLQGQKIDGTTLTQLNKE VLDNLVTTNPKLREKFQVVHQLLEDTQGTMNQKDTNRTV
3️⃣ Length & Composition
- Length: 174 amino acids
- Most frequent residue: Glycine (G)
5️⃣ Protein Family
NanoLuc belongs to:
- Luciferase family
- Oxidoreductases
Function:
- Catalyzes oxidative reactions
- Chemical energy → photon emission
6️⃣ Structure Page
PDB ID: 5IBO
(RCSB structure page referenced)
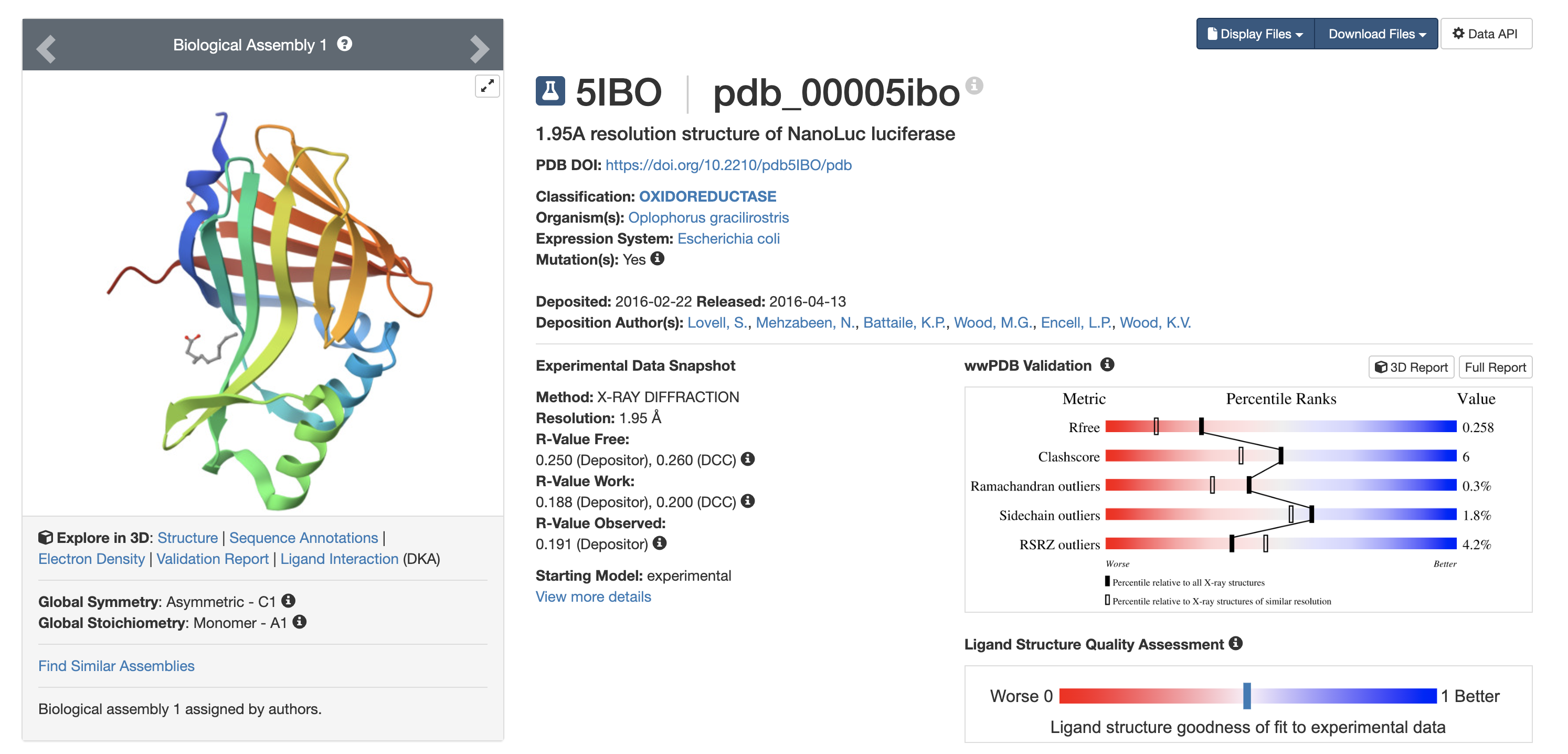

7️⃣ Structure Quality
- Method: X-ray crystallography
- Resolution: 1.95 Å
- Released: 2016
Resolution < 2.5 Å = high-quality structure
1.95 Å is considered reliable.
8️⃣ Other Molecules in Structure
Yes.
Bound ligand: Decanoic acid
Why this matters:
- Indicates defined binding pocket
- Active site accommodates hydrophobic molecules
Design implications for ice cream:
- Does luciferin need micro-encapsulation?
- Does fat content affect substrate diffusion?
- Could dairy fat interact with hydrophobic pocket?
Structural data → formulation constraints.
🔟 PyMOL Visualization
Observation:
- Beta-sheet dominant structure
- Consistent with beta-barrel fold
Interpretation:
- Beta-rich cores = tightly packed
- Likely structurally stable
Implication: May retain activity under:
- Slight warming
- Partial melting
- Oxygen exposure
1️⃣1️⃣ Residue Surface Analysis
Hydrophobic residues:
- Cluster in interior
Polar/charged residues:
- Dominant on surface
Interpretation:
- Hydrophobic core → structural stability
- Hydrophilic surface → water compatibility
From ice cream perspective:
- Ice cream = water + fat + sugar
- Hydrophilic exterior = favorable
- Hydrophobic core remains shielded
Encouraging for enzyme stability in food matrix.