Week 3 HW: Lab Automation
Python Script
Having no prior experience in python, I decided to use TA Donovan’s website. I chose the mRFP1 Bacteria to design the Transformers Autobots logo on the agar plate. I manually drew the desired shape and manipulated using only the cursor. I ended up with the following:
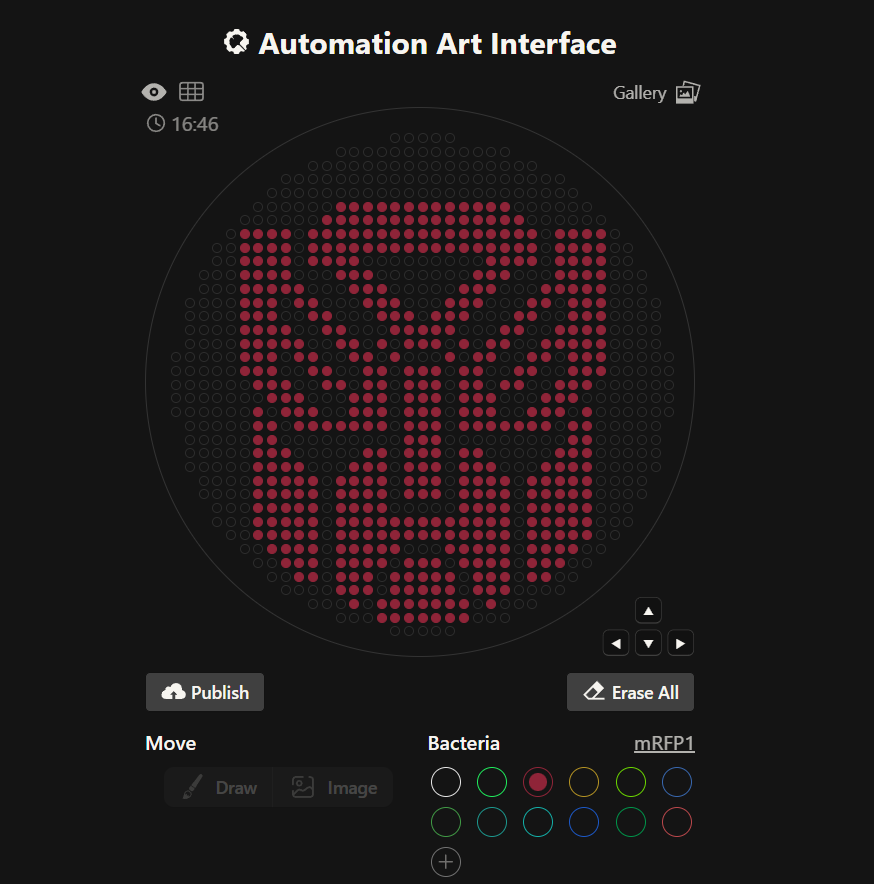

After making the logo of one faction, I decided it would be fun to attempt again with the Transformers Decepticons logo, as well. I chose the Azurite bacteria for this one. Again after careful trial and error (especially to make diagonal lines), I finished with the following:
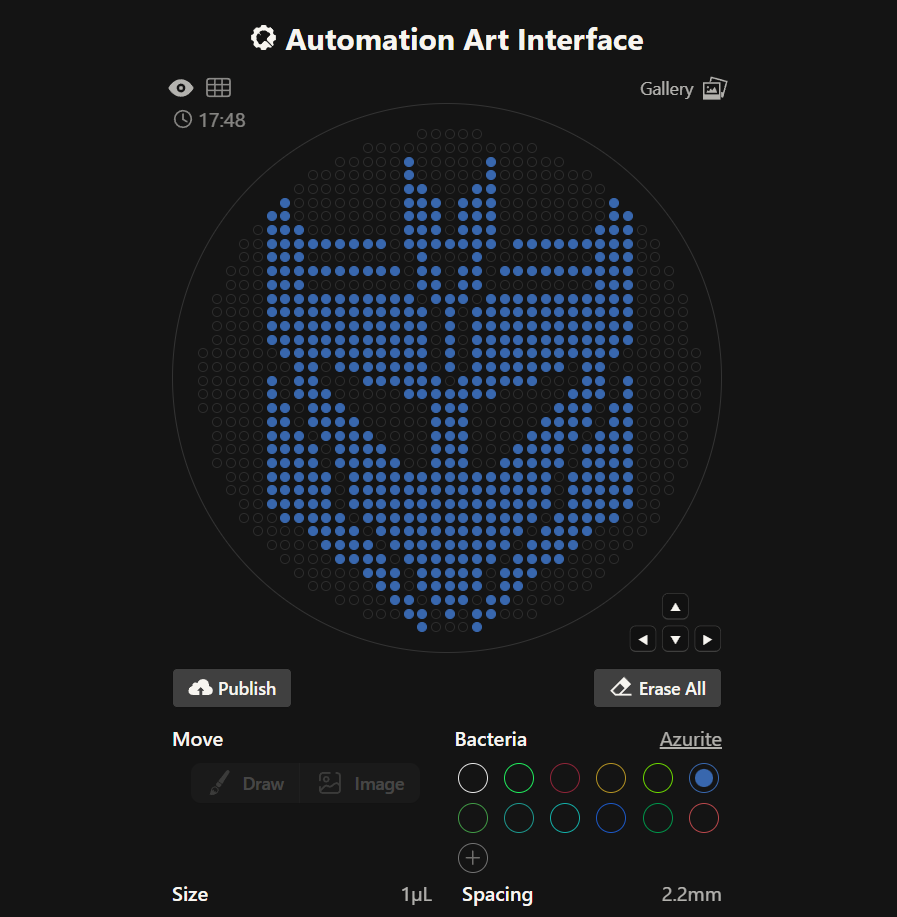

- Script for Design 1 on a 96 Deep-Well Plate:
from opentrons import types
import string
metadata = {
'protocolName': '{YOUR NAME} - Opentrons Art - HTGAA',
'author': 'HTGAA',
'source': 'HTGAA 2026',
'apiLevel': '2.20'
}
Z_VALUE_AGAR = 2.0
POINT_SIZE = 1
mrfp1_points = [(-13.2,28.6), (-11,28.6), (-8.8,28.6), (-6.6,28.6), (-4.4,28.6), (-2.2,28.6), (0,28.6), (2.2,28.6), (4.4,28.6), (6.6,28.6), (8.8,28.6), (11,28.6), (13.2,28.6), (-15.4,26.4), (-13.2,26.4), (-11,26.4), (-8.8,26.4), (-6.6,26.4), (-4.4,26.4), (-2.2,26.4), (0,26.4), (2.2,26.4), (4.4,26.4), (6.6,26.4), (8.8,26.4), (11,26.4), (13.2,26.4), (15.4,26.4), (-28.6,24.2), (-26.4,24.2), (-24.2,24.2), (-22,24.2), (-17.6,24.2), (-15.4,24.2), (-13.2,24.2), (-11,24.2), (-8.8,24.2), (-6.6,24.2), (-4.4,24.2), (-2.2,24.2), (0,24.2), (2.2,24.2), (4.4,24.2), (6.6,24.2), (8.8,24.2), (11,24.2), (13.2,24.2), (15.4,24.2), (17.6,24.2), (22,24.2), (24.2,24.2), (26.4,24.2), (28.6,24.2), (-28.6,22), (-26.4,22), (-24.2,22), (-22,22), (-17.6,22), (-15.4,22), (-13.2,22), (-11,22), (-8.8,22), (-6.6,22), (-4.4,22), (-2.2,22), (0,22), (2.2,22), (4.4,22), (6.6,22), (8.8,22), (11,22), (13.2,22), (15.4,22), (17.6,22), (22,22), (24.2,22), (26.4,22), (28.6,22), (-28.6,19.8), (-26.4,19.8), (-24.2,19.8), (-22,19.8), (-17.6,19.8), (-15.4,19.8), (-13.2,19.8), (-11,19.8), (11,19.8), (13.2,19.8), (15.4,19.8), (17.6,19.8), (22,19.8), (24.2,19.8), (26.4,19.8), (28.6,19.8), (-28.6,17.6), (-26.4,17.6), (-24.2,17.6), (-22,17.6), (-13.2,17.6), (-11,17.6), (-8.8,17.6), (8.8,17.6), (11,17.6), (13.2,17.6), (22,17.6), (24.2,17.6), (26.4,17.6), (28.6,17.6), (-28.6,15.4), (-26.4,15.4), (-24.2,15.4), (-22,15.4), (-19.8,15.4), (-11,15.4), (-8.8,15.4), (-6.6,15.4), (6.6,15.4), (8.8,15.4), (11,15.4), (19.8,15.4), (22,15.4), (24.2,15.4), (26.4,15.4), (28.6,15.4), (-28.6,13.2), (-26.4,13.2), (-24.2,13.2), (-19.8,13.2), (-17.6,13.2), (-8.8,13.2), (-6.6,13.2), (-4.4,13.2), (4.4,13.2), (6.6,13.2), (8.8,13.2), (17.6,13.2), (19.8,13.2), (24.2,13.2), (26.4,13.2), (28.6,13.2), (-28.6,11), (-26.4,11), (-24.2,11), (-17.6,11), (-15.4,11), (-6.6,11), (-4.4,11), (-2.2,11), (2.2,11), (4.4,11), (6.6,11), (15.4,11), (17.6,11), (24.2,11), (26.4,11), (28.6,11), (-28.6,8.8), (-26.4,8.8), (-24.2,8.8), (-22,8.8), (-15.4,8.8), (-13.2,8.8), (-4.4,8.8), (-2.2,8.8), (0,8.8), (2.2,8.8), (4.4,8