Week 2 HW: DNA Design Challenge
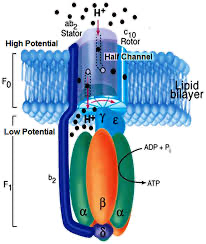
I chose the ATP synthase beta subunit because it’s essentially a biological motor and connects to my broader interest in energy systems:
Protons flow down their gradient across the mitochondrial membrane, almost like current moving through a circuit, and that flow physically spins part of the protein like a tiny turbine. That rotation drives changes in the beta subunits, which catalyze the formation of ATP from ADP and phosphate.
So it’s literally energy stored in a gradient being converted into mechanical motion and then into chemical energy. I find that idea really compelling, it’s molecular thermodynamics in action, where fundamental physics laws become something tangible inside living cells.
From NCBI I obtained the protein sequence:
https://www.ncbi.nlm.nih.gov/protein/NP_001677.2/
https://www.ncbi.nlm.nih.gov/protein/NP_001677.2?report=fasta
MLGFVGRVAAAPASGALRRLTPSASLPPAQLLLRAAPTAVHPVRDYAAQTSPSPKAGAATGRIVAVIGAV VDVQFDEGLPPILNALEVQGRETRLVLEVAQHLGESTVRTIAMDGTEGLVRGQKVLDSGAPIKIPVGPET LGRIMNVIGEPIDERGPIKTKQFAPIHAEAPEFMEMSVEQEILVTGIKVVDLLAPYAKGGKIGLFGGAGV GKTVLIMELINNVAKAHGGYSVFAGVGERTREGNDLYHEMIESGVINLKDATSKVALVYGQMNEPPGARA RVALTGLTVAEYFRDQEGQDVLLFIDNIFRFTQAGSEVSALLGRIPSAVGYQPTLATDMGTMQERITTTK KGSITSVQAIYVPADDLTDPAPATTFAHLDATTVLSRAIAELGIYPAVDPLDSTSRIMDPNIVGSEHYDV ARGVQKILQDYKSLQDIIAILGMDELSEEDKLTVSRARKIQRFLSQPFQVAEVFTGHMGKLVPLKETIKG FQQILAGEYDHLPEQAFYMVGPIEEAVAKADKLAEEHSS
We know we go from 3 DNA bases → RNA → 1 Codon → 1 Amino Acid → 1 Protein letter
We can find the nucleotide record
https://www.ncbi.nlm.nih.gov/nuccore/NM_001686.4
https://www.ncbi.nlm.nih.gov/nuccore/NM_001686.4?report=fasta
AGTCTCCACCCGGACTACGCCATGTTGGGGTTTGTGGGTCGGGTGGCCGCTGCTCCGGCCTCCGGGGCCT TGCGGAGACTCACCCCTTCAGCGTCGCTGCCCCCAGCTCAGCTCTTACTGCGGGCCGCTCCGACGGCGGT CCATCCTGTCAGGGACTATGCGGCGCAAACATCTCCTTCGCCAAAAGCAGGCGCCGCCACCGGGCGCATC GTGGCGGTCATTGGCGCAGTGGTGGACGTCCAGTTTGATGAGGGACTACCACCAATTCTAAATGCCCTGG AAGTGCAAGGCAGGGAGACCAGACTGGTTTTGGAGGTGGCCCAGCATTTGGGTGAGAGCACAGTAAGGAC TATTGCTATGGATGGTACAGAAGGCTTGGTTAGAGGCCAGAAAGTACTGGATTCTGGTGCACCAATCAAA ATTCCTGTTGGTCCTGAGACTTTGGGCAGAATCATGAATGTCATTGGAGAACCTATTGATGAAAGAGGTC CCATCAAAACCAAACAATTTGCTCCCATTCATGCTGAGGCTCCAGAGTTCATGGAAATGAGTGTTGAGCA GGAAATTCTGGTGACTGGTATCAAGGTTGTCGATCTGCTAGCTCCCTATGCCAAGGGTGGCAAAATTGGG CTTTTTGGTGGTGCTGGAGTTGGCAAGACTGTACTGATCATGGAGTTAATCAACAATGTCGCCAAAGCCC ATGGTGGTTACTCTGTGTTTGCTGGTGTTGGTGAGAGGACCCGTGAAGGCAATGATTTATACCATGAAAT