Week 2 Lab: DNA Read Write Edit

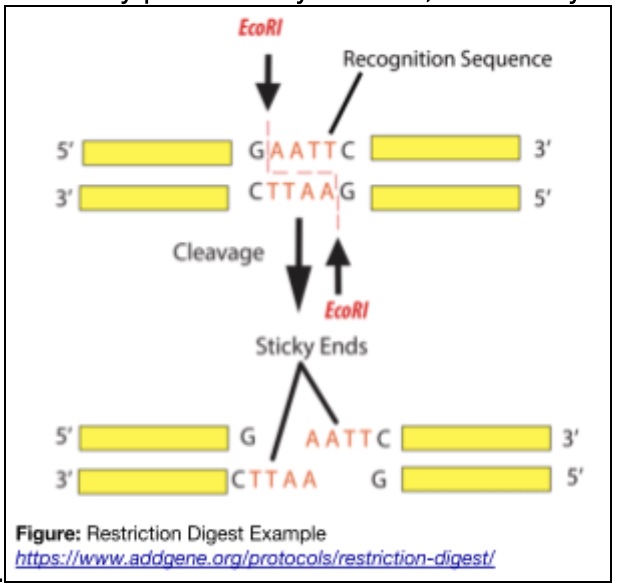
In Lab 2 I learned how restriction enzymes can be used to cut DNA at very specific sequences, almost like precise molecular scissors. These enzymes recognize short DNA sequences called restriction sites and cleave the DNA at or near those locations, allowing us to deliberately fragment genetic material in a controlled way.
I also learned that restriction enzymes, or endonucleases, naturally come from bacteria. In their original context, they act as a defense mechanism by cutting up invading viral DNA, protecting the bacterial cell from infection. It was interesting to see how a biological immune strategy becomes a foundational lab tool.
We then discussed how CRISPR can be thought of as a generalized or programmable restriction enzyme. Instead of being limited to one fixed recognition site, CRISPR systems can be guided to almost any DNA sequence, making them far more flexible and powerful for gene editing.
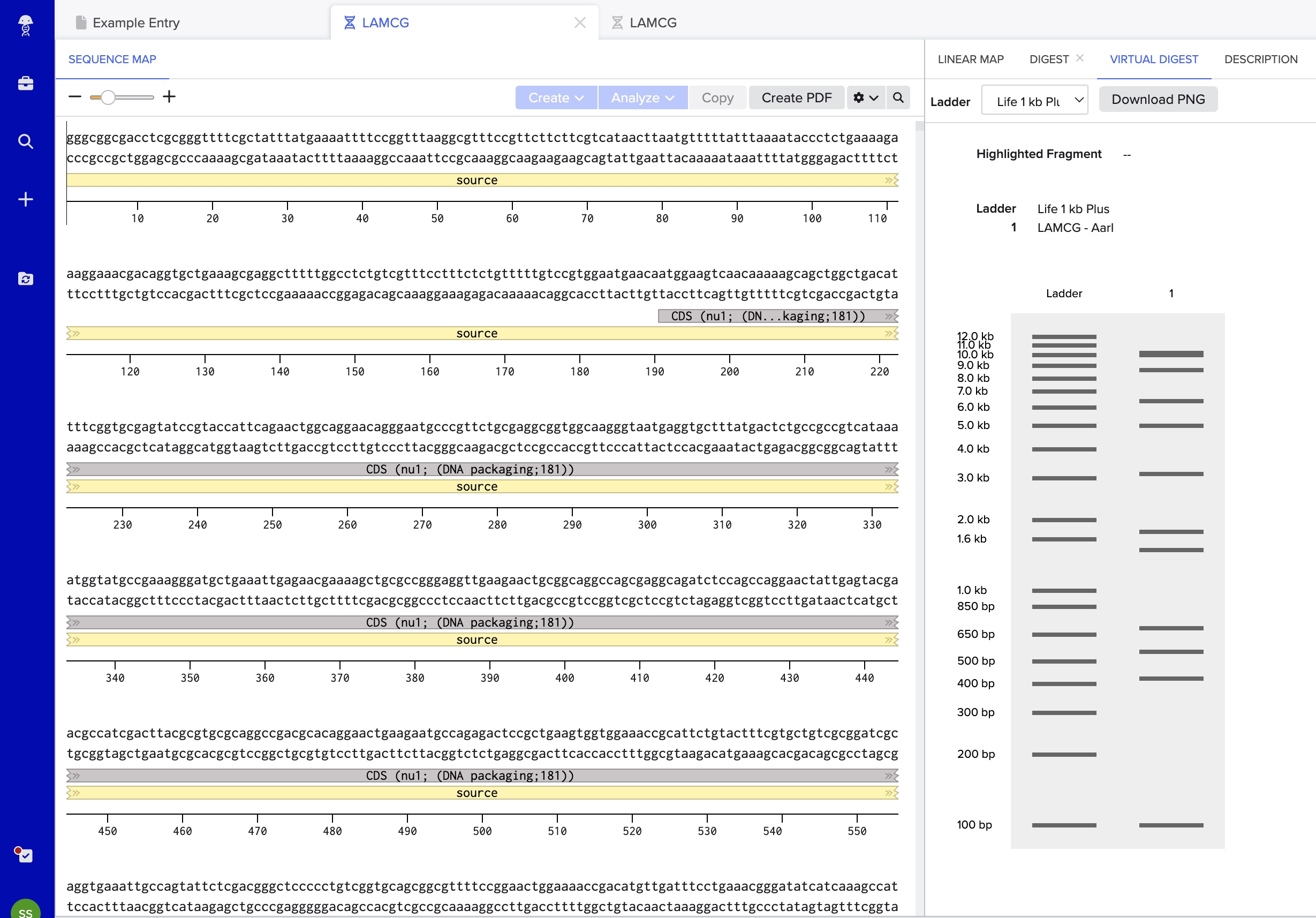
I also learned how to use Benchling to simulate restriction enzyme digests in silico. We uploaded DNA sequences and tested different enzyme combinations to see how the DNA would be cut and what fragment sizes we would expect before actually running the gel.
To run a virtual digest, the DNA sequence has to be uploaded in a standard format, usually either FASTA or GenBank.