.8), (13.2,8.8), (15.4,8.8), (22,8.8), (24.2,8.8), (26.4,8.8), (28.6,8.8), (-28.6,6.6), (-26.4,6.6), (-24.2,6.6), (-22,6.6), (-19.8,6.6), (-13.2,6.6), (-11,6.6), (-6.6,6.6), (-2.2,6.6), (0,6.6), (2.2,6.6), (6.6,6.6), (11,6.6), (13.2,6.6), (19.8,6.6), (22,6.6), (24.2,6.6), (26.4,6.6), (28.6,6.6), (-28.6,4.4), (-26.4,4.4), (-24.2,4.4), (-19.8,4.4), (-17.6,4.4), (-11,4.4), (-8.8,4.4), (-4.4,4.4), (4.4,4.4), (8.8,4.4), (11,4.4), (17.6,4.4), (19.8,4.4), (24.2,4.4), (26.4,4.4), (28.6,4.4), (-28.6,2.2), (-26.4,2.2), (-24.2,2.2), (-17.6,2.2), (-15.4,2.2), (-8.8,2.2), (-6.6,2.2), (-2.2,2.2), (0,2.2), (2.2,2.2), (6.6,2.2), (8.8,2.2), (15.4,2.2), (17.6,2.2), (24.2,2.2), (26.4,2.2), (28.6,2.2), (-26.4,0), (-24.2,0), (-22,0), (-15.4,0), (-13.2,0), (-8.8,0), (-6.6,0), (-2.2,0), (0,0), (2.2,0), (6.6,0), (8.8,0), (13.2,0), (15.4,0), (22,0), (24.2,0), (26.4,0), (-24.2,-2.2), (-22,-2.2), (-19.8,-2.2), (-11,-2.2), (-8.8,-2.2), (-6.6,-2.2), (-2.2,-2.2), (0,-2.2), (2.2,-2.2), (6.6,-2.2), (8.8,-2.2), (11,-2.2), (19.8,-2.2), (22,-2.2), (24.2,-2.2), (-26.4,-4.4), (-22,-4.4), (-19.8,-4.4), (-17.6,-4.4), (-11,-4.4), (-8.8,-4.4), (-6.6,-4.4), (-2.2,-4.4), (0,-4.4), (2.2,-4.4), (6.6,-4.4), (8.8,-4.4), (11,-4.4), (17.6,-4.4), (19.8,-4.4), (22,-4.4), (26.4,-4.4), (-26.4,-6.6), (-24.2,-6.6), (-19.8,-6.6), (-17.6,-6.6), (-15.4,-6.6), (-13.2,-6.6), (-11,-6.6), (-8.8,-6.6), (-6.6,-6.6), (-2.2,-6.6), (0,-6.6), (2.2,-6.6), (6.6,-6.6), (8.8,-6.6), (11,-6.6), (13.2,-6.6), (15.4,-6.6), (17.6,-6.6), (19.8,-6.6), (24.2,-6.6), (26.4,-6.6), (-26.4,-8.8), (-24.2,-8.8), (-2.2,-8.8), (0,-8.8), (2.2,-8.8), (24.2,-8.8), (26.4,-8.8), (-26.4,-11), (-24.2,-11), (-22,-11), (-8.8,-11), (-6.6,-11), (-2.2,-11), (0,-11), (2.2,-11), (6.6,-11), (8.8,-11), (22,-11), (24.2,-11), (26.4,-11), (-26.4,-13.2), (-24.2,-13.2), (-22,-13.2), (-19.8,-13.2), (-11,-13.2), (-8.8,-13.2), (-6.6,-13.2), (-2.2,-13.2), (0,-13.2), (2.2,-13.2), (6.6,-13.2), (8.8,-13.2), (11,-13.2), (19.8,-13.2), (22,-13.2), (24.2,-13.2), (26.4,-13.2), (-26.4,-15.4), (-24.2,-15.4), (-22,-15.4), (-19.8,-15.4), (-17.6,-15.4), (-13.2,-15.4), (-11,-15.4), (-8.8,-15.4), (-6.6,-15.4), (-2.2,-15.4), (0,-15.4), (2.2,-15.4), (6.6,-15.4), (8.8,-15.4), (11,-15.4), (13.2,-15.4), (17.6,-15.4), (19.8,-15.4), (22,-15.4), (24.2,-15.4), (26.4,-15.4), (-26.4,-17.6), (-24.2,-17.6), (-22,-17.6), (-19.8,-17.6), (-17.6,-17.6), (-13.2,-17.6), (-11,-17.6), (-8.8,-17.6), (-6.6,-17.6), (-2.2,-17.6), (0,-17.6), (2.2,-17.6), (6.6,-17.6), (8.8,-17.6), (11,-17.6), (13.2,-17.6), (17.6,-17.6), (19.8,-17.6), (22,-17.6), (24.2,-17.6), (26.4,-17.6), (-26.4,-19.8), (-24.2,-19.8), (-22,-19.8), (-19.8,-19.8), (-17.6,-19.8), (-13.2,-19.8), (-11,-19.8), (-8.8,-19.8), (-6.6,-19.8), (6.6,-19.8), (8.8,-19.8), (11,-19.8), (13.2,-19.8), (17.6,-19.8), (19.8,-19.8), (22,-19.8), (24.2,-19.8), (26.4,-19.8), (-26.4,-22), (-24.2,-22), (-22,-22), (-19.8,-22), (-17.6,-22), (-13.2,-22), (-11,-22), (-8.8,-22), (-6.6,-22), (-4.4,-22), (-2.2,-22), (0,-22), (2.2,-22), (4.4,-22), (6.6,-22), (8.8,-22), (11,-22), (13.2,-22), (17.6,-22), (19.8,-22), (22,-22), (24.2,-22), (26.4,-22), (-26.4,-24.2), (-24.2,-24.2), (-22,-24.2), (-19.8,-24.2), (-17.6,-24.2), (-13.2,-24.2), (-11,-24.2), (-8.8,-24.2), (-6.6,-24.2), (-4.4,-24.2), (-2.2,-24.2), (0,-24.2), (2.2,-24.2), (4.4,-24.2), (6.6,-24.2), (8.8,-24.2), (11,-24.2), (13.2,-24.2), (17.6,-24.2), (19.8,-24.2), (22,-24.2), (24.2,-24.2), (26.4,-24.2), (-24.2,-26.4), (-22,-26.4), (-19.8,-26.4), (-17.6,-26.4), (-13.2,-26.4), (-11,-26.4), (-8.8,-26.4), (-6.6,-26.4), (-4.4,-26.4), (4.4,-26.4), (6.6,-26.4), (8.8,-26.4), (11,-26.4), (13.2,-26.4), (17.6,-26.4), (19.8,-26.4), (22,-26.4), (24.2,-26.4), (-22,-28.6), (-19.8,-28.6), (-17.6,-28.6), (-13.2,-28.6), (-11,-28.6), (-8.8,-28.6), (-6.6,-28.6), (-2.2,-28.6), (0,-28.6), (2.2,-28.6), (6.6,-28.6), (8.8,-28.6), (11,-28.6), (13.2,-28.6), (17.6,-28.6), (19.8,-28.6), (22,-28.6), (-19.8,-30.8), (-17.6,-30.8), (-13.2,-30.8), (-11,-30.8), (-8.8,-30.8), (-4.4,-30.8), (-2.2,-30.8), (0,-30.8), (2.2,-30.8), (4.4,-30.8), (8.8,-30.8), (11,-30.8), (13.2,-30.8), (17.6,-30.8), (19.8,-30.8), (-13.2,-33), (-11,-33), (-8.8,-33), (-4.4,-33), (-2.2,-33), (0,-33), (2.2,-33), (4.4,-33), (8.8,-33), (11,-33), (13.2,-33), (-11,-35.2), (-6.6,-35.2), (-4.4,-35.2), (-2.2,-35.2), (0,-35.2), (2.2,-35.2), (4.4,-35.2), (6.6,-35.2), (11,-35.2), (-6.6,-37.4), (-4.4,-37.4), (-2.2,-37.4), (0,-37.4), (2.2,-37.4), (4.4,-37.4), (6.6,-37.4)]
point_name_pairing = [("mrfp1", mrfp1_points)]
# Robot deck setup constants
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
# Place the PCR tubes in this order
well_colors = {
'A1': 'sfGFP',
'A2': 'mRFP1',
'A3': 'mKO2',
'A4': 'Venus',
'A5': 'mKate2_TF',
'A6': 'Azurite',
'A7': 'mCerulean3',
'A8': 'mClover3',
'A9': 'mJuniper',
'A10': 'mTurquoise2',
'A11': 'mBanana',
'A12': 'mPlum',
'B1': 'Electra2',
'B2': 'mWasabi',
'B3': 'mScarlet_I',
'B4': 'mPapaya',
'B5': 'eqFP578',
'B6': 'tdTomato',
'B7': 'DsRed',
'B8': 'mKate2',
'B9': 'EGFP',
'B10': 'mRuby2',
'B11': 'TagBFP',
'B12': 'mChartreuse_TF',
'C1': 'mLychee_TF',
'C2': 'mTagBFP2',
'C3': 'mEGFP',
'C4': 'mNeonGreen',
'C5': 'mAzamiGreen',
'C6': 'mWatermelon',
'C7': 'avGFP',
'C8': 'mCitrine',
'C9': 'mVenus',
'C10': 'mCherry',
'C11': 'mHoneydew',
'C12': 'TagRFP',
'D1': 'mTFP1',
'D2': 'Ultramarine',
'D3': 'ZsGreen1',
'D4': 'mMiCy',
'D5': 'mStayGold2',
'D6': 'PA_GFP'
}
volume_used = {
'mrfp1': 0
}
def update_volume_remaining(current_color, quantity_to_aspirate):
rows = string.ascii_uppercase
for well, color in list(well_colors.items()):
if color == current_color:
if (volume_used[current_color] + quantity_to_aspirate) > 250:
# Move to next well horizontally by advancing row letter, keeping column number
row = well[0]
col = well[1:]
# Find next row letter
next_row = rows[rows.index(row) + 1]
next_well = f"{next_row}{col}"
del well_colors[well]
well_colors[next_well] = current_color
volume_used[current_color] = quantity_to_aspirate
else:
volume_used[current_color] += quantity_to_aspirate