GATTGAATCTGGTGTTATCAACTTAAAAGATGCCACCTCTAAGGTAGCGCTGGTATATGGTCAAATGAAT GAACCACCTGGTGCTCGTGCCCGGGTAGCTCTGACTGGGCTGACTGTGGCTGAATACTTCAGAGACCAAG AAGGTCAAGATGTACTGCTATTTATTGATAACATCTTTCGCTTCACCCAGGCTGGTTCAGAGGTGTCTGC ATTATTGGGCCGAATCCCTTCTGCTGTGGGCTATCAGCCTACCCTGGCCACTGACATGGGTACTATGCAG GAAAGAATTACCACTACCAAGAAGGGATCTATCACCTCTGTACAGGCTATCTATGTGCCTGCTGATGACT TGACTGACCCTGCCCCTGCTACTACGTTTGCCCATTTGGATGCTACCACTGTACTGTCGCGTGCCATTGC TGAGCTGGGCATCTATCCAGCTGTGGATCCTCTAGACTCCACCTCTCGTATCATGGATCCCAACATTGTT GGCAGTGAGCATTACGATGTTGCCCGTGGGGTGCAAAAGATCCTGCAGGACTACAAATCCCTCCAGGATA TCATTGCCATCCTGGGTATGGATGAACTTTCTGAGGAAGACAAGTTGACCGTGTCCCGTGCACGGAAAAT ACAGCGTTTCTTGTCTCAGCCATTCCAGGTTGCTGAGGTCTTCACAGGTCATATGGGGAAGCTGGTACCC CTGAAGGAGACCATCAAAGGATTCCAGCAGATTTTGGCAGGTGAATATGACCATCTCCCAGAACAGGCCT TCTATATGGTGGGACCCATTGAAGAAGCTGTGGCAAAAGCTGATAAGCTGGCTGAAGAGCATTCATCGTG AGGGGTCTTTGTCCTCTGTACTGTCTCTCTCCTTGCCCCTAACCCAAAAAGCTTCATTTTTCTGTGTAGG CTGCACAAGAGCCTTGATTGAAGATATATTCTTTCTGAACAGTATTTAAGGTTTCCAATAAAATGTACAC CCCTCAGAA
Multiple codons can code for the same amino acid, but different organisms prefer certain codons over others. So we have to optimize codon usage for that specific organism otherwise translation might be inefficient, we want to use tRNA’s that are plentiful – which bind to that specific codon attaching the specific amino acid.
I have chosen E.coli as the organism to optimize the protein sequence for. Since we use them in the fluorescent bacteria artwork lab!
Above is the entire mRNA sequence, but we need the coding sequence (CDS) – the mRNA sequence has additional information like a start and end codon and untranslated regions. We can go to the CDS record instead, obtain the coding sequence and then use our codon optimization on it.
https://www.ncbi.nlm.nih.gov/CCDS/CcdsBrowse.cgi?REQUEST=CCDS&DATA=CCDS8924.1
https://www.idtdna.com/CodonOpt
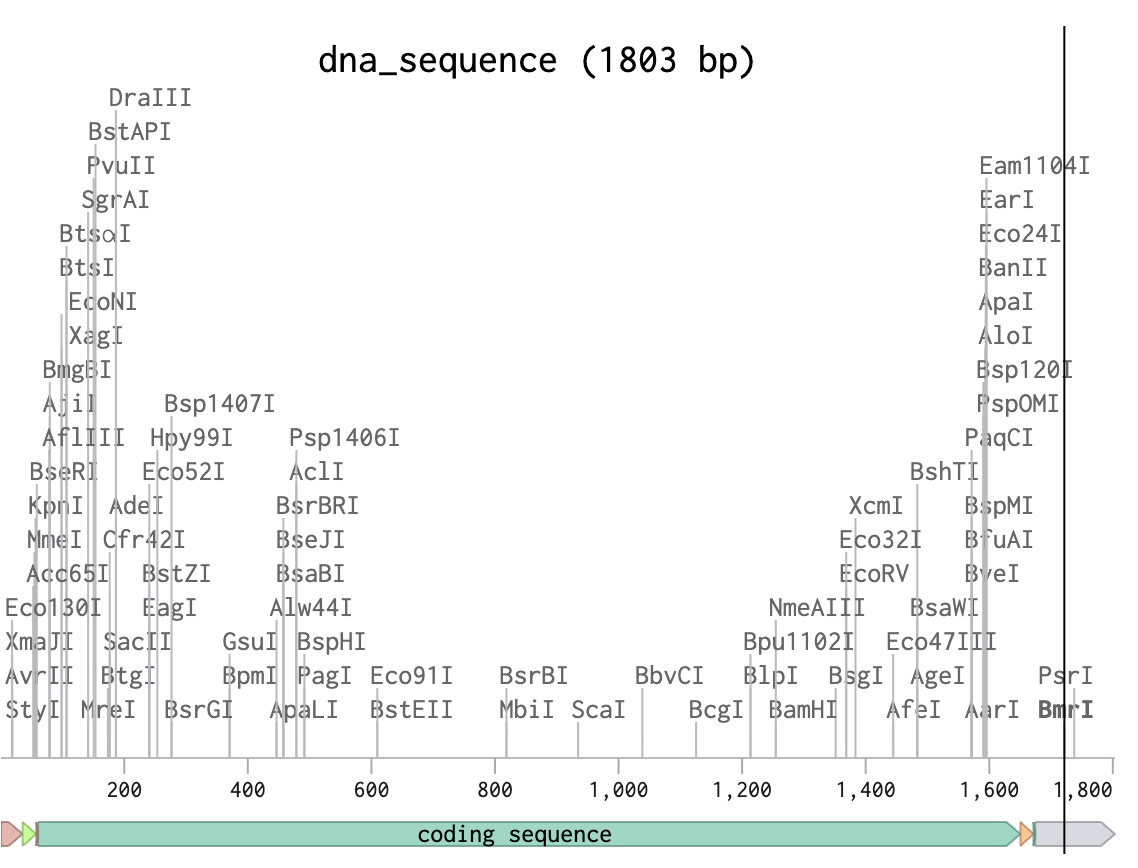
We get this result:
We can use cell dependent expressions, like cloning the optimized DNA sequence into a plasmid vector and introducing it into a host organism such as E.coli. Once inside the promoter recruits RNA polymerase and transcribes the DNA sequence into mRNA. The ribosomes then binds to the mRNA and tRNA’s match codons and deliver amino acids. The amino acids are then linked together to form the protein. The bacteria would then produce ATP synthase beta subunit as part of their cellular machinery.
