Week 2 HW: DNA Read, Write, and Edit
Part 1: Benchling & In-silico Gel Art
For this assignment, I performed a virtual restriction digest on Lambda DNA using Benchling. My creative goal was to produce a pattern that mirrors the anatomy of a scorpion, inspired by my final project on the BmK CT scorpion peptide.
Creative Vision: The Genetic Scorpion 🦂
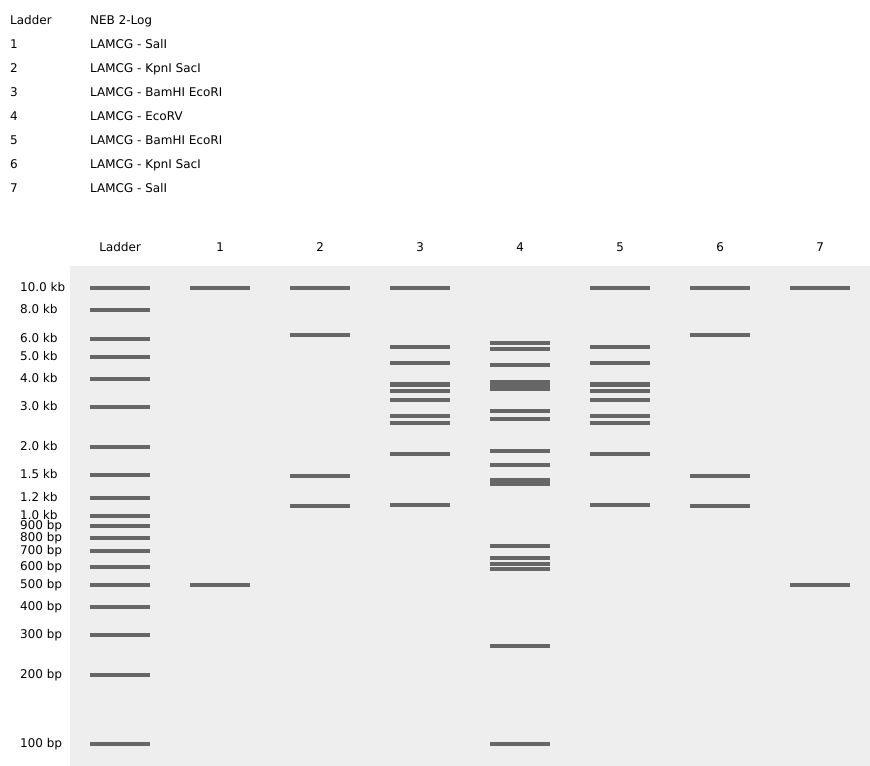
To achieve this symmetrical design, I strategically assigned different enzymes and double-digests across 7 lanes:
- Lanes 1 & 7 (The Pincers): I used SalI to create distinct high-molecular-weight bands.
- Lanes 2 & 6 (The Legs): A double digest of KpnI and SacI created a wider spread of bands, mimicking the scorpion’s legs.
- Lanes 3 & 5 (The Torso): Combining BamHI and EcoRI resulted in a dense cluster of bands to represent the main body segments.
- Lane 4 (The Stinger): I used EcoRV, which produces 21 fragments, creating a solid vertical pillar in the center that represents the scorpion’s iconic stinger.
Methodology
- Sequence: Imported the Lambda DNA (48,502 bp) sequence into Benchling.
- Analysis: Used the Restriction Analysis tool to map sites for EcoRI, HindIII, BamHI, KpnI, EcoRV, SacI, and SalI.
- Simulation: Conducted virtual gel electrophoresis using a NEB 2-Log Ladder as a size reference.
- Result: The final digest achieved a balanced, symmetrical pattern that visually aligns with my biotech research interests.

Part 3: DNA Design Challenge
3.1. Choose your protein
The protein I chose for this assignment is BmK CT, a chlorotoxin-like peptide derived from the venom of the Chinese scorpion Olivierus martensii (previously called Mesobuthus martensii). It is a small bioactive peptide which functions primarily as a targeted anti-tumor agent against glioma cells, with key roles in inhibiting cell migration, invasion, and proliferation.
