Week 6 Genetic Circuits Part I: Assembly Technologies
Assignment: DNA Assembly
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion High-Fidelity PCR Master Mix is a concentrated, 2X formulation designed for high-performance DNA amplification. Its primary components include:
- Phusion DNA Polymerase: A recombinant enzyme consisting of a Pyrococcus-like proofreading polymerase fused to a dsDNA-binding domain (Sso7d). This fusion technology enhances processivity, allowing the enzyme to synthesize long fragments rapidly while maintaining a low error rate (50 × lower than Taq).
- Deoxynucleotide Triphosphates (dNTPs): Equimolar concentrations of dATP, dCTP, dGTP, and dTTP serve as the chemical precursors for the nascent DNA strand.
- Magnesium Chloride (MgCl2): A critical divalent cation cofactor. Mg2+ ions coordinate with the phosphate groups of the dNTPs and the enzyme’s active site, facilitating the nucleophilic attack of the 3’-OH primer end on the incoming nucleotide.
- Reaction Buffers (HF or GC): These maintain optimal pH (8.8–9.3) and ionic strength. The HF (High-Fidelity) Buffer is the standard for most templates, while the GC Buffer contains additives (like DMSO) that lower the melting temperature of DNA to assist in the denaturation of templates with secondary structures or high GC content.
2. What are some factors that determine primer annealing temperature during PCR?
The optimal annealing temperature is vital for balancing primer specificity with yield. It is dictated by: Melting Temperature (T_m): The temperature at which 50% of the primer-template duplex is dissociated. Annealing temperature is typically set 3–5 °C below melting temperature.
- Base Composition (GC Content): Because Guanine-Cytosine pairs involve three hydrogen bonds compared to two for Adenine-Thymine, higher GC content increases the thermal stability of the primer.
- Primer Length: Longer sequences possess higher cumulative hydrogen bonding, raising the energy required for dissociation.
- Salt Concentration: Monovalent (K+) and divalent (Mg2+) cations neutralize the negatively charged phosphate backbone of DNA, reducing electrostatic repulsion and stabilizing the duplex, which effectively raises the melting temperature.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
Both methods yield linear DNA fragments, but they differ fundamentally in mechanism and application:
| Feature | PCR Amplification | Restriction Enzyme Digest |
|---|---|---|
| Mechanism | De novo enzymatic synthesis of specific DNA regions. | Hydrolysis of phosphodiester bonds at specific recognition sites. |
| Input Material | Requires only trace amounts of template DNA. | Requires a significant mass of pre-existing DNA (e.g., plasmid). |
| Product Ends | Typically blunt-ended (unless primers include overhangs). | Can produce sticky (overhanging) or blunt ends depending on the enzyme. |
| Precision | High flexibility; primers can be designed for any sequence. | Limited by the presence of specific palindromic recognition sites. |
| Primary Use | Scaling DNA quantity and adding functional “tails.” | Cloning into vectors or diagnostic mapping of DNA fragments. |
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
To ensure compatibility with Gibson Assembly, fragments must satisfy the following criteria:
- Overlapping Homology: Fragments must share 20–40 bp of identical sequence at their termini. For PCR products, these overlaps are engineered into the 5’ ends of the primers.
- Sequence Purity: PCR products must be purified (via column or gel extraction) to remove residual primers and dNTPs, which could interfere with the Gibson Master Mix’s precise ratio of exonuclease and polymerase activity.
- End Compatibility: While Gibson can join blunt or sticky ends, the presence of the correct homologous overlap is the only requirement for the 5’ → 3’ exonuclease to “chew back” the DNA and allow for annealing.
5. How does the plasmid DNA enter the E. coli cells during transformation?
Plasmid DNA enters E. coli cells through two primary laboratory methods:
- Chemical Transformation (Heat Shock): Cells are treated with Calcium Chloride to coat the negatively charged cell membrane with Ca2+ ions, facilitating DNA adhesion. A rapid increase in temperature 42 °C creates a thermal imbalance that induces the formation of transient pores in the membrane, through which the DNA-calcium complex is internalized.
- Electroporation: Cells are subjected to a high-voltage electric pulse (typically 1.8–2.5 kV). This causes dielectric breakdown of the cellular membrane, creating localized “nanopores” that allow the plasmid DNA to enter the cytoplasm via electrophoresis.
6. Describe another assembly method in detail (such as Golden Gate Assembly).
Golden Gate Assembly is a highly efficient molecular cloning technique that allows for the simultaneous, directional assembly of multiple DNA fragments into a single vector. This method relies on Type IIS restriction enzymes (such as BsaI or BpiI), which are unique because they cleave DNA at a specific distance outside of their non-palindromic recognition sequences. By strategically placing these recognition sites at the ends of DNA fragments, the enzyme generates unique, user-defined 4-base overhangs (sticky ends) that are eliminated from the final product during the reaction. Because the recognition sites are removed during cleavage, the desired assembly is “lossless” and cannot be re-cut by the enzyme, effectively driving the reaction toward the final circular product. The process occurs in a single-tube “one-pot” reaction where restriction digestion and T4 DNA ligation happen concurrently through biochemical cycling. This method is particularly favored for synthetic biology and modular cloning (MoClo) because it allows for the seamless “scarless” joining of many parts with nearly 100% efficiency.
