Week 3 HW: Lab automation
Homework Week 3
Post-Lab Questions
One of the great parts about having an automated robot is being able to precisely mix, deposit, and run reactions without much intervention, and design and deploy experiments remotely. For this week, we’d like for you to do the following:
1. Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Initially tried to do a Sonic 3 & Knuckles logo (Classic sonic game) silhouette. The design with some tweaks was hopeful. In the end, I went with the Batman Beyond logo, as it was simple and only had one colour (given the limitations of our node). The final code (with the help of Gemini) I used is below:
*### *### YOUR CODE HERE to create your design
############################################################################## Simple Design: Batman Beyond Logo ##############################################################################
spacing = 1.7 design_points = []
#* We use absolute values abs(i) to ensure perfect left-right symmetry for i in range(-18, 19): # Horizontal span (~72mm total) for j in range(-15, 12): # Vertical span
x = abs(i)
# 1. Top Wing Edge (slopes up to the points)
if j < (0.5 * x) + 3:
# 2. Bottom "V" Shape
if j > (1.2 * x) - 16:
# 3. Inner Wing Cutouts (The 'U' shapes next to the head)
# If we are not in the cutout zone, add the point
is_cutout = (2 < x < 7) and (j > -2)
# 4. The Head (Center spike)
is_head = (x <= 1) and (j < 5)
if not is_cutout or is_head:
design_points.append((i * spacing, j * spacing, 'Red'))
#* EXECUTION points_for_color = [p for p in design_points if p[2] == ‘Red’]
if points_for_color: pipette_20ul.pick_up_tip() pipette_20ul.aspirate(15, location_of_color(‘Red’))
for x, y, c in points_for_color:
if pipette_20ul.current_volume < 0.5:
pipette_20ul.aspirate(15, location_of_color('Red'))
target = center_location.move(types.Point(x=x, y=y))
dispense_and_detach(pipette_20ul, 0.5, target)
pipette_20ul.drop_tip()
############################################################################## END OF CODE ##############################################################################
The output:

With some manual tweaks, what I initially wanted to do:

However, the end product was pretty atrocious and would take way too much time to fix, given I’m doing this in the last minute. So atrocious that I won’t paste it.
2. Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
While your description/project idea doesn’t need to be set in stone, we would like to see core details of what you would automate. This is due at the start of lecture and does not need to be tested on the Opentrons yet.
(Answer 1)
Paper: An Automated Versatile Diagnostic Workflow for Infectious Disease Detection in Low-Resource Settings
Miren Urrutia Iturritza, Phuthumani Mlotshwa, Jesper Gantelius, Tobias Alfvén, Edmund Loh, Jens Karlsson, Chris Hadjineophytou, Krzysztof Langer, Konstantinos Mitsakakis, Aman Russom, Håkan N. Jönsson, Giulia Gaudenzi
https://doi.org/10.3390/mi15060708
This paper describes how researchers built an automated diagnostic workflow for detection of infectious diseases in low-resource settings [1]. Specifically, they tested for Neisseria meningitidis; a gram-negative bacterium that cause serious meningitis and blood infections in humans.
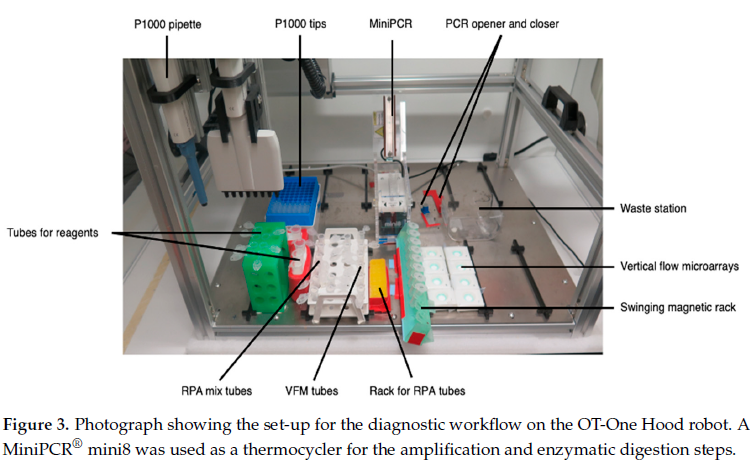
For their workflow, they used Opentrons OT-One-S Hood. This is an open-source liquid handling robot, which can be bought at a relatively low cost. The researchers wrote custom software developed at SciLifeLab Nanobiotechnology division [2] to create scripts for their workflow.
Materials and reagents were organized onto the OT-One-S Hood robot, with racks and tubes with primers, buffers, and enzymes, the MiniPCR® mini8 thermal cycler, magnetic bead racks, waste containers, and microarray holders, to analyze Neisseria meningitidis DNA in both clinical and spiked samples. “Clinical” samples refere to specimens collected from individuals, where “spiked” samples were lab prepared samples where a known amount of Neisseria meningitidis DNA.
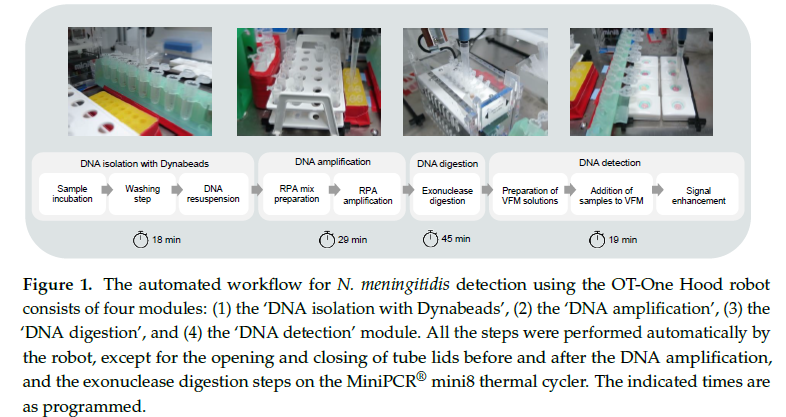
The robot then performs all the necessary pipetting steps, RNA amplication of ctrA gene (as its conserved, species-specific gene essential for capsule formation, making it a reliable marker [3]), enzymatic digestion, and deposition onto paper-based microarrays. The only manual steps were the opening and closing of tube lids before and after the DNA amplification, and the exonuclease digestion steps on the MiniPCR® mini8 thermal cycler [1].
The study showed that, automated liquid handling can detect Neisseria meningitidis in low-resource settings, though accuracy and reproducibility were not fully validated.