What makes BmK CT especially promising is that it can promote apoptosis and enhance the sensitivity of glioma cells to chemotherapeutic agents like temozolomide. Importantly, studies suggest that it exhibits minimal toxicity toward normal glial cells, highlighting its potential as a selective therapeutic candidate for high-grade gliomas.
The sequence was obtained from UniProt (Entry: Q9UAD0). I focused on the biologically active mature peptide consisting of 35 amino acids:
CGPCFTTDANMARKCRECCGGIGKCFGPQCLCNRI
This peptide is cysteine-rich, suggesting the formation of multiple disulfide bonds that stabilize its tertiary structure.
3.2. Reverse Translate
Using the Reverse Translate tool from the Sequence Manipulation Suite, the following 105 bp DNA sequence (35 × 3 nucleotides) was generated:
tgcggcccgtgctttaccaccgatgcgaacatggcgcgcaaatgccgcgaatgctgcggcggcattggcaaatgctttggcccgcagtgcctgtgcaaccgcatt
3.3. Codon Optimization
When expressing a foreign gene in a host organism, it’s important to consider codon usage bias. This refers to the preference of the host to translate certain synonymous codons more efficiently than others. Codon optimization replaces some codons in the sequence with those preferred by the host, without changing the amino acid sequence, to ensure efficient translation.
For BmK CT, I chose Escherichia coli (K-12 strain MG1655) as the host because it is a standard, well-characterized chassis in synthetic biology that grows rapidly and is easy to handle. Using the Codon Optimization Tool from VectorBuilder, I optimized the sequence while avoiding Type IIs restriction enzyme sites (BsaI, Esp3I, and BbsI) to make the sequence compatible with modular cloning workflows.
Improved Sequence (CAI: 0.99, GC Content: 59.05%):
TGCGGCCCGTGCTTTACCACCGATGCGAATATGGCGCGTAAATGCCGCGAATGCTGCGGCGGCATTGGCAAATGCTTTGGCCCGCAGTGCCTGTGCAATCGCATT
3.4. You have a sequence! Now what?
Once the optimized gene is synthesized, it can be produced using a hybrid approach that combines both cell-dependent and cell-free systems.
Cell-Dependent Recombinant Expression
In a traditional approach, the optimized gene is cloned into an expression vector (like pET-28a) under a strong promoter and transformed into E. coli.
- Transcription: DNA is transcribed into mRNA by RNA polymerase.
- Translation: mRNA is translated by ribosomes into the peptide.
- Purification: The peptide is purified using chromatographic methods (e.g., IMAC via a His-tag).
Cell-Free Expression Systems
Alternatively, the optimized DNA can be directly introduced into a cell-free transcription–translation system. This offers:
- Rapid prototyping and reduced toxicity constraints.
- Precise control over reaction conditions.
- Compatibility with high-throughput screening and automated cloud-lab paradigms.
This hybrid approach ensures the scalability of living cells while providing the flexibility needed to produce potentially toxic bioactive peptides like BmK CT.
Part 4: Prepare a Twist DNA Synthesis Order (Practice)
In this practice exercise, I simulated the workflow for ordering a synthetic gene, moving from a custom expression cassette design in Benchling to a clonal gene construct on the Twist Bioscience portal.
Step 1: Account Creation
I successfully set up accounts on Twist Bioscience and Benchling to facilitate DNA design and synthesis simulation. ✅
Step 2: Build Your DNA Insert Sequence
I designed a complete Expression Cassette for my codon-optimized BmK CT sequence, optimized for an E. coli expression system. Using Benchling, I sequentially assembled the following components into a single linear DNA sequence:
- Promoter:
TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGC - RBS:
CATTAAAGAGGAGAAAGGTACC - Start Codon:
ATG - Coding Sequence (CDS): Codon-optimized BmK CT
- 7x His Tag:
CATCACCATCACCATCATCAC(to enable protein purification) - Stop Codon:
TAA - Terminator:
CCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA
Linear Map: View on Benchling
Step 3: Simulation on Twist Bioscience
I simulated the ordering process by selecting the “Genes” category and choosing the “Clonal Genes” option. Unlike gene fragments, clonal genes arrive already inserted into a circular vector, which allows for direct transformation into E. coli.