break
def run(protocol):
# Load labware, modules and pipettes
protocol.home()
# Tips
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
# Pipettes
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
# Deep Well Plate
temperature_plate = protocol.load_labware('nest_96_wellplate_2ml_deep', 6)
# Agar Plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate')
agar_plate.set_offset(x=0.00, y=0.00, z=Z_VALUE_AGAR)
# Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
# Helper function (dispensing)
def dispense_and_jog(pipette, volume, location):
assert(isinstance(volume, (int, float)))
# Go above the location
above_location = location.move(types.Point(z=location.point.z + 2))
pipette.move_to(above_location)
# Go downwards and dispense
pipette.dispense(volume, location)
# Go upwards to avoid smearing
pipette.move_to(above_location)
# Helper function (color location)
def location_of_color(color_string):
for well,color in well_colors.items():
if color.lower() == color_string.lower():
return temperature_plate[well]
raise ValueError(f"No well found with color {color_string}")
# Print pattern by iterating over lists
for i, (current_color, point_list) in enumerate(point_name_pairing):
# Skip the rest of the loop if the list is empty
if not point_list:
continue
# Get the tip for this run, set the bacteria color, and the aspirate bacteria of choice
pipette_20ul.pick_up_tip()
max_aspirate = int(18 // POINT_SIZE) * POINT_SIZE
quantity_to_aspirate = min(len(point_list)*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Iterate over the current points list and dispense them, refilling along the way
for i in range(len(point_list)):
x, y = point_list[i]
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_jog(pipette_20ul, POINT_SIZE, adjusted_location)
if pipette_20ul.current_volume == 0 and len(point_list[i+1:]) > 0:
quantity_to_aspirate = min(len(point_list[i:])*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Drop tip between each color
pipette_20ul.drop_tip()- Script for Design 2 on a 96 Deep-Well Plate:
from opentrons import types
import string
metadata = {
'protocolName': '{YOUR NAME} - Opentrons Art - HTGAA',
'author': 'HTGAA',
'source': 'HTGAA 2026',
'apiLevel': '2.20'
}
Z_VALUE_AGAR = 2.0
POINT_SIZE = 1
azurite_points = [(-6.6,35.2), (6.6,35.2), (-6.6,33), (6.6,33), (-6.6,30.8), (-4.4,30.8), (4.4,30.8), (6.6,30.8), (-26.4,28.6), (-6.6,28.6), (-4.4,28.6), (-2.2,28.6), (2.2,28.6), (4.4,28.6), (6.6,28.6), (26.4,28.6), (-28.6,26.4), (-26.4,26.4), (-6.6,26.4), (-4.4,26.4), (-2.2,26.4), (2.2,26.4), (4.4,26.4), (6.6,26.4), (26.4,26.4), (28.6,26.4), (-28.6,24.2), (-26.4,24.2), (-24.2,24.2), (-6.6,24.2), (-4.4,24.2), (-2.2,24.2), (2.2,24.2), (4.4,24.2), (6.6,24.2), (24.2,24.2), (26.4,24.2), (28.6,24.2), (-28.6,22), (-26.4,22), (-24.2,22), (-22,22), (-19.8,22), (-17.6,22), (-15.4,22), (-13.2,22), (-11,22), (-6.6,22), (-4.4,22), (-2.2,22), (0,22), (2.2,22), (4.4,22), (6.6,22), (11,22), (13.2,22), (15.4,22), (17.6,22), (19.8,22