I learned in more detail how gel electrophoresis works and why DNA moves through the gel the way it does. Because DNA has a negatively charged phosphate backbone, it migrates toward the positive electrode when an electric field is applied. The agarose gel acts like a molecular sieve, so smaller DNA fragments move faster and travel further than larger ones, allowing the fragments to separate by size.
I weighed out agarose and mixed it with 1x TAE buffer to make a 1 percent solution. I microwaved it in short bursts until it fully dissolved, let it cool slightly, added SYBR Safe stain, poured it into the gel tray with a comb inserted, and allowed it to solidify to form wells.
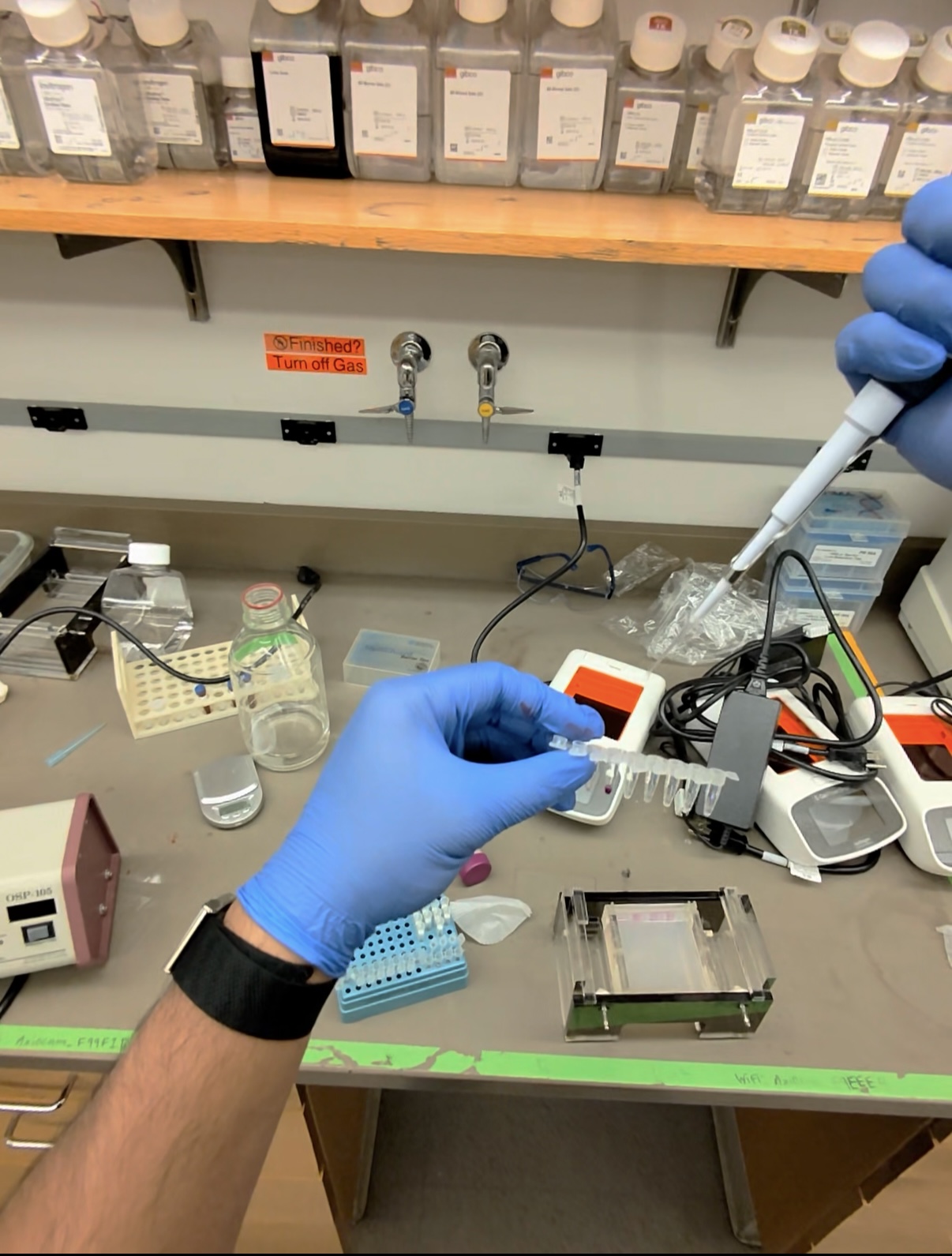
I prepared the DNA digestion mixture by combining lambda DNA, the correct enzyme buffer, the chosen restriction enzyme or enzymes, and nuclease free water. I then incubated the tubes at 37 degrees Celsius so the enzymes could cut the DNA into fragments.

After the gel set, I removed the comb, filled the gel box with 1x TAE buffer, and mixed my DNA samples with loading dye. I carefully loaded each well without puncturing the gel and ran the gel at around 80 to 115 volts for about 45 minutes to separate the DNA fragments by size.

Once the run was complete, I transferred the gel to a blue light transilluminator, and captured an image of the separated DNA bands to analyze the pattern of fragments - there was a lot of noise but the experiment was fun nonetheless.