- Importing Sequence: I uploaded the FASTA file of the Benchling cassette.
- Vector Selection: I chose the pTwist Amp High Copy cloning vector as the circular backbone.
- Export: I downloaded the final construct as a GenBank (.gb) file for verification.
Recombinant Vector Overview: Below is the visualization of the final construct as it appears in the Twist portal:
Step 4: Final Plasmid Verification in Benchling
After re-importing the Twist-generated GenBank file back into Benchling, I verified the final circular plasmid containing my custom expression cassette.
Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) Target for Sequencing
I want to sequence the metagenomic DNA from the venom glands of Northern Pakistani scorpions (specifically species related to Mesobuthus martensii). Rationale: While BmK CT is documented, local Pakistani species might harbor unique genetic variants with higher therapeutic potency or different binding affinities for glioma cells. Sequencing this DNA helps explore local biodiversity and bioprospecting for novel drug precursors.
(ii) Sequencing Technology: Oxford Nanopore (ONT)
I have chosen Oxford Nanopore Technologies (ONT), a third-generation, single-molecule sequencing platform.
Input & Preparation:
- Material: High Molecular Weight (HMW) genomic DNA from venom gland tissue.
- Steps: 1. Optional fragmentation (kept long for ONT), 2. End-repair and A-tailing, 3. Adapter ligation with motor proteins, and 4. Tethering to the flow cell membrane.
Mechanism & Base Calling:
- DNA strands pass through a protein nanopore, causing characteristic disruptions in the ionic current.
- AI/ML algorithms decode these electrical signals into base sequences (A, T, G, C) in real-time.
Output:
- FASTQ files containing long-read sequences, ideal for variant analysis and de novo assembly.
5.2 DNA Write
(i) Target for Synthesis
I want to synthesize a codon-optimized genetic construct for expressing the BmK CT peptide in E. coli. Rationale: Synthesis allows the inclusion of a T7 promoter for high-level expression and a 6xHis-tag for streamlined purification. This is a direct therapeutic application for glioma drug discovery.
(ii) Synthesis Technology: Silicon-based Electrochemical Synthesis
I would utilize the technology implemented by Twist Bioscience.
Essential Steps:
- Phosphoramidite chemistry performed on a high-density silicon chip.
- Computer-controlled electrochemical activation of specific pixels to add nucleotides one at a time.
- Short synthesized oligos are harvested and assembled into the full-length gene.
Limitations:
- Error rates can increase with length, necessitating sequence verification.
- Large constructs (>3 kb) are significantly more challenging to synthesize directly.
5.3 DNA Edit
(i) Target for Editing
I want to edit the genome of E. coli expression strains (e.g., BL21) to enhance the secretion of BmK CT. Rationale: Normally, recombinant peptides accumulate in the periplasm; by editing secretion pathways or outer membrane proteins, the peptide could be secreted directly into the medium, simplifying downstream purification.
(ii) Editing Technology: CRISPR-Cas9
Mechanism & Essential Steps:
- Recognition: A custom Guide RNA (gRNA) targets the specific genomic sequence.
- Cleavage: The Cas9 enzyme generates a Double-Strand Break (DSB).
- Repair: Homology-Directed Repair (HDR) inserts the desired mutation using a provided donor template.
Preparation & Input:
- Design: Designing gRNA via Benchling to minimize off-target effects.
- Inputs: Cas9 enzyme/plasmid, custom gRNA, donor DNA template, and competent cells.
Limitations:
- Efficiency: HDR efficiency can be low in certain bacterial strains.
- Precision: Potential for off-target effects requires careful validation.
Sources & Acknowledgments
I used a combination of published literature, AI tools (including ChatGPT and Google Gemini), and discussions with peers to compile and refine this assignment.