Week 7 Homework — Genetic Circuits Part II: Neuromorphic Circuits
Part 1: Intracellular Artificial Neural Networks (IANNs)
Q1. What advantages do IANNs have over traditional genetic circuits whose input/output behaviors are Boolean functions?
Traditional genetic circuits implement Boolean logic — outputs are strictly binary (gene ON or gene OFF). IANNs offer several key advantages over this approach.
First, IANNs produce analog, continuous outputs proportional to input signal strength, allowing nuanced regulation based on the level of a signal rather than just its presence or absence. Second, each input can be assigned a weight via promoter strength, RBS efficiency, or RNA stability, so the cell can differentially prioritize signals — something impossible with standard Boolean gates. Third, IANNs can classify complex, high-dimensional input combinations that would require exponentially many Boolean gates to approximate. Fourth, weights can be tuned synthetically or evolutionarily to change circuit behavior without rewiring the topology. Finally, analog computation with soft thresholds is more robust to stochastic gene expression noise than hard Boolean switching, which is especially important given the inherently noisy environment of a living cell.
Q2. Describe a useful application for an IANN, including detailed input/output behavior and limitations.
Application: Intracellular cancer cell classifier
Goal: Program a therapeutic cell to detect whether it resides inside a cancer cell and trigger selective apoptosis only when the cancer-specific molecular signature is detected.
Inputs:
- X₁ = intracellular miR-21 concentration (an oncomiR elevated in many cancers)
- X₂ = p53 mRNA level (a tumor suppressor depleted in many cancers)
Input/output behavior: Layer 1 weights X₁ positively and X₂ negatively. The perceptron fires — producing an intermediate transcription factor — only when miR-21 is high AND p53 is low, which constitutes the cancer-specific signature. Layer 2 translates that transcription factor output into a pro-apoptotic protein such as a caspase activator. The output is continuous and graded: low basal expression occurs in normal cells, while high expression (leading to cell death) occurs in cancer cells. The analog nature of the IANN allows partial activation at borderline signal levels rather than crashing to a binary output, giving the circuit proportional rather than all-or-nothing behavior.
Limitations:
