Week 3 HW: Lab Automation
Opentrons Artwork

Post-Lab Questions
Question 1.
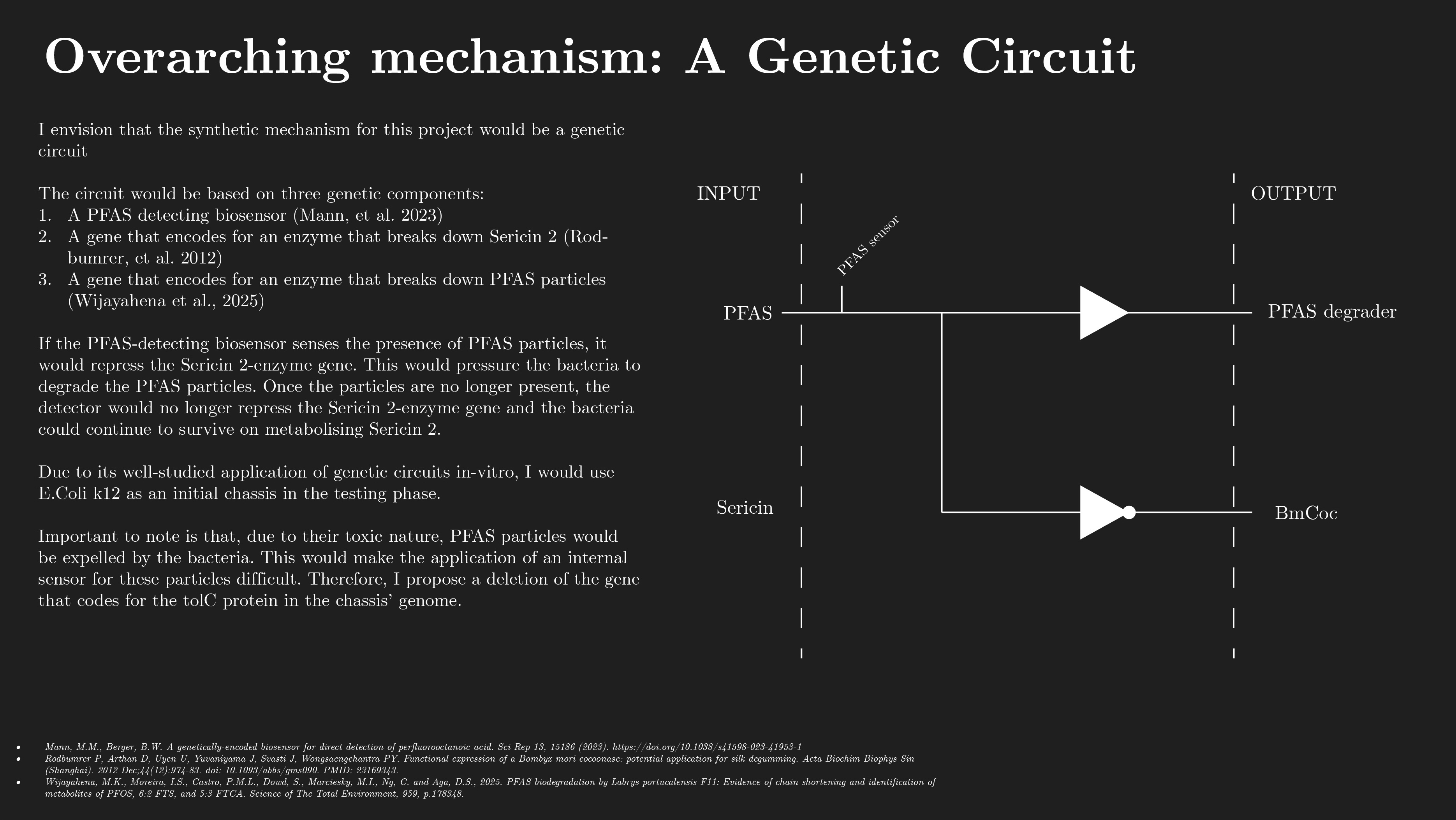
My chosen article, entitled ‘AssemblyTron: flexible automation of DNA assembly with Opentrons OT-2 lab robots’, details novel development within the field of synthetic biology and automated lab operations in regards to DNA assembly.
Compared to other steps in the DBTL cycle of synbio, building has progressed the least in terms of automation, optimization and algorithmical performance. This step often involves DNA assembly, which continues to be done manually, with low throughput and often unreliable results.
The article presents AssemblyTron; an open-source Python package to integrate j5 DNA assembly design software outputs with build implementations in Opentrons liquid handling robotics with minimal human intervention.
It describes how AssemblyTron is capable of the following:
Several scarless, multipart DNA assemblies, beginning from fragment amplification.
Polymerase chain reactions across a range of fragment lengths and annealing temperatures by utilising an optimal annealing gradient calculation algorithm.
Golden Gate and homology dependent in vivo assemblies.
Site-directed mutagenesis reactions via homology-dependent IVA
John A Bryant, Mason Kellinger, Cameron Longmire, Ryan Miller, R Clay Wright, AssemblyTron: flexible automation of DNA assembly with Opentrons OT-2 lab robots, Synthetic Biology, Volume 8, Issue 1, 2023, ysac032, https://doi.org/10.1093/synbio/ysac032
Question 2.
For this assignment I decided to focus on one of my three final graduation project, investigating the possibilities to utilize Opentrons to safely and efficiently conduct lab research:
Final project option 2 - Isolating the DNA sequence that codes for the enzyme responsible for PFAS degradation in L. portucalensis F11
For this option, if I could get access to F11, I would use opentrons to inoculate 96-well plates with the bacteria in a medium solution that contains PFAS as the sole carbon source. I would then measure OD600 timepoints to determine cell density (Myers, et al. 2020) to find the optimal timing for RNA extraction.
When the optimal timing for RNA extraction is reached, I could then automate the cell lysing process.
For the extraction of excessive rRNA, I would automate the pipetting process of the hybridisation of DNA probes to the rRNA. Then, using the Opentrons magnetic module (Opentrons, 2026), I could remove the rRNA by using streptavidin-coated magnetic beads.
Finally, I would utilise opentrons automation to library prep for Oxford Nanopore Sequencing.
References:
Myers, J.A., Kitajima, M., Dai, L.C., & Wright, M.S. (2020) 'Robust estimation of bacterial cell count from optical density', Communications Biology, 3(1), p. 446. Available at: https://doi.org/10.1038/s42003-020-01127-5 (Accessed: 24 February 2026).
Opentrons (2026). Opentrons Flex Magnetic Block. [online] Opentrons.com. Available at: https://opentrons.com/products/opentrons-flex-magnetic-block-gen1?utm_source=google&utm_medium=cpc&utm_campaign={campaignname}&utm_term=&hsa_cam=21988025885&hsa_grp=&hsa_mt=&hsa_src=x&hsa_ad=&hsa_acc=2303351826&hsa_net=adwords&hsa_kw =&hsa_tgt=&hsa_ver=3&gad_source=1&gad_campaignid=21981643080&gbraid=0AAAAADemOsWv5oA02ZBRucCPg3Hfq31a8&gclid=Cj0KCQiAtfXMBhDzARIsAJ0jp3DelnoEGaaIAaFZozVi7Rp9gGhAOWiMezpLkMCDgPqzlIMIj1niM3oaAvkeEALw_wcB
Oxford Nanopore Technologies (2026) Platform technology. Available at:
https://nanoporetech.com/platform/technology (Accessed: 24 February 2026)
Final Project Ideas