Week 1 HW: Principles and Practices
Project Title: BAP – The Bits and Atoms Pen
BAP (Bits and Atoms Pen) is a hybrid tool that integrates digital fabrication interfaces with biological microfluidic systems into a single handheld device. Originally conceived during the HTGAA program in 2017, the project explores the convergence of computation, fabrication, and living systems. For my final project, I plan to further develop BAP conceptually and experimentally as a platform that bridges digital control (“bits”) and biological matter (“atoms”).
On one side, BAP functions as a digital interface tool. It includes a nib compatible with touch screens, spatial sensors, and a microcontroller that enables communication with computers and digital fabrication machines. The device acts as both an input tool and a physical extension of design software, allowing users to interact seamlessly with fabrication systems.
On the opposite side, BAP incorporates a conceptual microfluidic architecture capable of sustaining engineered bacteria within isolated microchannels. These bacteria would produce primary color proteins (e.g., red, green, blue). Through controlled microfluidic mixing, the device could generate a full visible color spectrum in real time. This same principle could be extended beyond pigments: microfluidic channels could mix therapeutic compounds, enabling on-demand pharmaceutical formulation or customized biochemical outputs.
The project is not intended as an immediate deployable medical device, but rather as a speculative and research-driven platform exploring how programmable biology and digital fabrication might coexist safely within designed objects. The publication will focus on design principles, biosafety considerations, microfluidic feasibility, containment strategies, and ethical implications of embedding living systems into everyday tools.
Ultimately, BAP aims to provoke new conversations about how humans may responsibly design tools that operate simultaneously in digital and biological domains, while prioritizing biosecurity, environmental protection, and safe laboratory practices.
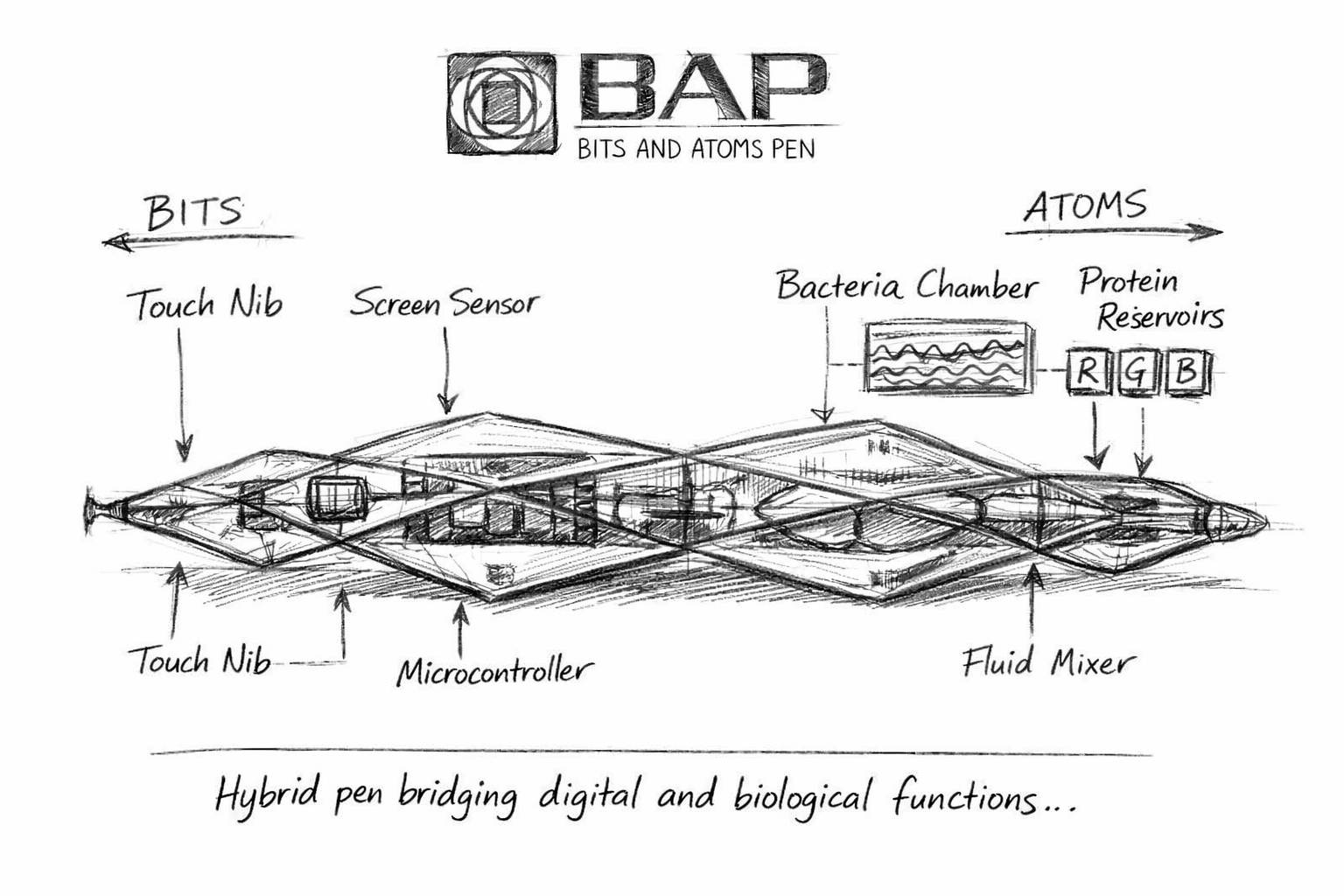
 Conceptual sketch of the BAP (Bits and Atoms Pen). This image was generated using artificial intelligence based on an original visual concept created by the author (2017). The AI-assisted rendering was used to reinterpret the initial design as a preliminary sketch for publication purposes.
