HTGAA 2026: Lab Automation & DNA Design
1. Laboratory Automation: Opentrons Bio-Art
Using the HTGAA26 Opentrons Colab as a framework, I developed a custom automation protocol to translate digital designs into biological patterns.
Implementation Documentation
- Technical Script: sami_tanveer_opentrons.py
- Protocol Logic: The script utilizes API Level 2.20 and a P20 Single-Channel Gen2 pipette. It features an optimized
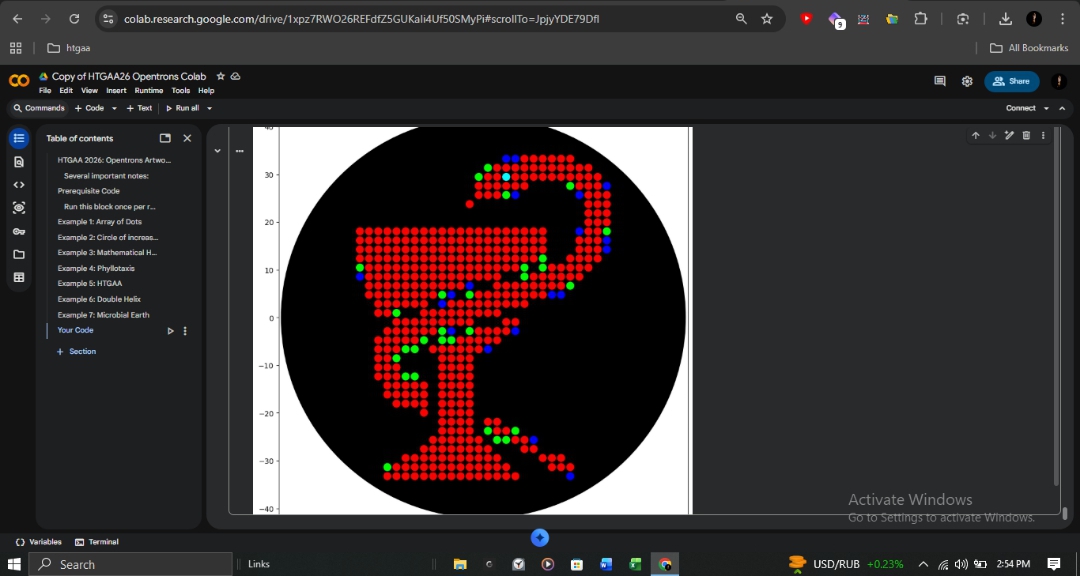
draw_pointsfunction that handles coordinate-based dispensing with batched aspiration to ensure mechanical efficiency and prevent cross-contamination between fluorescent strains. - Design Interface: The design was mapped using the Opentrons Art GUI, ensuring precise coordinate placement for Red (mRFP1), Green (mClover3), Blue (Azurite), and Cyan (sfGFP) reporters.
Visual Reference of Design Interface:

2. Post-Lab Questions & Research
Q1: Published Paper — SHERLOCK (Nature Protocols, 2019)
Paper: SHERLOCK: Specific High-sensitivity Enzymatic Reporter unLOCKing (PubMed: 31548639).
SHERLOCK is a CRISPR-based diagnostic tool that utilizes the collateral cleavage activity of Cas13. This paper is foundational to my final project because it demonstrates a detection method that is both highly sensitive and low-cost (~$0.60/reaction), making it ideal for the Pakistani clinical context.
Automation Connection: The paper highlights the transition from manual detection to high-throughput plates. Utilizing an Opentrons robot allows for the parallelization of hundreds of reactions, ensuring that carrier screening is consistent, rapid, and free from the human pipetting errors common in manual diagnostics.
Q2: Final Project — Automated Thalassemia Biosensor
Motivation
Thalassemia is a critical public health burden in Pakistan (5–8% carrier rate). Current testing (HPLC/Sequencing) is centralized and expensive ($50–$200). My project aims to develop a $3, 90-minute portable biosensor validated via automated workflows.
Technical Workflow
- DNA Extraction: Rapid lysis from patient blood.
- LAMP Amplification: Isothermal amplification at 65°C, removing the need for thermal cyclers.
- Cas13 Detection: Programmed with gRNA targeting common HBB mutations.
- Visual Readout: Result visualized via UV torch (glow = positive).
3. Project Presentation Assets
This section visualizes the engineering roadmap and clinical justification for the Automated Thalassemia Biosensor. These assets bridge the gap between initial molecular design and high-throughput diagnostic implementation.
Slide 1: The Clinical Burden (Pakistan Context)
Strategic Context: Identifying the socio-economic barriers to thalassemia screening in Pakistan. This slide establishes the “Why” behind the project—replacing expensive, centralized testing with a $3 portable alternative tailored for rural health clinics.

Slide 2: Molecular Diagnostic Logic (SHERLOCK)
Technical Mechanism: Leveraging the Cas13 enzyme for programmable DNA detection. This slide details the transition from a patient sample to a visible fluorescent readout using the SHERLOCK protocol.


Slide 3: High-Throughput Automation Roadmap
Systems Engineering: A comprehensive plan for scaling validation. By utilizing the Ginkgo Nebula cloud laboratory (including the Echo acoustic handler and PHERAstar plate reader), we move from manual testing to a parallelized 384-well screening environment.


4. Technical Resources & Methodology
- Automation Protocol: sami_tanveer_opentrons.py
- Experimental Design Hub: [Copy of HTGAA26 Opentrons Colab](./Copy of HTGAA26 Opentrons Colab)
AI Methodology
I utilized Claude (Anthropic) and Google Gemini for script optimization—specifically for refactoring coordinate lists into the draw_points batching logic and structuring the Ginkgo Nebula high-throughput workflow. All clinical objectives and molecular strategies are my original work.