Off-target activation may occur in normal cells with transiently elevated miR-21, for example during inflammation. There is also a delivery challenge: the IANN construct must reach every target cell. Weight tuning in vivo is not yet routine — circuit parameters must be designed computationally and validated empirically before deployment. Finally, the metabolic burden of maintaining the circuit may impair host cell fitness over time.
Q3. Multilayer perceptron diagram
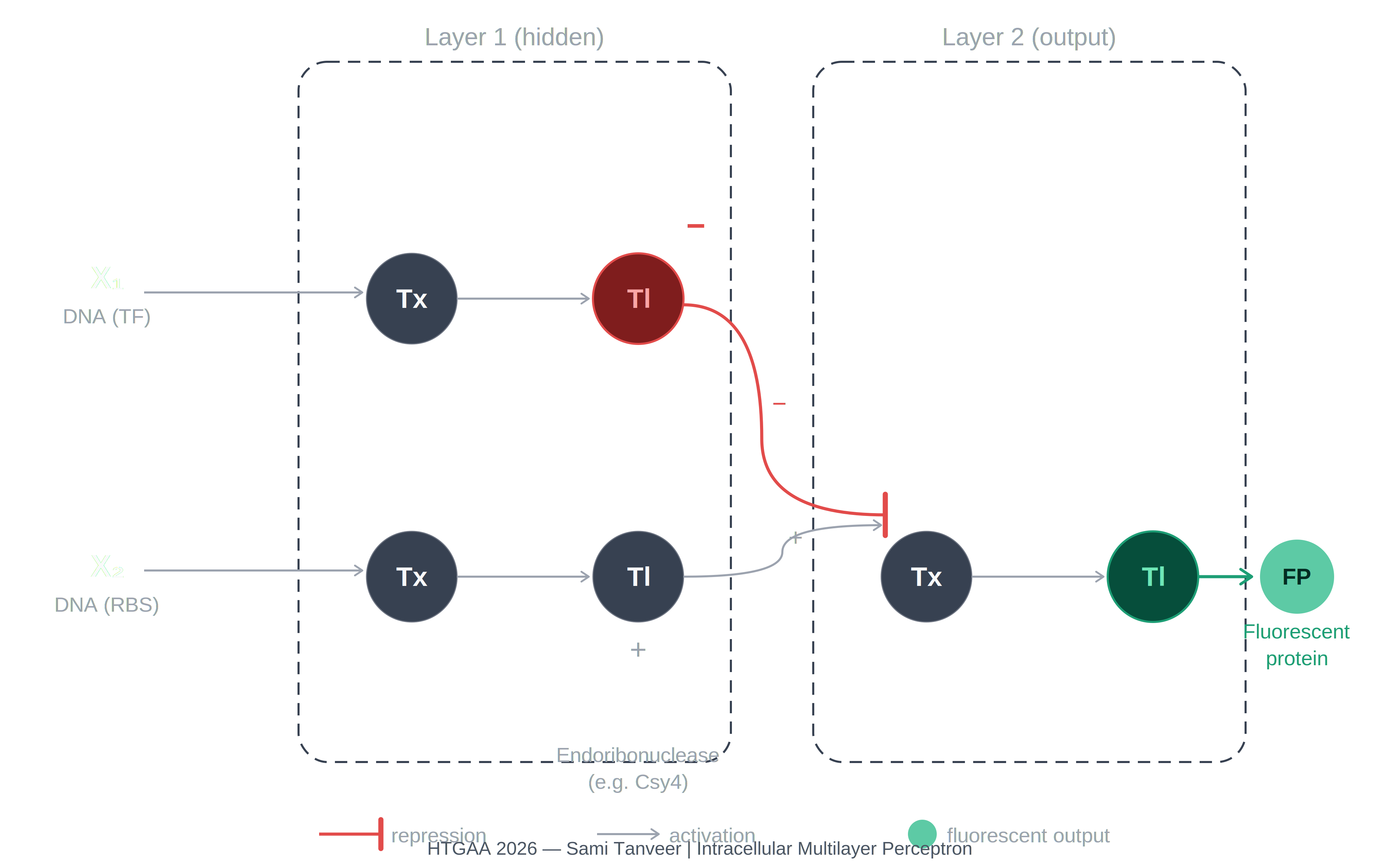

The diagram depicts an intracellular two-layer perceptron extending the single-layer architecture shown in the assignment reference image (1774856540029_image.png).
Layer 1 — hidden layer:
X₁ (DNA encoding a transcription factor) undergoes Tx → Tl and produces Csy4 endoribonuclease, represented as the red inhibitory node. X₂ (DNA encoding a regulatory element) undergoes Tx → Tl and produces an activating signal represented as the gray positive node. Csy4 carries a repression signal (−) forward into Layer 2.
Layer 2 — output layer:
The layer receives the repression signal from Csy4 (−) and the activation signal from the X₂ branch (+). These are integrated at the Layer 2 Tx node. The downstream Tl step produces the fluorescent protein output, represented as the teal circle (FP). The fluorescent protein is expressed only when the repression from Csy4 is absent or outweighed by the activating input from X₂.
The key difference from the single-layer perceptron is the insertion of the endoribonuclease as a tunable hidden-layer node that integrates Layer 1 signals before passing a regulatory output to Layer 2, enabling more complex input-output computation than a single weighted integration step.
Part 2: Fungal Materials
Q1. Examples of existing fungal materials, their uses, advantages, and disadvantages over traditional counterparts
Mycelium — the vegetative, thread-like root network of fungi — is the basis for most current fungal biomaterials.
Mycelium composites (e.g., Ecovative Mylo, Bolt Threads): Used as packaging foam replacements and leather-like textiles. The mycelium is grown on agricultural waste substrates such as corn stalks or hemp hurds, then heat-killed to stop growth, producing a rigid or flexible material depending on processing. These are fully biodegradable, carbon-negative to produce, require low energy manufacturing, contain no petrochemicals, and can be grown into near-net shapes to reduce waste. Their disadvantages include lower mechanical strength and water resistance compared to synthetic foams or conventional leather, inconsistent properties between batches, and currently higher cost than conventional materials at scale.