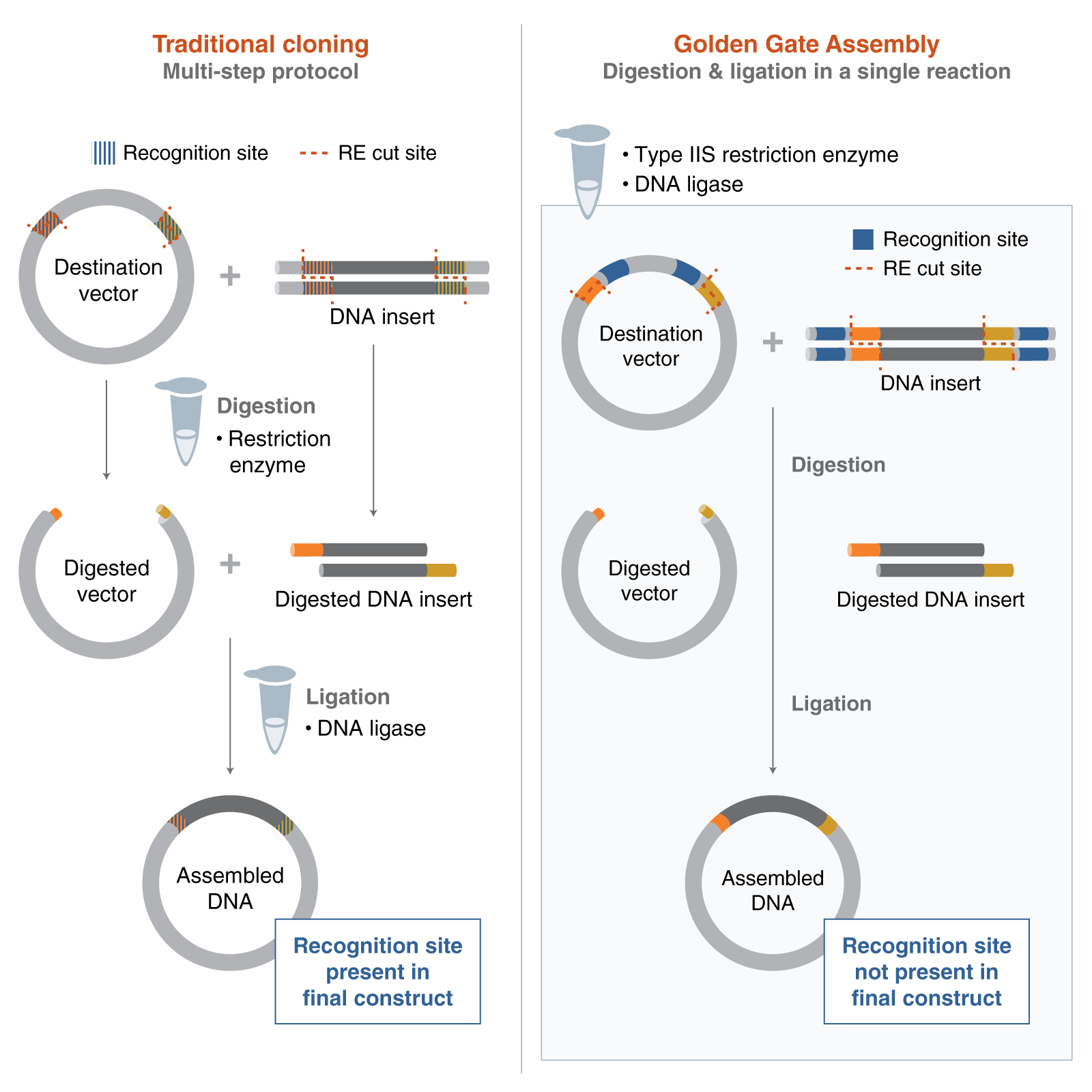

The Mechanism of Type IIS Cleavage
The key to Golden Gate is that the enzyme binds to a recognition site but cuts a few nucleotides away. This allows you to design the “overhang” to be whatever sequence you need for perfect complementarity with the next fragment.
The “One-Pot” Reaction Cycle
Because the ligation of the correct fragments destroys the original restriction sites, the reaction is essentially a “one-way street.” If the enzyme cuts a fragment and it ligates back to its original orientation, the site is reformed and the enzyme will simply cut it again. Once the correct assembly is formed, the sites are gone, and the product remains stable.
Comparison: Golden Gate vs. Gibson Assembly
While Gibson Assembly uses overlaps and “chew-back” enzymes, Golden Gate uses specific 4-base sticky ends. This makes Golden Gate extremely reliable for assembling very small fragments or repetitive sequences that might confuse the Gibson exonuclease.