Expression Cassette
https://benchling.com/s/seq-QDGibA4g7TjoTuX3lb5A?m=slm-Gx8zqXYh9sr4lxSK0Xqu
We can view the full plasmid sequence for our clonal genes (circular dna) and pTwist Amp High Copy cloning vector in Benchling:
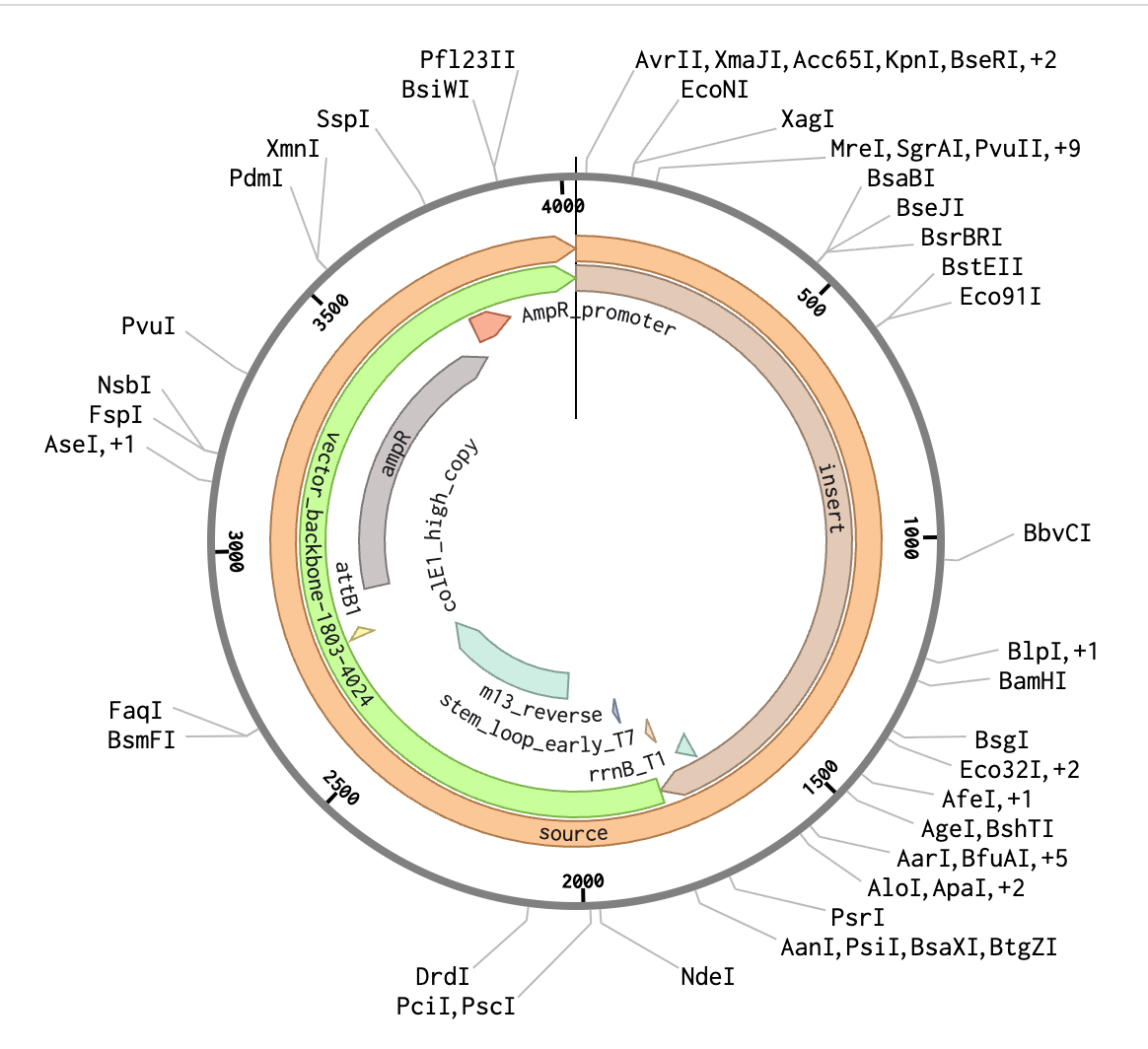
https://benchling.com/s/seq-wsl9w63Z5DcxN7rlp5cG?m=slm-ndl9y5U2FSsJgNYW6z7
I would choose to sequence the DNA of extremophiles that thrive in high-radiation or high-temperature environments. By sequencing genes involved in radiation resistance, DNA repair, and protein stabilization, we could better understand the molecular mechanisms that allow biological systems to survive under extreme stress. This knowledge could help inform the engineering of radiation-resistant biological materials or bio-hybrid systems designed to operate in harsh energy environments. Studying these organisms connects molecular biology with broader challenges in advanced energy systems.