), (22,22), (24.2,22), (26.4,22), (28.6,22), (-28.6,19.8), (-26.4,19.8), (-24.2,19.8), (-4.4,19.8), (4.4,19.8), (24.2,19.8), (26.4,19.8), (28.6,19.8), (-28.6,17.6), (-26.4,17.6), (-24.2,17.6), (-22,17.6), (-19.8,17.6), (-17.6,17.6), (-15.4,17.6), (-13.2,17.6), (-11,17.6), (-8.8,17.6), (-4.4,17.6), (-2.2,17.6), (2.2,17.6), (4.4,17.6), (8.8,17.6), (11,17.6), (13.2,17.6), (15.4,17.6), (17.6,17.6), (19.8,17.6), (22,17.6), (24.2,17.6), (26.4,17.6), (28.6,17.6), (-28.6,15.4), (-26.4,15.4), (-24.2,15.4), (-4.4,15.4), (-2.2,15.4), (2.2,15.4), (4.4,15.4), (24.2,15.4), (26.4,15.4), (28.6,15.4), (-28.6,13.2), (-26.4,13.2), (-24.2,13.2), (-22,13.2), (-19.8,13.2), (-17.6,13.2), (-15.4,13.2), (-13.2,13.2), (-11,13.2), (-8.8,13.2), (-6.6,13.2), (-2.2,13.2), (0,13.2), (2.2,13.2), (6.6,13.2), (8.8,13.2), (11,13.2), (13.2,13.2), (15.4,13.2), (17.6,13.2), (19.8,13.2), (22,13.2), (24.2,13.2), (26.4,13.2), (28.6,13.2), (-28.6,11), (-26.4,11), (-24.2,11), (-22,11), (-19.8,11), (-17.6,11), (-15.4,11), (-13.2,11), (-11,11), (-8.8,11), (-6.6,11), (-4.4,11), (0,11), (4.4,11), (6.6,11), (8.8,11), (11,11), (13.2,11), (15.4,11), (17.6,11), (19.8,11), (22,11), (24.2,11), (26.4,11), (28.6,11), (-28.6,8.8), (-26.4,8.8), (-24.2,8.8), (-22,8.8), (-19.8,8.8), (-17.6,8.8), (-15.4,8.8), (-13.2,8.8), (-11,8.8), (-8.8,8.8), (-6.6,8.8), (-4.4,8.8), (0,8.8), (4.4,8.8), (6.6,8.8), (8.8,8.8), (11,8.8), (13.2,8.8), (15.4,8.8), (17.6,8.8), (19.8,8.8), (22,8.8), (24.2,8.8), (26.4,8.8), (28.6,8.8), (-28.6,6.6), (-26.4,6.6), (-24.2,6.6), (-22,6.6), (-19.8,6.6), (-17.6,6.6), (-15.4,6.6), (-13.2,6.6), (-11,6.6), (-8.8,6.6), (-6.6,6.6), (-4.4,6.6), (0,6.6), (4.4,6.6), (6.6,6.6), (8.8,6.6), (11,6.6), (13.2,6.6), (15.4,6.6), (17.6,6.6), (19.8,6.6), (22,6.6), (24.2,6.6), (26.4,6.6), (28.6,6.6), (-26.4,4.4), (-24.2,4.4), (-22,4.4), (-19.8,4.4), (-17.6,4.4), (-15.4,4.4), (-13.2,4.4), (-11,4.4), (-8.8,4.4), (-6.6,4.4), (-4.4,4.4), (0,4.4), (4.4,4.4), (6.6,4.4), (8.8,4.4), (11,4.4), (13.2,4.4), (15.4,4.4), (17.6,4.4), (19.8,4.4), (22,4.4), (24.2,4.4), (26.4,4.4), (-24.2,2.2), (-22,2.2), (-17.6,2.2), (-15.4,2.2), (-13.2,2.2), (-11,2.2), (-8.8,2.2), (-6.6,2.2), (-4.4,2.2), (0,2.2), (4.4,2.2), (6.6,2.2), (8.8,2.2), (11,2.2), (13.2,2.2), (15.4,2.2), (17.6,2.2), (22,2.2), (24.2,2.2), (-28.6,0), (-24.2,0), (-22,0), (-13.2,0), (-11,0), (-8.8,0), (-6.6,0), (-4.4,0), (-2.2,0), (2.2,0), (4.4,0), (6.6,0), (8.8,0), (11,0), (13.2,0), (22,0), (24.2,0), (28.6,0), (-28.6,-2.2), (-24.2,-2.2), (-22,-2.2), (-19.8,-2.2), (-6.6,-2.2), (-4.4,-2.2), (-2.2,-2.2), (0,-2.2), (2.2,-2.2), (4.4,-2.2), (6.6,-2.2), (19.8,-2.2), (22,-2.2), (24.2,-2.2), (28.6,-2.2), (-28.6,-4.4), (-26.4,-4.4), (-22,-4.4), (-19.8,-4.4), (-17.6,-4.4), (-2.2,-4.4), (0,-4.4), (2.2,-4.4), (17.6,-4.4), (19.8,-4.4), (22,-4.4), (26.4,-4.4), (28.6,-4.4), (-28.6,-6.6), (-26.4,-6.6), (-22,-6.6), (-19.8,-6.6), (-17.6,-6.6), (-15.4,-6.6), (-2.2,-6.6), (0,-6.6), (2.2,-6.6), (15.4,-6.6), (17.6,-6.6), (19.8,-6.6), (22,-6.6), (26.4,-6.6), (28.6,-