For this task, I wanted to do something that was both beautiful and scientific (and programmatic, since I do have some knowledge of that), so I chose to draw a fractal, since they are often found in nature as well (alveolar hierarchy, snowflakes, plants). A fractal that I have always liked aesthetically, as well as being easy to draw in Python, is the Sierpinski triangle.
I made different drawings, with different cycles, varying the volume dispensed. Some of the volumes dispensed are unrealistic, but I wanted to see the fractal with more iterations. To construct the triangle and the different iterations, I simply drew the vertices of each triangle, recursively calculating the positions of the vertices of the internal triangles by taking the midpoints of each side. A limitation of my code at the moment is that in some cases it dispenses bioink multiple times in the same position if a vertex is part of two triangles at the same time.
from opentrons import types
import math
metadata = { # see https://docs.opentrons.com/v2/tutorial.html#tutorial-metadata
'author': 'Rodrigo Antonio Arredondo Garza',
'protocolName': 'Fractal Life',
'description': 'Draw a Sierpinski triangle parametrically (Change iterations and dimensions easily).',
'source': 'HTGAA 2026 Opentrons Lab',
'apiLevel': '2.20'
}
##############################################################################
### Robot deck setup constants - don't change these
##############################################################################
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
well_colors = {
'A1' : 'Red',
'B1' : 'Green',
'C1' : 'Orange'
}
def run(protocol):
##############################################################################
### Load labware, modules and pipettes
##############################################################################
# Tips
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
# Pipettes
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
# Modules
temperature_module = protocol.load_module('temperature module gen2', COLORS_DECK_SLOT)
# Temperature Module Plate
temperature_plate = temperature_module.load_labware('opentrons_96_aluminumblock_generic_pcr_strip_200ul',
'Cold Plate')
# Choose where to take the colors from
color_plate = temperature_plate
# Agar Plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate') ## TA MUST CALIBRATE EACH PLATE!
# Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
##############################################################################