I would use Illumina sequencing to sequence the DNA since it provides high accuracy and high throughput and is well suited for whole-genome sequencing and variant detection. It’s a second generation technique, sequencing millions of short DNA fragments in parallel using sequencing-by-synthesis. Illumina sequencing reads DNA by copying it one base at a time and taking a picture after each base is added.
The input would be the extracted genomic DNA.
Preparation steps:
- Fragment DNA into short pieces
- Ligate sequencing adapters
- PCR amplify fragments
- Load onto flow cell for cluster amplification
Essential sequencing steps:
- DNA fragments bind to flow cell
- Bridge amplification forms clusters
- Fluorescently labeled nucleotides are added one at a time
- A camera detects the fluorescent signal for each incorporated base
- The color signal determines the base
The output would be millions of short sequence reads containing nucleotide sequences and quality scores which can be assembled into a genome or aligned to a reference.
I would want to synthesize a cluster of genes involved in enhanced DNA repair and protein stabilization from extremophiles and express them in a model organism. By combining multiple protective pathways, we could engineer cells with improved resistance to radiation and thermal stress. The idea would to use this to develop radiation-resistant biomaterials or biological components for extreme energy environments. We could build a genetic circuit that enables engineered bacteria to sense and respond to radiation stress. This circuit could include radiation response promoters, DNA repair genes and protective protein pathways that activate under high oxidative or ionizing radiation conditions.
To synthesize this genetic circuit, we could use Twist combined with phosphoramidite solid-phase DNA synthesis and Gibson Assembly for multi-fragment assembly.
Essential steps:
- Design optimized DNA sequence computationally
- Chemically synthesize short oligonucleotides (base-by-base addition)
- Cleave and purify oligos
- Assemble fragments into full-length gene (e.g., Gibson Assembly)
- Clone into plasmid backbone
- Sequence-verify construct
Limitations:
Length limits: Direct chemical synthesis is reliable only for short fragments with longer genes requiring assembly. There’s also base errors so we would need to do sequencing validation and it can be very expensive for large gene clusters and take a large amount of time.
I would edit the genomes of photosynthetic microorganisms such as algae to improve their efficiency in converting light energy into chemical fuels. I could target genes involved in photosystem efficiency, carbon fixation pathways, and hydrogen production.
Photosynthesis is essentially a natural solar energy conversion system, but it is quite inefficient. We could modify regulatory genes to reduce energy losses or redirect metabolic pathways toward hydrogen or biofuel production, so we could have biological systems that convert sunlight into storable chemical energy more efficiently.
I am interested as it connects directly to large-scale energy systems and treating living cells as programmable energy conversion platforms, similar to designing more efficient reactors or turbines.

WE could use CRISPR-Cas12a for genome editing in cyanobacteria.
How does it edit dna
- Design guide RNAs targeting specific genes.
- Deliver Cas12a and guide RNAs into the cells.
- Cas12a cuts the DNA at precise locations.
- The cell repairs the cut using a donor DNA template to insert optimized sequence
Design:
- Identify metabolic bottlenecks in photosynthesis or fuel production.
- Design guide RNAs.
- Design donor DNA templates if inserting new sequences.
Inputs:
- Cas enzyme
- Guide RNAs
- Donor DNA (if needed)
- Host cells (e.g., cyanobacteria)
Limitations
- Off-target edits may occur.
- Large pathway rewiring is complex.
- Efficiency gains may be modest due to thermodynamic constraints.