Week 3 HW: Lab Automation
Assignment: Python Script for Opentrons Artwork
This section documents my process of creating bio-art of the Markhor, Pakistan’s national animal, using the Opentrons OT-2 liquid handling robot.
1. Image Generation
I started by using Google Gemini to generate a high-quality image of a Markhor with a clean white background to serve as the reference for my design.


2. Pixel Art Conversion
I then requested the AI to convert the high-resolution image into pixel art, which is necessary for the robot to dispense discrete “dots” of bacteria.
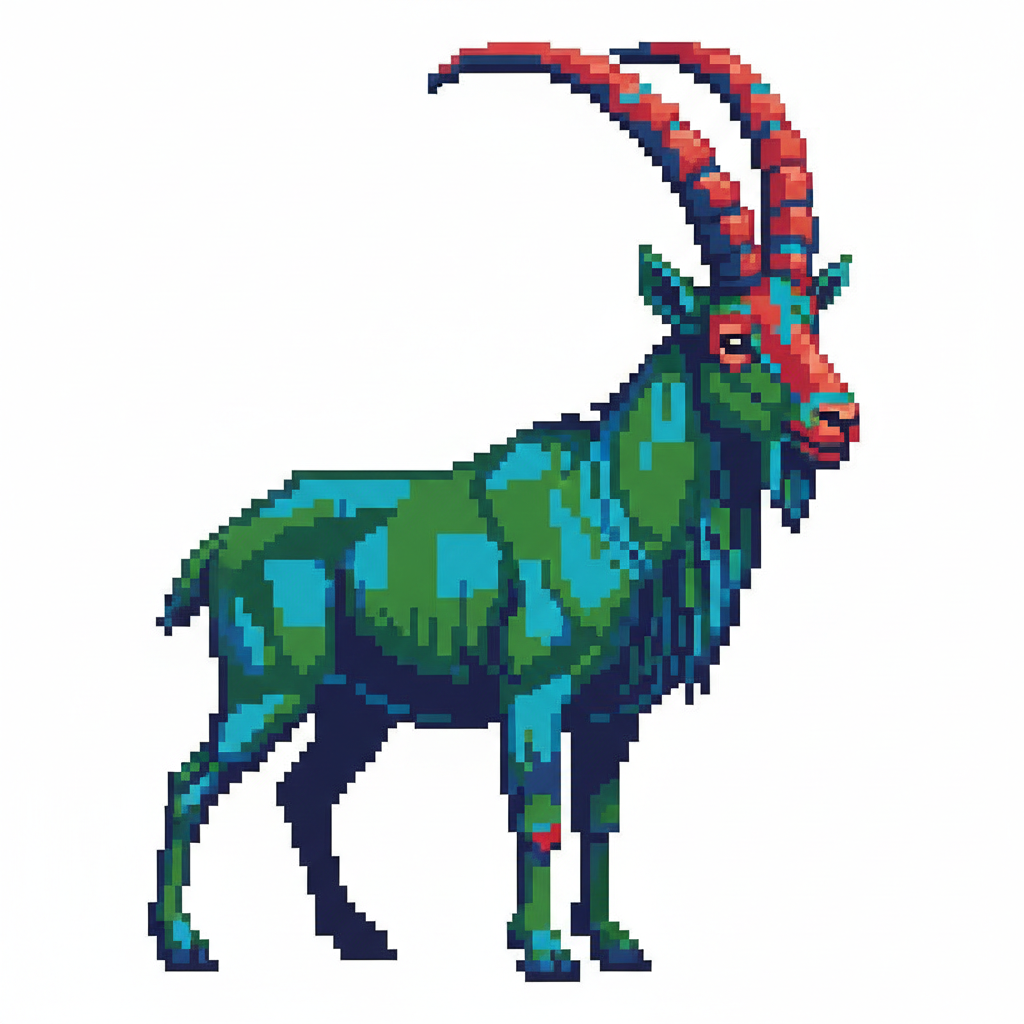


3. Coordinate Generation via GUI
I uploaded the pixel art to the Opentrons Art GUI. The initial generation included many colors, so I tweaked the design to limit it to three primary fluorescent colors: Blue, Green, and Red. During this stage, I also manually added the Markhor’s ears, which were not correctly captured in the original AI image.