### Patterning
##############################################################################
###
### Helper functions for this lab
###
# pass this e.g. 'Red' and get back a Location which can be passed to aspirate()
def location_of_color(color_string):
for well,color in well_colors.items():
if color.lower() == color_string.lower():
return color_plate[well]
raise ValueError(f"No well found with color {color_string}")
# For this lab, instead of calling pipette.dispense(1, loc) use this: dispense_and_detach(pipette, 1, loc)
def dispense_and_detach(pipette, volume, location):
"""
Move laterally 5mm above the plate (to avoid smearing a drop); then drop down to the plate,
dispense, move back up 5mm to detach drop, and stay high to be ready for next lateral move.
5mm because a 4uL drop is 2mm diameter; and a 2deg tilt in the agar pour is >3mm difference across a plate.
"""
assert(isinstance(volume, (int, float)))
above_location = location.move(types.Point(z=location.point.z + 5)) # 5mm above
pipette.move_to(above_location) # Go to 5mm above the dispensing location
pipette.dispense(volume, location) # Go straight downwards and dispense
pipette.move_to(above_location) # Go straight up to detach drop and stay high
###
### YOUR CODE HERE to create your design
###
# Don't forget to end with a drop_tip()
def sierpinski(side_length, depth, color_string):
MAX_RADIUS = 38
drop_counter = 0
max_drops = 20
h = side_length * math.sqrt(3) / 2
p1 = (-side_length/2, -h/3)
p2 = (side_length/2, -h/3)
p3 = (0, 2*h/3)
triangles = []
def subdivide(a, b, c, d):
if d == 0:
triangles.append((a, b, c))
return
ab = midpoint(a, b)
bc = midpoint(b, c)
ca = midpoint(c, a)
subdivide(a, ab, ca, d-1)
subdivide(ab, b, bc, d-1)
subdivide(ca, bc, c, d-1)
def midpoint(p, q):
return ((p[0]+q[0])/2, (p[1]+q[1])/2)
subdivide(p1, p2, p3, depth)
pipette_20ul.pick_up_tip()
nonlocal_drop = {'count': 0}
def safe_dispense(x, y):
r = math.sqrt(x**2 + y**2)
if r > MAX_RADIUS:
return
if nonlocal_drop['count'] % max_drops == 0:
pipette_20ul.aspirate(20, location_of_color(color_string))
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_detach(pipette_20ul, 1, adjusted_location)
nonlocal_drop['count'] += 1
for a, b, c in triangles:
for p in [a, b, c]:
safe_dispense(p[0], p[1])
pipette_20ul.drop_tip()
sierpinski(
side_length=65,
depth=5,
color_string='Orange'
)Visual Outputs
BIOPRINTED PATTERNS
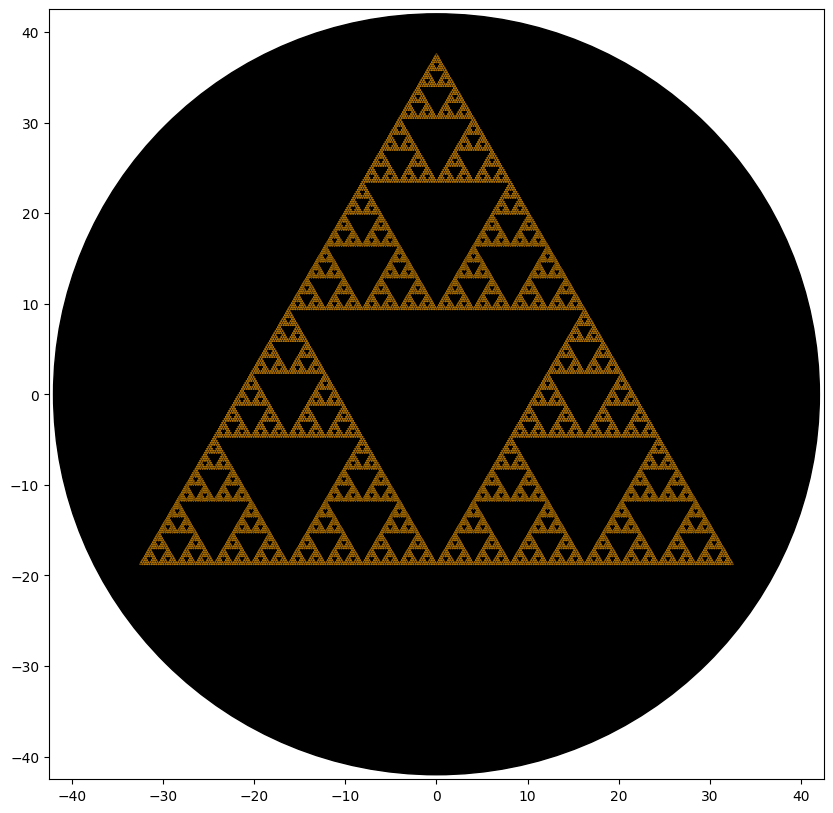

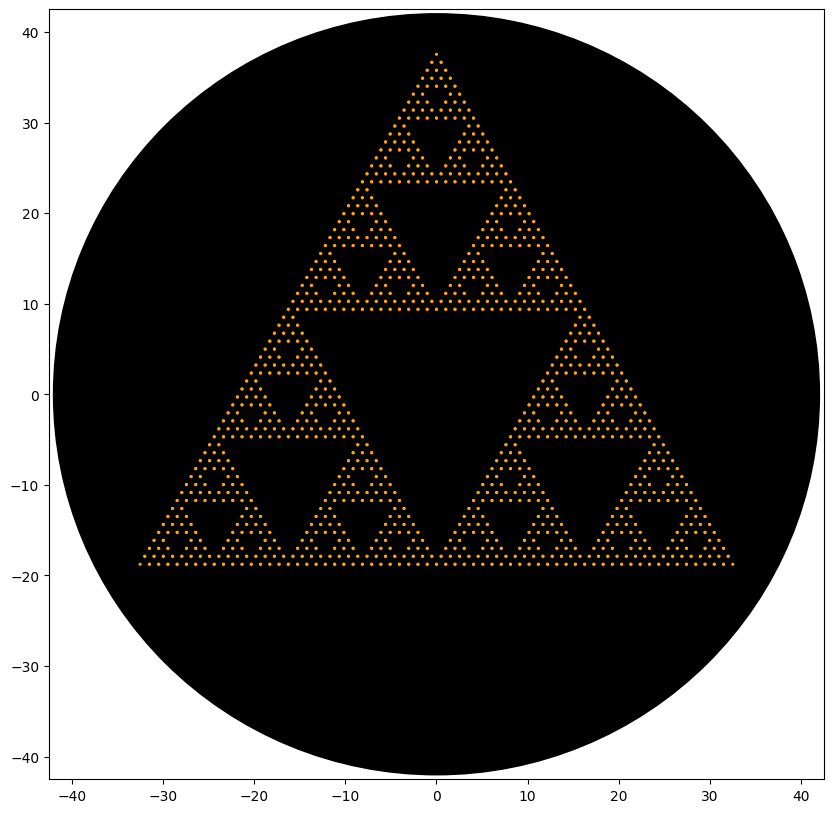