6.6), (-28.6,-8.8), (-26.4,-8.8), (-24.2,-8.8), (-19.8,-8.8), (-17.6,-8.8), (-15.4,-8.8), (-13.2,-8.8), (-2.2,-8.8), (0,-8.8), (2.2,-8.8), (13.2,-8.8), (15.4,-8.8), (17.6,-8.8), (19.8,-8.8), (24.2,-8.8), (26.4,-8.8), (28.6,-8.8), (-28.6,-11), (-26.4,-11), (-24.2,-11), (-22,-11), (-17.6,-11), (-15.4,-11), (-13.2,-11), (-11,-11), (-2.2,-11), (0,-11), (2.2,-11), (11,-11), (13.2,-11), (15.4,-11), (17.6,-11), (22,-11), (24.2,-11), (26.4,-11), (28.6,-11), (-28.6,-13.2), (-26.4,-13.2), (-24.2,-13.2), (-22,-13.2), (-17.6,-13.2), (-15.4,-13.2), (-13.2,-13.2), (-11,-13.2), (-8.8,-13.2), (-2.2,-13.2), (0,-13.2), (2.2,-13.2), (8.8,-13.2), (11,-13.2), (13.2,-13.2), (15.4,-13.2), (17.6,-13.2), (22,-13.2), (24.2,-13.2), (26.4,-13.2), (28.6,-13.2), (-28.6,-15.4), (-26.4,-15.4), (-24.2,-15.4), (-22,-15.4), (-19.8,-15.4), (-15.4,-15.4), (-13.2,-15.4), (-11,-15.4), (-8.8,-15.4), (-6.6,-15.4), (-4.4,-15.4), (-2.2,-15.4), (0,-15.4), (2.2,-15.4), (4.4,-15.4), (6.6,-15.4), (8.8,-15.4), (11,-15.4), (13.2,-15.4), (15.4,-15.4), (19.8,-15.4), (22,-15.4), (24.2,-15.4), (26.4,-15.4), (28.6,-15.4), (-28.6,-17.6), (-26.4,-17.6), (-24.2,-17.6), (-22,-17.6), (-19.8,-17.6), (-15.4,-17.6), (-13.2,-17.6), (-11,-17.6), (-8.8,-17.6), (-6.6,-17.6), (-4.4,-17.6), (-2.2,-17.6), (0,-17.6), (2.2,-17.6), (4.4,-17.6), (6.6,-17.6), (8.8,-17.6), (11,-17.6), (13.2,-17.6), (15.4,-17.6), (19.8,-17.6), (22,-17.6), (24.2,-17.6), (26.4,-17.6), (28.6,-17.6), (-28.6,-19.8), (-26.4,-19.8), (-24.2,-19.8), (-22,-19.8), (-19.8,-19.8), (-15.4,-19.8), (-13.2,-19.8), (-11,-19.8), (-8.8,-19.8), (-6.6,-19.8), (-4.4,-19.8), (-2.2,-19.8), (0,-19.8), (2.2,-19.8), (4.4,-19.8), (6.6,-19.8), (8.8,-19.8), (11,-19.8), (13.2,-19.8), (15.4,-19.8), (19.8,-19.8), (22,-19.8), (24.2,-19.8), (26.4,-19.8), (28.6,-19.8), (-26.4,-22), (-24.2,-22), (-22,-22), (-19.8,-22), (-17.6,-22), (-13.2,-22), (-11,-22), (-8.8,-22), (-6.6,-22), (-4.4,-22), (-2.2,-22), (0,-22), (2.2,-22), (4.4,-22), (6.6,-22), (8.8,-22), (11,-22), (13.2,-22), (17.6,-22), (19.8,-22), (22,-22), (24.2,-22), (26.4,-22), (-22,-24.2), (-19.8,-24.2), (-17.6,-24.2), (-15.4,-24.2), (-11,-24.2), (-8.8,-24.2), (-6.6,-24.2), (-4.4,-24.2), (-2.2,-24.2), (0,-24.2), (2.2,-24.2), (4.4,-24.2), (6.6,-24.2), (8.8,-24.2), (11,-24.2), (15.4,-24.2), (17.6,-24.2), (19.8,-24.2), (22,-24.2), (-19.8,-26.4), (-17.6,-26.4), (-15.4,-26.4), (-13.2,-26.4), (-8.8,-26.4), (-6.6,-26.4), (-4.4,-26.4), (-2.2,-26.4), (0,-26.4), (2.2,-26.4), (4.4,-26.4), (6.6,-26.4), (8.8,-26.4), (13.2,-26.4), (15.4,-26.4), (17.6,-26.4), (19.8,-26.4), (-17.6,-28.6), (-15.4,-28.6), (-13.2,-28.6), (-11,-28.6), (-6.6,-28.6), (-4.4,-28.6), (-2.2,-28.6), (0,-28.6), (2.2,-28.6), (4.4,-28.6), (6.6,-28.6), (11,-28.6), (13.2,-28.6), (15.4,-28.6), (17.6,-28.6), (-13.2,-30.8), (-11,-30.8), (-8.8,-30.8), (-4.4,-30.8), (-2.2,-30.8), (0,-30.8), (2.2,-30.8), (4.4,-30.8), (8.8,-30.8), (11,-30.8), (13.2,-30.8), (-11,-33), (-8.8,-33), (-4.4,-33), (-2.2,-33), (0,-33), (2.2,-33), (4.4,-33), (8.8,-33), (11,-33), (-8.8,-35.2), (-6.6,-35.2), (-2.2,-35.2), (0,-35.2), (2.2,-35.2), (6.6,-35.2), (8.8,-35.2), (-6.6,-37.4), (-4.4,-37.4), (0,-37.4), (4.4,-37.4), (6.6,-37.4), (-4.4,-39.6), (4.4,-39.6)]