References
Urrutia Iturritza M, Mlotshwa P, Gantelius J, Alfvén T, Loh E, Karlsson J, Hadjineophytou C, Langer K, Mitsakakis K, Russom A, et al. An automated versatile diagnostic workflow for infectious disease detection in low-resource settings. Micromachines. 2024;15(6):708. doi:10.3390/mi15060708.
Langer K, Joensson HN. Rapid production and recovery of cell spheroids by automated droplet microfluidics. SLAS Technol. 2020;25:111–122.
Rivas L, Reuterswärd P, Rasti R, Herrmann B, Mårtensson A, Alfvén T, Gantelius J, Andersson-Svahn H. A vertical flow paper-microarray assay with isothermal DNA amplification for detection of Neisseria meningitidis. Talanta. 2018;183:192–200.
(Answer 2)
For the automation of my project, I plan to use automation tools to develop and test a CRISPR-based biosensor that would be capable of detecting multi-drug-resistant tuberculosis (MDR-TB) signatures. This workflow would involve high-throughput liquid handling and cell-free protein synthesis. Possible steps would include:
(i) Module setup: This would include arranging reagents, tip racks, thermal cyclers, magnetic bead racks, and microarray holders on an Opentrons OT-2 deck [1]. This would be supplemented by temperature modules for incubation and heater-shaker modules for mixing and precise reaction control
(ii) Automated reaction setup: The robot will then perform pipetting of cell-free lysate, DNA templates, CRISPR guides, and cofactors into 96- or 384-well plate. Then multiple combinations of lineage-specific SNP guides and resistance mutation guides will be tested to evaluate ‘AND-gate logic’.
(iii) Incubation: External devices like a plate reader or miniPCR thermal cycler amplification will be loaded. Then Python scripts will be used to control timing, mixtures, and incubation periods.
(iv) Signal detection and analysis: Fluorescent outputs will be measure using devices such as Spark or PHERAstar FSX for high-throughput plate analysis [1]. This will be a measure of change in fluorescence colour which would indicate successful target detection and amplification.
(v) Microfluidic integration (if possible): If possible, will look to integrate 3D printed holders for small microfluidic chips. These can serve as small test cartridges for running multiple tests at once while minimizing manual handling and contamination risk in low-resource settings.
References
[1] Course Recitation Slides. Lab Automation Overview. Course presentation, [institution or course name if known]; Year [cited 2026 Feb 23]. Available from: https://docs.google.com/presentation/d/e/2PACX-1vQc3zo7Z0b6HK7YeC56p_n2RbHNjUHh1HI66DH0cHbFk0db1HlbF7gILE__NCvhUiYMjIGSOHwHPv2_/pub?start=false&loop=false&delayms=3000#slide=id.g2b9b763dcde_1_131