Conceptual sketch of the BAP (Bits and Atoms Pen). This image was generated using artificial intelligence based on an original visual concept created by the author (2017). The AI-assisted rendering was used to reinterpret the initial design as a preliminary sketch for publication purposes.
Proposed options for the final project direction:
Option 1: Conceptual Design + Risk Framework Publication
Option 2: Prototype with Non-living Safe Color Fluids (No bacteria)
Option 3: Contained Microfluidic Prototype Using Non-pathogenic Strains (BSL1, strictly laboratory-based)
| Does the option: | Option 1 – Conceptual | Option 2 – Non-living Fluids | Option 3 – BSL1 Lab Prototype |
|---|
| Enhance Biosecurity | High | High | Medium (requires oversight) |
| • By preventing incidents | Strong (framework) | Strong (no living systems) | Moderate (containment needed) |
| • By helping respond | Conceptual guidance | Limited | Moderate (lab protocols) |
| Foster Lab Safety | High | High | Medium–High |
| • By preventing incident | Yes (risk mapping) | Yes | Yes (if BSL1 compliant) |
| • By helping respond | Theoretical | Minimal | Practical protocols |
| Protect the environment | Strong | Strong | Moderate (needs waste control) |
| • By preventing incidents | Yes | Yes | Yes (with sterilization) |
| • By helping respond | Ethical framing | Limited | Requires SOPs |
| Other considerations | | | |
| • Minimizing costs and burdens to stakeholders | Very High | High | Lower (lab costs) |
| • Feasibility? | Very High | High | Medium |
| • Not impede research | Yes | Yes | Possibly regulated |
| • Promote constructive applications | Strong conceptual | Strong design demo | Strong research potential |
| • Ethical clarity | Very strong | Strong | Requires bioethics review |
| • Dual-use risk | Minimal | Minimal | Must be evaluated |
| • Regulatory complexity | None | Low | Moderate |
Homework questions
Professor Jacobson
- What is the polymerase error rate? How does this compare to the length of the human genome? How does biology address this discrepancy?
DNA polymerase with proofreading has an approximate error rate of about 1 in 10⁶ nucleotides. When I compare this to the size of the human genome (around 3.2 billion base pairs), it becomes clear that polymerase fidelity alone would not be sufficient. Without additional correction systems, thousands of errors could occur during one full genome replication.
Biology addresses this through multiple layers of error correction. Polymerases can proofread their own work, and after replication there are repair systems such as mismatch repair that detect and fix remaining mistakes. In addition, biological systems are relatively robust and can tolerate some mutations. Together, these mechanisms reduce the effective mutation rate to a level compatible with long-term genome stability.
- How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice, why do many of these different codes fail to encode the protein properly?
Because there are 64 codons for only 20 amino acids, the genetic code is redundant. For a protein of about 345 amino acids, there are an astronomically large number of possible DNA sequences that could encode the same protein.
However, not all of these sequences function equally well. Factors such as codon bias, GC content, RNA secondary structure, unintended regulatory signals, and synthesis constraints can affect gene expression. Even if two DNA sequences encode the same amino acid chain, they may behave differently in a biological system.
Dr. LeProust
- What is the most commonly used method for oligonucleotide synthesis?
The most commonly used method is solid-phase phosphoramidite synthesis. This is a chemical process that builds DNA step by step by adding one nucleotide at a time in repeated cycles.
- Why is it difficult to produce oligonucleotides longer than 200 nucleotides by direct synthesis?
Each synthesis cycle is highly efficient but not perfect. Since each step has slightly less than 100% efficiency, small errors accumulate as the sequence length increases. As a result, the percentage of full-length, correct oligonucleotides drops significantly beyond about 200 nucleotides.
- Why can’t a 2000 bp gene be created by direct oligonucleotide synthesis?
A 2000 base pair gene cannot be synthesized directly as a single oligonucleotide because the accumulated errors and truncated products would be too high. Instead, long genes are assembled from shorter oligos using methods such as PCR assembly or Gibson assembly.
Professor George Church
- What are the 10 essential amino acids in all animals? How does this affect your view of the “Lysine Contingency”?
The essential amino acids in animals are histidine, isoleucine, leucine, lysine, methionine, phenylalanine, threonine, tryptophan, and valine (and in some contexts arginine). These must be obtained from the diet because animals cannot synthesize them.
The idea of a “Lysine Contingency” highlights how metabolic dependence can be used as a containment strategy. Since lysine is essential, engineering organisms that depend on external lysine availability could act as a biological safety mechanism. I see this as a practical way of using natural biological limits to improve biosafety in synthetic systems.