point_name_pairing = [("azurite", azurite_points)]
# Robot deck setup constants
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
# Place the PCR tubes in this order
well_colors = {
'A1': 'sfGFP',
'A2': 'mRFP1',
'A3': 'mKO2',
'A4': 'Venus',
'A5': 'mKate2_TF',
'A6': 'Azurite',
'A7': 'mCerulean3',
'A8': 'mClover3',
'A9': 'mJuniper',
'A10': 'mTurquoise2',
'A11': 'mBanana',
'A12': 'mPlum',
'B1': 'Electra2',
'B2': 'mWasabi',
'B3': 'mScarlet_I',
'B4': 'mPapaya',
'B5': 'eqFP578',
'B6': 'tdTomato',
'B7': 'DsRed',
'B8': 'mKate2',
'B9': 'EGFP',
'B10': 'mRuby2',
'B11': 'TagBFP',
'B12': 'mChartreuse_TF',
'C1': 'mLychee_TF',
'C2': 'mTagBFP2',
'C3': 'mEGFP',
'C4': 'mNeonGreen',
'C5': 'mAzamiGreen',
'C6': 'mWatermelon',
'C7': 'avGFP',
'C8': 'mCitrine',
'C9': 'mVenus',
'C10': 'mCherry',
'C11': 'mHoneydew',
'C12': 'TagRFP',
'D1': 'mTFP1',
'D2': 'Ultramarine',
'D3': 'ZsGreen1',
'D4': 'mMiCy',
'D5': 'mStayGold2',
'D6': 'PA_GFP'
}
volume_used = {
'azurite': 0
}
def update_volume_remaining(current_color, quantity_to_aspirate):
rows = string.ascii_uppercase
for well, color in list(well_colors.items()):
if color == current_color:
if (volume_used[current_color] + quantity_to_aspirate) > 250:
# Move to next well horizontally by advancing row letter, keeping column number
row = well[0]
col = well[1:]
# Find next row letter
next_row = rows[rows.index(row) + 1]
next_well = f"{next_row}{col}"
del well_colors[well]
well_colors[next_well] = current_color
volume_used[current_color] = quantity_to_aspirate
else:
volume_used[current_color] += quantity_to_aspirate
break
def run(protocol):
# Load labware, modules and pipettes
protocol.home()
# Tips
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
# Pipettes
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
# Deep Well Plate
temperature_plate = protocol.load_labware('nest_96_wellplate_2ml_deep', 6)
# Agar Plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate')
agar_plate.set_offset(x=0.00, y=0.00, z=Z_VALUE_AGAR)
# Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
# Helper function (dispensing)
def dispense_and_jog(pipette, volume, location):
assert(isinstance(volume, (int, float)))
# Go above the location
above_location = location.move(types.Point(z=location.point.z + 2))
pipette.move_to(above_location)
# Go downwards and dispense
pipette.dispense(volume, location)
# Go upwards to avoid smearing
pipette.move_to(above_location)
# Helper function (color location)
def location_of_color(color_string):
for well,color in well_colors.items():
if color.lower() == color_string.lower():
return temperature_plate[well]
raise ValueError(f"No well found with color {color_string}")
# Print pattern by iterating over lists
for i, (current_color, point_list) in enumerate(point_name_pairing):