- View My Design: Markhor on Opentrons-Art
4. Python Scripting & Debugging
Using the Echo CSV coordinates from the GUI, I utilized Google Gemini (both the web interface and the integrated assistant in the HTGAA Colab) to write the final Python script.
The process involved significant debugging due to:
- Color Swapping: Adjusting the mapping to ensure the robot pulled the correct color from the source wells (A1, B1, C1).
- API Syntax: Fixing
NameErrorandKeyErrorrelated to the labware setup and coordinate indexing in the simulator.
Finally, I successfully simulated the design, ensuring the Markhor pattern was centered and clearly defined.
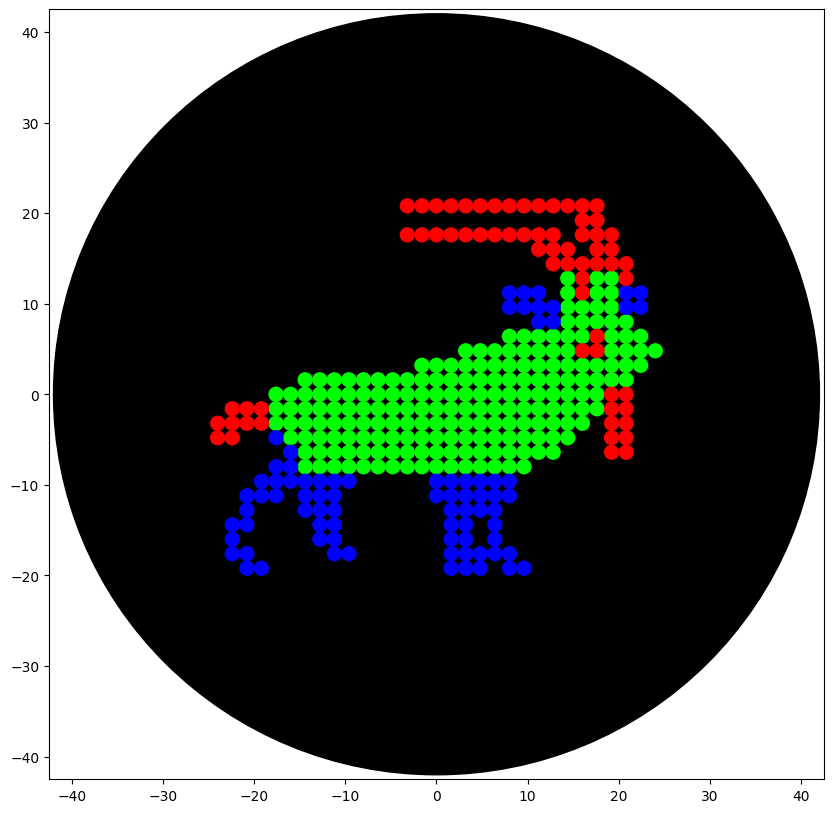

Post-Lab Questions: Research & Automation Strategy
1. Published Research Paper Analysis
I have reviewed the following paper regarding advanced laboratory automation:
Paper Name: Automating life science labs at the single-cell level through precise ultrasonic liquid sample ejection: PULSE
Link: https://doi.org/10.1038/s41378-024-00798-y
General Overview
- The paper introduces PULSE (Precise Ultrasonic Liquid Sample Ejection), a novel laboratory automation platform designed for biological experiments at the single-cell level.
- While current lab automation excels at bulk samples, it often lacks the precision and biocompatibility required for single-cell work, which can mask critical cellular heterogeneity.
- PULSE overcomes these limitations by using focused acoustic waves to eject nanoliter-sized droplets containing reagents or individual cells from a compact, disposable chip.
- This technology transforms standard titer plates into high-density microdroplet arrays, allowing researchers to programmably print “nanodrop pixels” with specific numbers of cells.
- The platform integrates software that synchronizes over 100 parameters, enabling automated and scalable workflows.
Major Applications
The authors demonstrate the versatility of PULSE across three major areas:
- Bio-fabrication: Precise deposition of different cell types to create 3D spheroids and patterned constructs.
- Precision Gating: Isolating single cells in nanodrops to track behavior over time (e.g., response to drugs) and retrieving them for downstream analysis.
- Deterministic Array Barcoding: Directly linking a single cell’s observable phenotype (via imaging) to its genotype (via sequencing) using pre-allocated, addressable primers.
