Week 6 HW: Genetic Circuits Part I: Assembly Technologies
Assignment: DNA Assembly
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The template DNA is the mUAV plasmid at Used at 20 ng/µL, with 0.8 µL added to the reaction. The primers are “colour forward” and “colour reverse”. Give that the stock concentration is 5 µL, using 2.5 µL of each primer in a total reaction volume of 25 µL results in a final primer concentration of 0.5 µM.
The Phusion HF Master mix is a solution containing DNA polymerase, nucleotides, and buffer Magnesium ions which enable accurate and efficient DNA amplification in PCR [1]. It was added at 12.5 µL from a 2X stock, resulting in a final concentration of 1X in the reaction.
Nuclease-free water is added (6.8 µL) to bring the total reaction volume up to 25 µL and ensure all components are at the correct final concentrations.
References
- New England Biolabs. Phusion High-Fidelity PCR Master Mix with HF Buffer [Internet]. Available from: https://www.neb.com/en-gb/products/m0531-phusion-high-fidelity-pcr-master-mix-with-hf-buffer
2. What are some factors that determine primer annealing temperature during PCR?
Annealing would depend on the melting point/temperature (Tm) of the primers; where annealing is generally done at 5 °C below the primer’s melting temperature [1]. Other factors include primer length, base composition (Guanine Content), salt and ion concentrations in the reaction (such as Mg2+ and monovalent salts).
References
- Integrated DNA Technologies. How do you calculate the annealing temperature for PCR? [Internet]. Available from: https://eu.idtdna.com/pages/support/faqs/how-do-you-calculate-the-annealing-temperature-for-pcr-?
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
| PCR | Restriction Enzyme Digests |
|---|---|
| Amplifies any region of DNA | Requires presence of specific restriction sites |
| Can introduce mutations, insertions, deletions, and overhangs for Gibson assembly | The enzymes target and cleave near these sites |
| Highly flexible and does not require specific sequences to introduce mutations [1] | Ends produced are either sticky or blunt |
| They allow for precise insertion of DNA fragments into vectors [2] |
For the is protocol, as it involves cloning, because it allows precise amplification of DNA fragments while introducing mutations and overlaps required for Gibson assembly. In contrast, restriction enzyme digestion would be limited to existing recognition and doesn’t easily introduce sequence changes.
References
National Human Genome Research Institute. Polymerase chain reaction (PCR) [Internet]. Bethesda (MD): NHGRI https://www.genome.gov/genetics-glossary/Polymerase-Chain-Reaction-PCR
Thermo Fisher Scientific. Restriction enzyme basics [Internet]. Waltham (MA):. Available from: https://www.thermofisher.com/uk/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/molecular-cloning/restriction-enzymes/restriction-enzyme-basics.html#:~:text=In%20cloning%2C%20restriction%20enzymes%20enable%20precise%20DNA,fundamental%20principle%20of%20recombinant%20DNA%20cloning%20technology.
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
Dpnl is a restriction enzyme that selectively digests methylated DNA, leaving unmethylated PCR products untouched [1]. Therefore, according to the lab protocol, adding 1 µL of dpnl to each sample to digest methylated DNA digests the mUAV template so that only newly created PCR fragments are introduced into the following Gibson Assembly step.
References
- University of Wisconsin–Madison. Lab 4: Background [Internet]. Biochemistry 551 Online Lab Manual; Available from: https://wisc.pb.unizin.org/biochemistry551online/chapter/lab-4-background/
5. How does the plasmid DNA enter the E. coli cells during transformation?
The most common forms of transformation are:
(i) Heat shock: Creating pores in cell wall through abrupt temperature changes
(ii) Electroporation: Generating pores via electrical voltage
These methods cause the wall to open up and create pores in the cell membrane, after which plasmids enter the bacteria through diffusion. After the initial heat/electric shock and entry, the pores eventually close up. Inside the bacteria, the plasmids replicate.
6. Describe another assembly method in detail (such as Golden Gate Assembly)
Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
Model this assembly method with Benchling or Asimov Kernel!
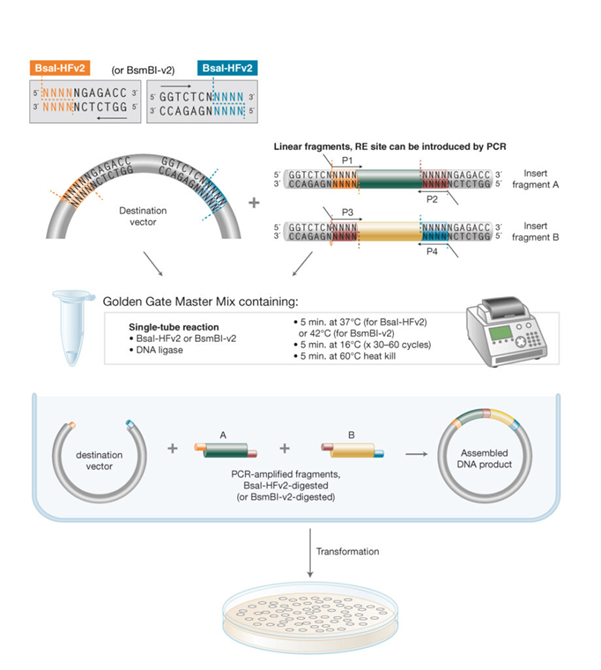
(1) The Golden Gate Assembly is a molecular cloning method that uses only the sequential or simultaneous activities of a single type IIS restriction enzyme and T4 DNA ligase [1], this enabled multiple inserts to be placed into the vector backbone in a single reaction.