# Skip the rest of the loop if the list is empty
if not point_list:
continue
# Get the tip for this run, set the bacteria color, and the aspirate bacteria of choice
pipette_20ul.pick_up_tip()
max_aspirate = int(18 // POINT_SIZE) * POINT_SIZE
quantity_to_aspirate = min(len(point_list)*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Iterate over the current points list and dispense them, refilling along the way
for i in range(len(point_list)):
x, y = point_list[i]
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_jog(pipette_20ul, POINT_SIZE, adjusted_location)
if pipette_20ul.current_volume == 0 and len(point_list[i+1:]) > 0:
quantity_to_aspirate = min(len(point_list[i:])*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Drop tip between each color
pipette_20ul.drop_tip()I have filled the form with links to both designs.
Post Lab Questions
- I chose a paper that was mentioned on the official Opentrons Resources. Paper Referenced

The paper describes that optimisation of drug delivery systems is a complex and multidimensional challenge that involves lots of factors, such asformulation composition, process parameters, and biological performance. The conventional approaches are limited, due to the high complexity, nonlinearity and multiple objective nature of the drug delivery problems of today's age. This article explores how Artificial Intelligence (AI) paired with Machine Learning (ML) can be used to improve formulation science by allowing data-driven, adaptive and efficient strategies. The paper lists both a conceptual and practical overview of ML-Guided optimisation workflows. The paper also discusses key challenges, such as data scarcity, experimental throughput, and model interpretability. Diverse Delivery applications are critically examined, highlighting how ML accelerates formulation development, reduce experimental burden, and uncover novel design spaces. The paper concludes, outlining the future of directions for integrating AI into pharmaceuticals and a focus on self-driving labs. The paper aims to aid drug delivery scientists with foundational knowledge and practical tools to harness AI and ML in the optimisation of advanced drug delivery systems.
- My main objective is to use lab automation in large production of my ideas. Once I have developed a concrete final idea, I do not possess access to a lab in my vicinity that would allow R&D to minors like myself. The goal is to be able to utilize the remote lab automation and create novel bioproducts. Additionally, a major reason I plan to use it is due to its precision. With my prior experiences in a wet lab (High School courses in University Labs), I have come to understand the impact of human error in the experiments. I wish to erase (or at least diminish) the variable of error with the help of lab automation. Additionally, these robots can perform multiple experiments that may contain components toxic to humans. This ensures utmost safety of the researchers, which is another priority.