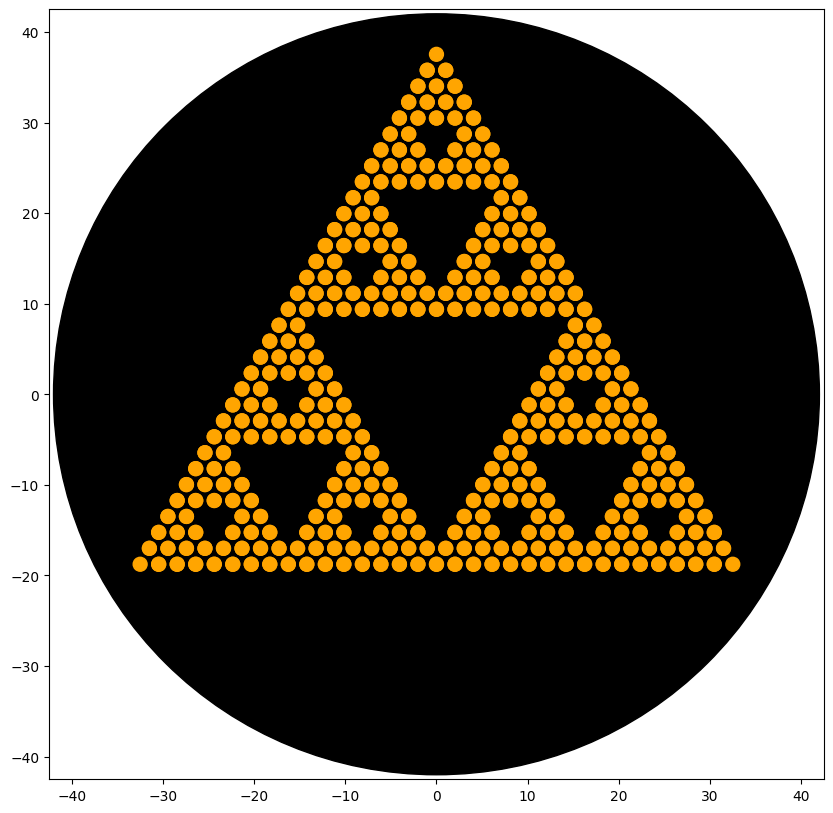

Post-Lab Analysis
I am describing a recent collaboration between Ginkgo and OpenAI, detailed in their announcement: GPT-5 Lowers Protein Synthesis Cost.
In this study, they integrate OpenAI's GPT-5 model with Ginkgo's cloud lab to optimize cell-free protein synthesis. The system forms a closed-loop control system in which an AI agent designs experiments that are automatically executed by the robotic infrastructure.
The results of these experiments are seamlessly returned to the model for continuous analysis and hypothesis generation. With this automated workflow, they achieved a remarkable 40% reduction in costs and a 27% increase in cell titer compared to the state of the art.
Much of the idea (and I suppose the ambitious part) of my final project is based on automation. I would like to be able to automate everything from experimental design to the execution of the experiments themselves. To do this, I plan to incorporate an AI agent (probably using frameworks such as Langgraph or Langchain) that would function as an experimental planner. It would propose experiments based on analysis of results and a world model, allowing it to decide on subsequent actions in a completely autonomous manner.
The agent will generate structured experiments by defining the drugs to be used, concentrations, combinations, and exposure times. It will autonomously create its own code to interact with the automated laboratory through a specialized API. This system requires an infrastructure that enables the automation of cell culture (media change, incubation, plating, etc.), reagent dispensing, and testing (viability, imaging, etc.). Using Ginkgo Nebula for this project seems to be a strong fit, as the AI agent could design the experiments with structured output (like Pydantic or JSON Schemas) supported perfectly by their API.
To provide a more realistic option for the scope of the course, instead of using iPSCs, E. coli bacteria equipped with a fluorescent stress reporter could be used, which would be evaluated and analyzed by the agent. Based on the results, the agent would design experiments iteratively, updating a probabilistic model that maps the dose/response relationship for individual compounds and combinations of compounds. The model used would be a Gaussian Process Regression, which is highly effective for small amounts of data and in situations where quantifying uncertainty is beneficial (allowing us to perform active learning).