Type IIS enzymes include BsaI, BsmBI, or BbsI [1, 2]. These cut DNA at a defined distance away from their recognition sites, rather than within them. This feature enables the generation of user-defined overhangs/fusion sites, which can be further designed to be unique and complementary guiding ordered and ligation of DNA parts with high specificity [2].
The reaction is done in 1 tube, where restriction digestion and ligation using T4 DNA ligase, increasing efficiency and reducing steps. Importantly, the recognition sites are removed during assembly, resulting in a seamless DNA construct [2]. The cyclic process of ligation (16 °C) and digestion (37 °C) facilitates repeated breakdown of incorrect assemblies and enhances growth of the selected product.
Tried to reference this whole process in this illustrated diagram:

To start, I tried entering J23100 Promoter sequence (35 nucleotides) into Benchling:
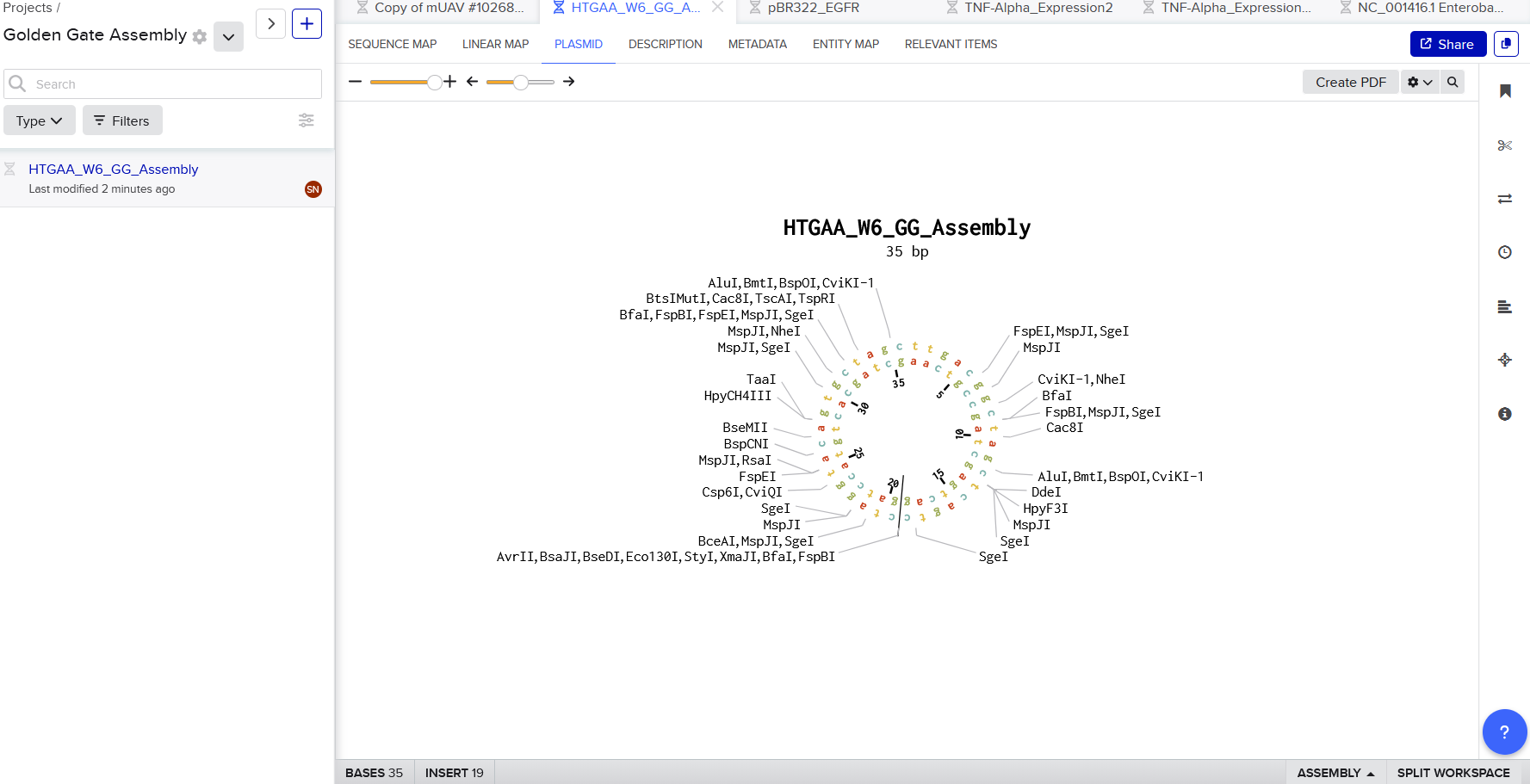

To facilitate Golden Gate Assembly, the promoter was then designed with flanking BsaI sites that allow the enzyme to create unique 4-base overhangs, ensuring the fragment inserts into the backbone in the correct orientation without leaving a ‘scar’ sequence. Therefore, I input (with some help from gemini to generate):
GGTCTCATCCCttgacggctagctcagtcctaggtacagtgctagcTACTTGAGACC
I was still having problems, so I added:
https://www.addgene.org/44335/ (as per suggestion from Gemini, given it is used in the CIDAR MoClo Parts Kit | https://www.addgene.org/kits/densmore-cidar-moclo/)
Now I’m getting these sticky end errors, too tired to solve it. But also, genuinely a bit lost and would like some more support on it.