Fungal-derived chitosan: Extracted from fungal cell walls of species such as Aspergillus and Rhizopus, used in wound dressings, food preservatives, and water treatment. It provides a renewable source that avoids the shellfish allergen concerns of crustacean-derived chitosan and has useful antimicrobial properties. However, fungal chitosan has lower molecular weight than crustacean-derived chitosan, which reduces viscosity and film-forming performance in some applications.
Mycelium biocomposites for construction: Experimental building insulation panels and structural bricks grown from mycelium combined with hemp or straw substrates. These are compostable at end of life, fire-resistant, and offer good acoustic and thermal insulation properties that can be grown locally from agricultural waste. They are not yet load-bearing at scale, are sensitive to moisture, and require sterile growth conditions during production.
Q2. What would you engineer fungi to do, and what are the advantages of synthetic biology in fungi over bacteria?
Engineering targets:
I would engineer filamentous fungi — specifically Aspergillus niger or Trichoderma reesei — to produce self-healing mycelium composites. The design would use stretch-sensitive promoters to detect mechanical damage and upregulate chitin synthase and cell wall repair genes, allowing the material to patch cracks autonomously. A second goal would be engineering mycelium to incorporate conductive nanoparticles during growth, producing biodegradable flexible electronics substrates that could replace conventional PCB materials.
Advantages of synthetic biology in fungi over bacteria:
Fungi perform eukaryotic post-translational processing including glycosylation, disulfide bonding, and complex protein folding that bacteria cannot, making them superior for producing functional complex proteins and enzymes. Filamentous fungi like Trichoderma are natural high-yield protein secretors — industrial strains already produce grams per liter of cellulase, a capacity bacteria cannot match without extensive engineering. Fungi also have large genomes with rich secondary metabolite pathways including polyketides, terpenes, and alkaloids, offering a broader toolkit for novel chemistry. Critically, mycelium naturally forms macroscale three-dimensional architectures, meaning engineering the material properties of a mycelium composite operates at the organism level rather than just the molecular level — something bacteria cannot replicate without significant external scaffolding. Finally, many fungi including Aspergillus and Saccharomyces hold Generally Recognized As Safe (GRAS) status, simplifying regulatory pathways for food and medical applications.
Part 3: First DNA Twist Order
Q1. Documentation guidelines review
Individual Final Project documentation guidelines reviewed. ✓
Q2. Google Form submission
Google Form submitted with the following:
- Project title: Cell-Free Toehold Switch Diagnostic for β-Thalassemia Fr 8-9 (+G) Mutation Detection
- Draft Aim 1: Design and validate a toehold switch riboregulator detecting the Fr 8-9 (+G) HBB frameshift mutation using a CFPS platform, confirmed by sfGFP fluorescence with synthetic trigger RNA as positive control and wild-type HBB mRNA as negative control
- Final project summary: submitted (see form document)
- HTGAA industry council selections: Twist Biosciences, Ginkgo Bioworks, New England Biolabs, Thermo Fisher Scientific, SecureDNA
- Benchling folder: HTGAA 2026 - Beta Thalassemia Diagnostic (shared folder, link sharing on)
Submitted prior to the March 20 deadline for MIT/Harvard/Wellesley students. ✓
Q3. DNA Design — Insert sequence and backbone vector
Project context: My individual final project is a cell-free toehold switch biosensor for detecting the HBB Fr 8-9 (+G) frameshift mutation associated with β-thalassemia, designed for point-of-care use in low-resource settings in northern Pakistan.