Key Findings
- High Precision: The platform can print single cells at speeds of 5-20 cells per second with an accuracy of 90.5-97.7%.
- Biocompatibility: RNA sequencing revealed that the acoustic ejection process causes no significant differential gene regulation up to 24 hours post-treatment, outperforming FACS.
- Accuracy: In deterministic barcoding experiments, the platform achieved a 95.6% accuracy in matching cell fluorescent phenotypes to genotypes.
2. Final Project Ideas
I am exploring three core directions for my final project, focusing on recombinant production, metabolic engineering, and industrial biocatalysis.
Idea 1: Recombinant Production of BmK CT Peptide for Glioma Therapy
- The Concept: To engineer E. coli K-12 for the high-yield production of the 35-residue mature BmK CT scorpion peptide.
- The Approach: Utilizing Twist Bioscience for codon-optimized gene synthesis and pET-series vectors for efficient recombinant expression.
- Impact: Establishing a scalable bio-manufacturing pipeline for a peptide with high affinity for brain tumor cells to facilitate further oncological research.
Idea 2: CryoDesign Yak: AI-Engineered Cold-Active Enzymes
- The Concept: Harnessing the unique metagenomic diversity of the Himalayan Yak microbiome to engineer psychrophilic enzymes for energy-efficient industrial bioprocessing.
- The Approach: Utilizing ESM-2 Transformers for structural motif identification and ProteinMPNN for inverse folding to optimize catalytic loop flexibility.
- Impact: Enabling room-temperature industrial catalysis (15°C–25°C) to slash heating energy demand, contributing to the EU Net Zero 2050 goal.
Idea 3: Automated Bio-manufacturing of Bryostatin-1 Precursors
- The Concept: Engineering a microbial chassis for the sustainable production of Bryostatin-1, a potent marine-derived anti-cancer precursor.
- The Approach: Implementing the biosynthetic gene cluster into a laboratory-friendly host and using Opentrons automation for screening optimal metabolic flux.
- Impact: Transitioning from destructive marine harvesting to a sustainable, automated bio-manufacturing paradigm.
Lab Automation Strategies
Automation Strategy for Idea 2: CryoDesign Yak (Cold-Active Enzymes)
My automation strategy focuses on a Closed-Loop ‘Design-Build-Test’ cycle for validating AI-designed cold-adapted enzymes. Precision in thermal management is the primary constraint for this project.
- Build (Ginkgo Nebula):
- I will utilize the Echo Acoustic Liquid Handler to nanodisperse synthetic DNA variants into 384-well plates.
- This will be coupled with the Bravo Automated Liquid Handling Platform to stamp in chilled cell-free protein synthesis (CFPS) reagents.
- Test (Local Opentrons):
- Since psychrophilic enzymes are heat-labile, I will use a 3D-printed chilled-block holder (maintained at 10°C) on the Opentrons deck.
- Process Flow:
- Opentrons P20 will perform high-accuracy dispensing of chromogenic substrates into the expressed enzyme variants.
- The Inheco Thermoshaker module will be used for controlled incubation at ambient ‘cold-start’ temperatures (15°C).
- Data will be captured via an integrated plate reader to measure the kinetic rate ($\Delta$ Abs / $\Delta$ time).
Automation Strategy for Idea 3: Bio-manufacturing of Bryostatin-1 Precursors
This strategy aims to automate the metabolic engineering and screening of Bryostatin-1 precursors in a laboratory-friendly microbial host. Since the biosynthetic gene cluster (BGC) is large and complex, automation is essential to find the optimal expression balance.
- Design & Build (Ginkgo Nebula):
- I will utilize the Echo Acoustic Liquid Handler to assemble combinatorial libraries of BGC components, including promoters, RBS, and biosynthetic modules.
- The Bravo Automated Liquid Handling Platform will be used for rapid transformation and plating of the engineered chassis.
- Test & Optimize (Local Opentrons):
- I will use the Opentrons OT-2 to automate the metabolic screening of small-molecule precursors.
- Process Flow:
- Opentrons P300 will handle the precise serial dilution of feeding precursors (e.g., acetate/propionate units) into the culture media.
- The Heater-Shaker Module will be programmed for automated induction and micro-fermentation at optimal temperatures.
- Post-fermentation, the Opentrons will perform an automated extraction protocol (solvent addition and supernatant collection) for downstream analysis via an integrated HPLC or plate reader.