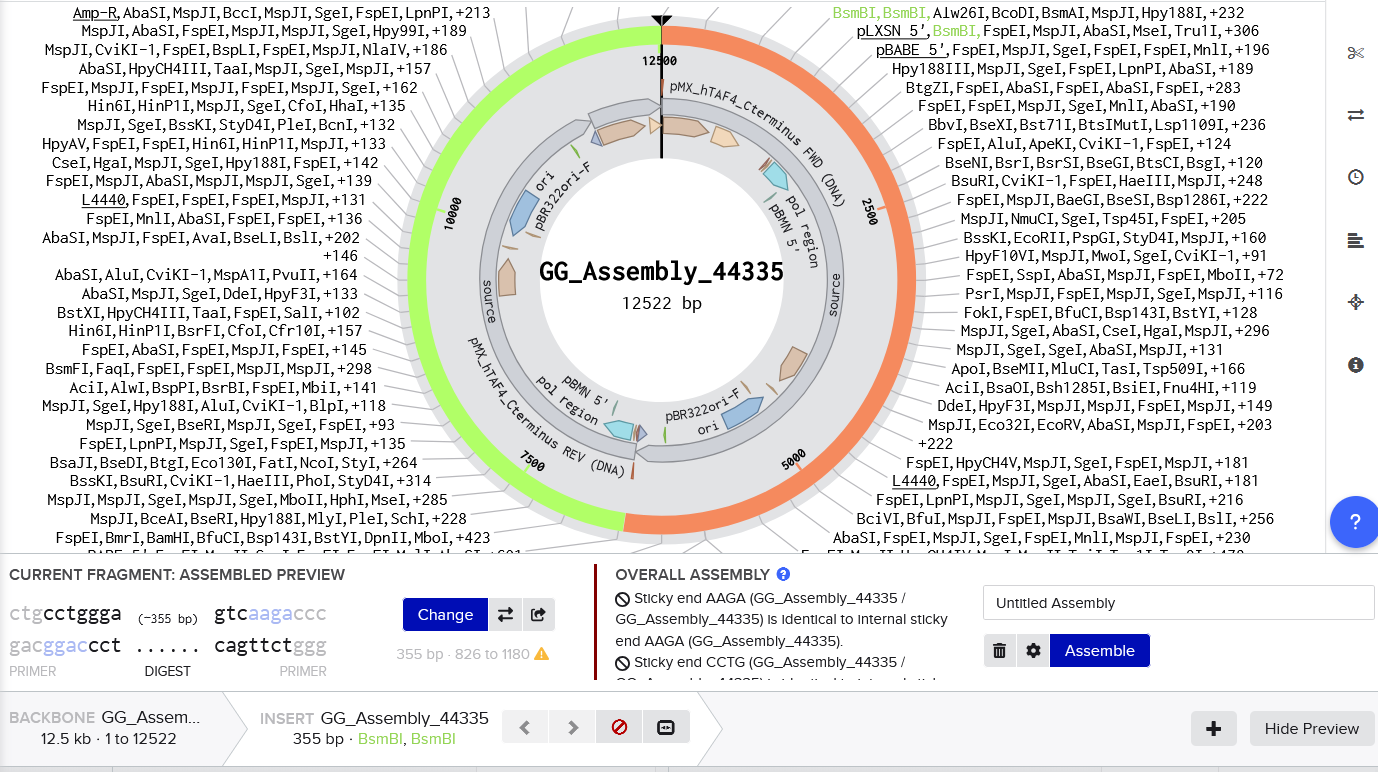

References
New England Biolabs. Golden Gate Assembly [Internet]. Ipswich (MA): New England Biolabs. Available from: https://www.neb-online.de/en/cloning-synthetic-biology/dna-assembly/golden-gate-assembly/
Laboratory Notes. Golden Gate Assembly [Internet]. Available from: https://www.laboratorynotes.com/golden-gate-assembly/