Sequences designed (Benchling — HTGAA 2026 Beta Thalassemia Diagnostic project):
Three sequences were designed and deposited:
- HBB_Fr8-9_plusG_trigger_30nt — the 30-nucleotide synthetic trigger RNA corresponding to the mutant HBB Fr 8-9 (+G) mRNA sequence, used as positive control
- HBB_toehold_switch_sfGFP_construct — the full toehold switch construct (v1) comprising the trigger-binding toehold region, hairpin loop with embedded RBS, and sfGFP reporter
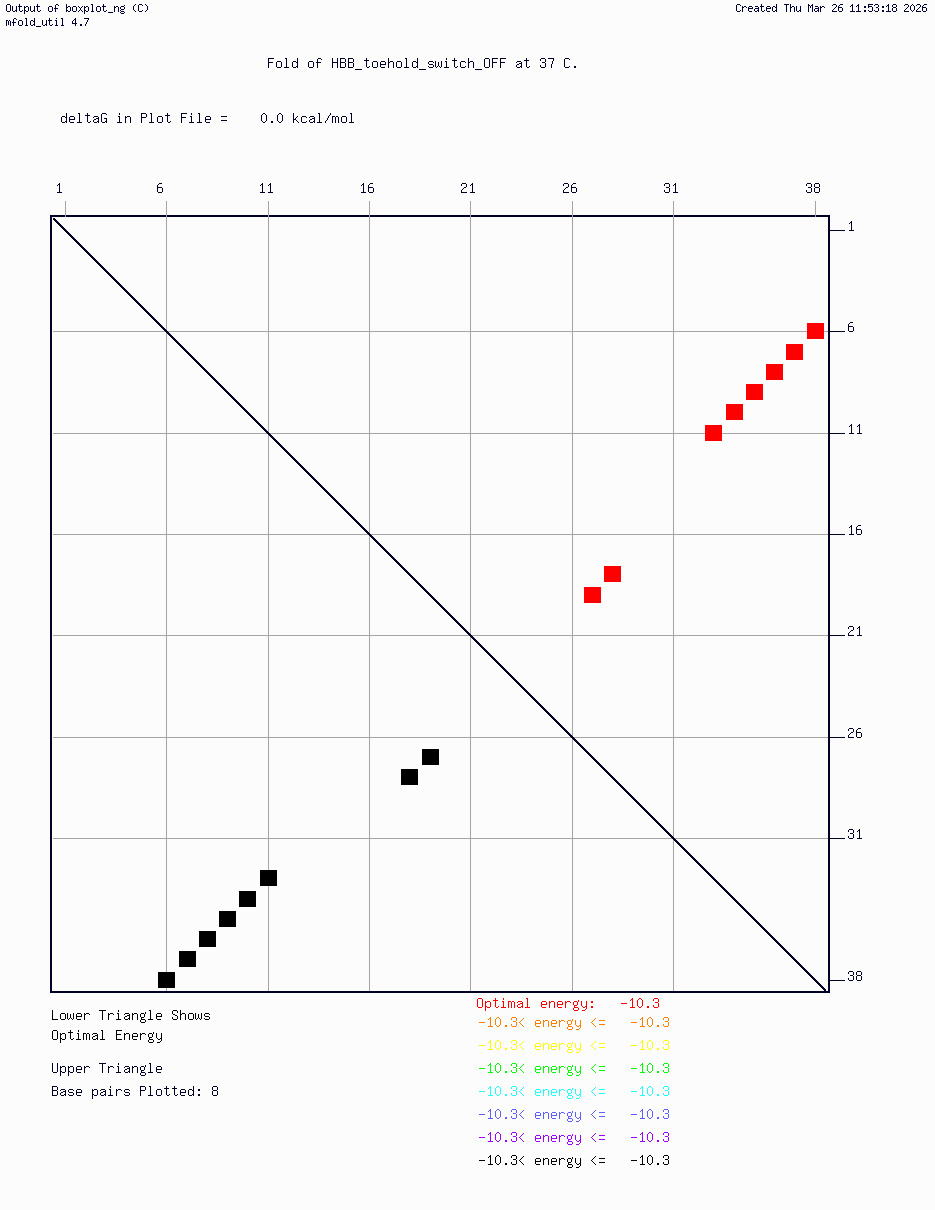
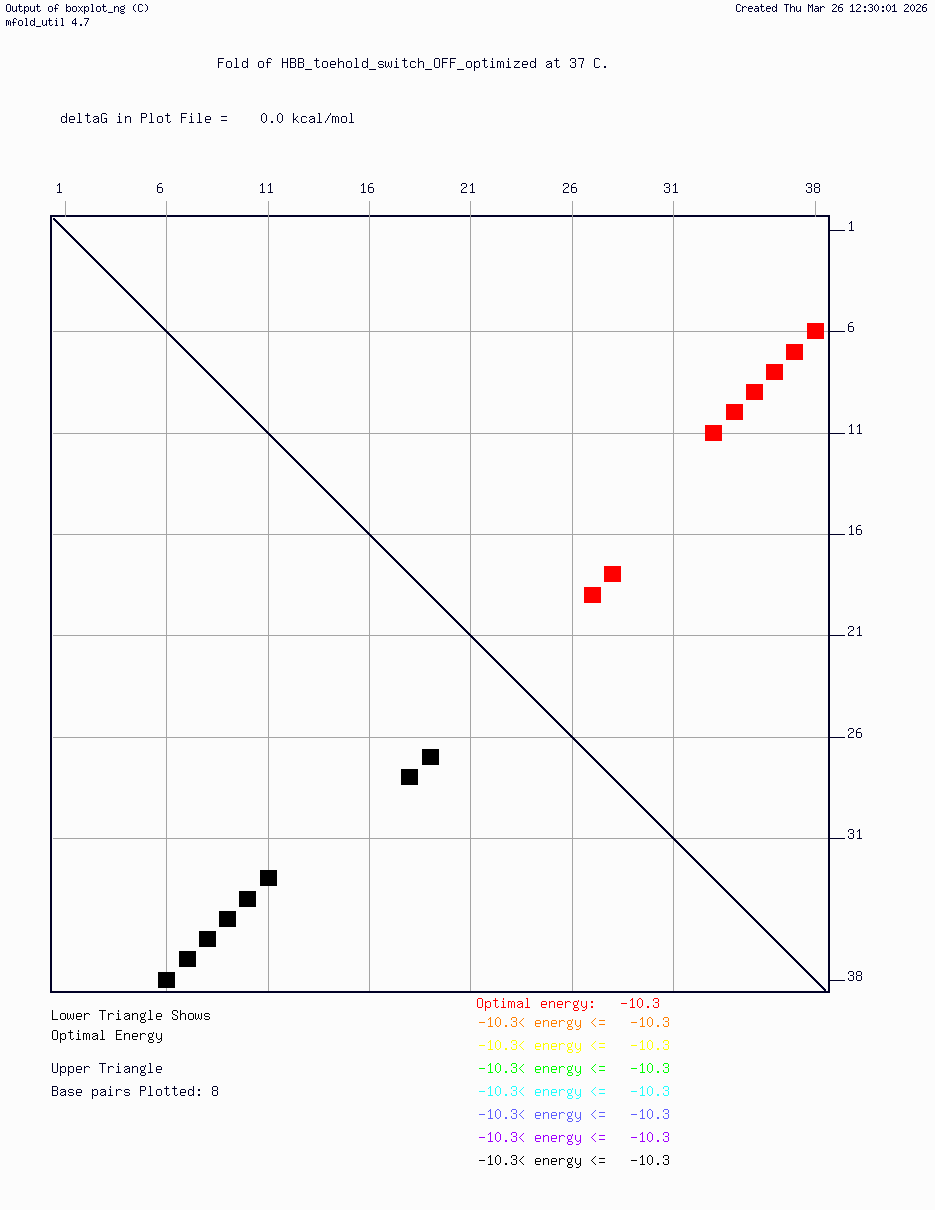
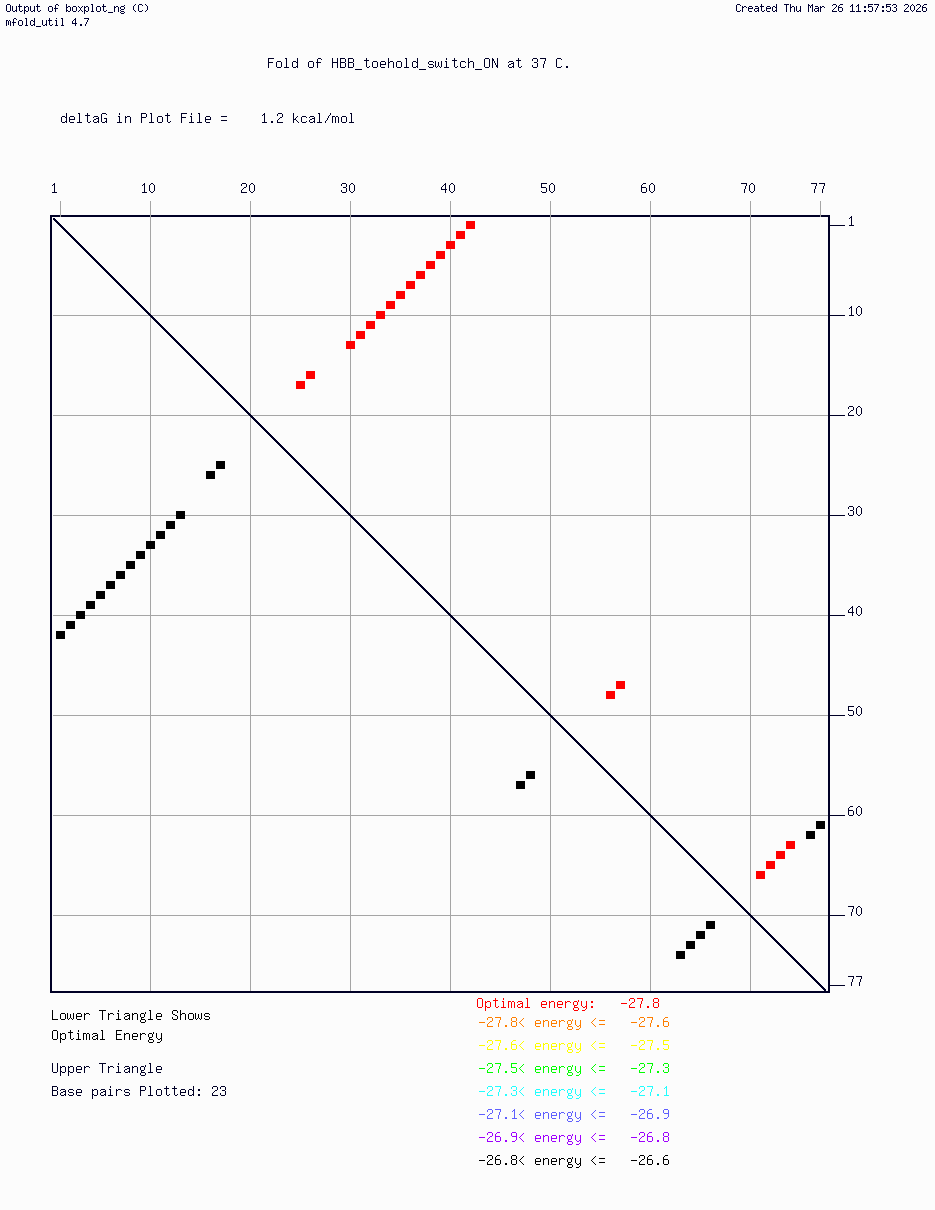
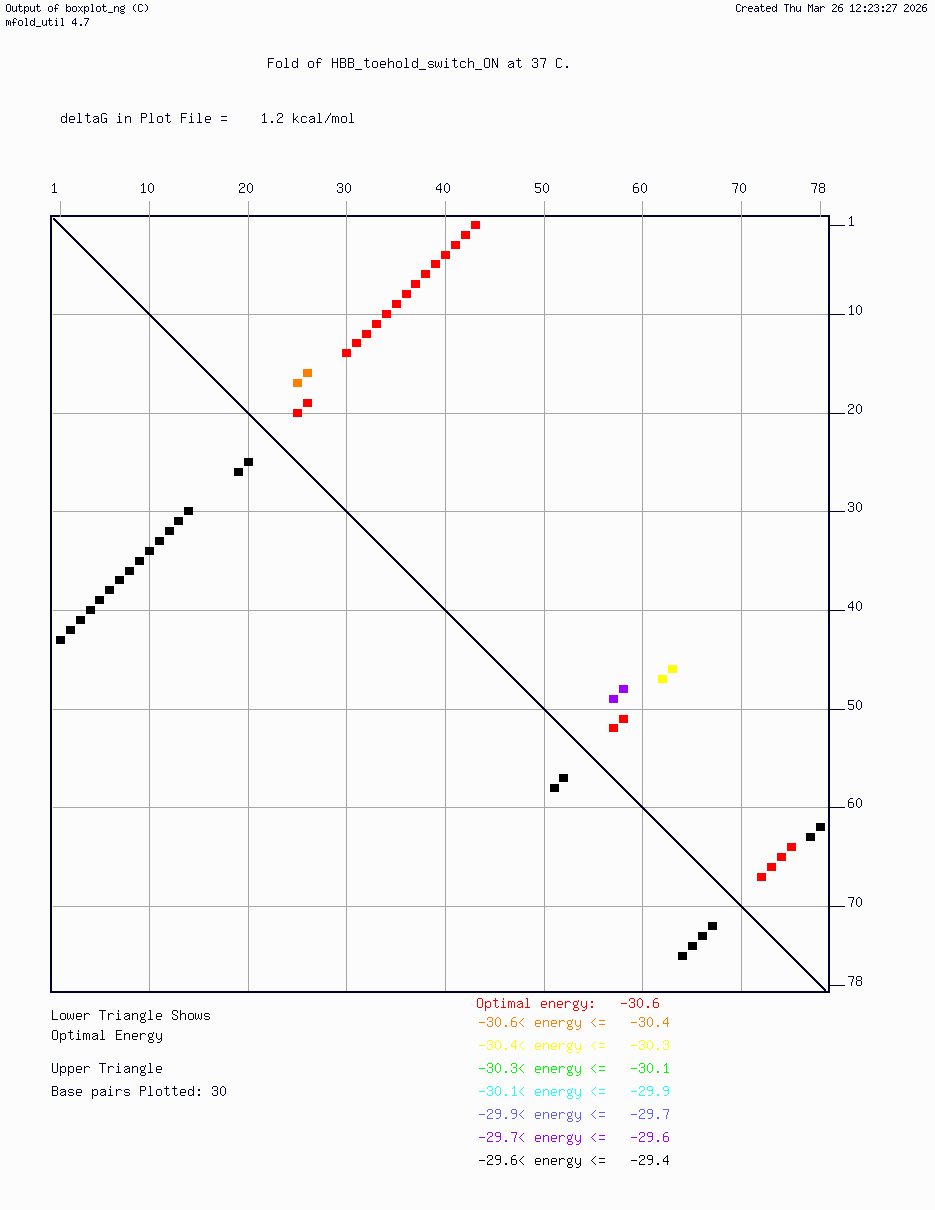
- HBB_Fr8-9_plusG_toehold_sfGFP_Version2 — iterated v2 construct following secondary structure validation at 37°C using mfold/UNAFold, confirming hairpin formation in the absence of trigger and RBS exposure upon trigger binding
Computational validation:
Secondary structure modeling was performed at 37°C using UNAFold/mfold. Results confirmed correct hairpin formation in the absence of the trigger sequence and predicted RBS exposure upon trigger binding in silico. The v2 construct was selected for Twist submission based on improved predicted folding fidelity over v1.
Backbone vector:
The insert is to be synthesized in pTwist Amp High Copy, the same backbone used in the Week 2 GFP expression cassette (Promoter J23106 + RBS B0034 + insert + Terminator B0015). This is documented on the HTGAA website under the beta-thalassemia biosensor project page.
Expression system: Cell-free (CFPS)
DNA order delivery: Ginkgo Bioworks cloud laboratory
Sequences submitted to Twist Bioscience via the HTGAA shared submission pipeline. ✓