Week 4 HW: Protein Design Part I
Part A
1. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
- Meat contains ~20% protein (Kenneth Carpenter et al. Britannica, 2026).
- In 500 grams of meat, ~100 grams is protein. That is equivalent to 100 grams of amino acids.
- By the mol formula, the number of mols of amino acids = Mass/RMM = 100 grams/100 Daltons = 100 grams/100 gram mol^-1 = 1 mol
- Therefore, number of molecules = Number of mols x Avogadro’s Number = 1 mol x 6.02214076×1023 mol−1 = 6.02214076×10^23
- Hence, 6.02214076×10^23 molecules of amino acid are present in 500 grams of meat.
2. Why do humans eat beef but do not become a cow, eat fish but do not become fish?
- The meat is usually cooked before consumption. This would lead to the denaturing of DNA.
- Even bypassing that, the human digestive system causes the substances to be completely broken down to the constituent molecules (amino acids, water, etc.), with the help of multiple enzymes. The simpler molecules (nutrients) are then absorbed by the cells.
- The DNA or RNA from the cells of the animal being consumed does not get a chance to enter our human cells. We also have our immune system to prevent the entry of foreign DNAs into the cells.
- The human cells are also genetically coded to only produce more human cells. It cannot produce cells of other species (although viruses have the ability to alter this).
3. Why are there only 20 natural amino acids?
- It is common knowledge that amino acids having 3 nitrogeneous bases, a total of 64 combinations (4^3) are possible. It is believed that the choice of 20 amino acids is to allow redundancy, where multiple codons code for the same amino acids, increasing efficiency and reducing mutations. However, this does not answer why the specific set of 20 amino acids. Any of the other 41 combinations (leaving 3 combinations for stop codons) could have been used instead, right?
- The Frozen Accident Theory (Francis Crick, 1968) implies that these were the first 20 (or the most in amount) during the preliminary stages of biology, and later ended up being locked (frozen) as the default set, as changing the amino acids would not be energy-feasible, and these set of 20 seemed to be the most efficient.
- Proteins can be made with a much smaller set of amino acids. A Japanese group headed by Satoshi Akanuma at Waseda University recently showed that a 13 amino acid alphabet can create folded, soluble, stable and catalytically active ‘proteins’, albeit not as active or stable as the parent proteins on which they were based (R Shibue et al. Sci. Rep., 2018).
- It is therefore concluded, that after the initial 13-14 default amino acids, the rest were adaptive. They became “natural” amino acids, when oxygen started to become a part of the biochemical reactions on prehistoric Earth. That begs the question, “why not more?” Technically, there are 2 more amino acids, parttaking in protein synthesis, but not in genetic code. Hence, it can be hypothesized that the process had not stopped, it just reached a point where incorporating new amino acids was extremely hard.
The limitation is in the recognition of the tRNA. Each tRNA molecule has a well-defined tertiary structure that is recognized by the enzyme aminoacyl tRNA synthetase, which adds the correct amino acid. From studying tRNA structures, it was concluded that the problem is finding ways to make new tRNA molecules that could recognise a new amino acid without picking up existing ones (A Saint-Léger et al. Sci. Adv., 2016). This was the ultimate problem and answer to why there are only 20 amino acids.
- This answer uses the references, resources and content from the article - Why are there 20 Amino Acids?
4. Where did amino acids come from before enzymes that make them, and before life started?
- In 1953, Miller and Urey attempted to re-create the conditions of primordial Earth. In a flask, they combined ammonia, hydrogen, methane, and water vapor plus electrical sparks (Miller 1953). They found that new molecules were formed, and they identified these molecules as eleven standard amino acids. From this observation, they posited that the first organisms likely arose in an environment similar to the one they constructed in their flask, widely regarded as the primordial soup.
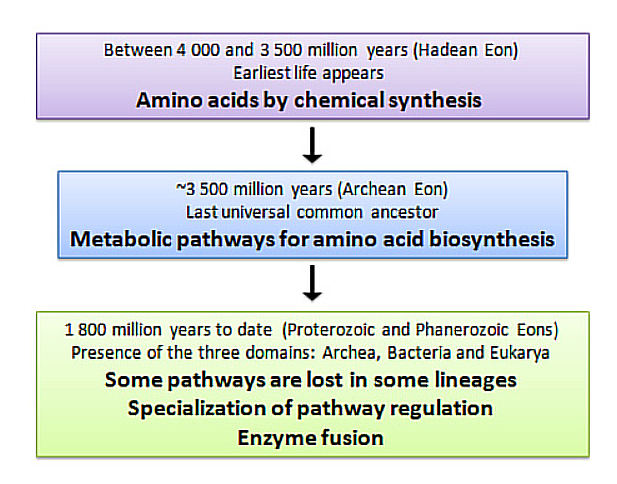
- This claim extended that in this soup, single-celled organisms evolved, while the necessary compounds depleted. In the competitive environment, only some organisms gained the ability to biosynthesize the required compounds (amino acids), while the rest died off, leading to the amino acids production by life we now know.
- The image from An Evolutionary Perspective on Amino Acids, illustrates this hypothesis:

5. If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
- An α-helix using D-amino acids would be left-handed, since D-amino acids are the mirrors of L-amino acids (Voet, Donald; Voet, Judith G. (2011). Biochemistry (4th ed.). Hoboken, NJ: John Wiley & Sons. p. 226.).
6. Can you discover additional helices in proteins?
- Additional Helices like pi-helix and 310 helix have been identified through research.
7. Why are most molecular helices right-handed?
- Despite the fact that both right-handed and left-handed α-helices are among the permitted conformations, the right-handed α-helix is energetically more favorable because of fewer steric clashes between the side chains and the main chain. Thus, all α-helices in proteins are right-handed (Robinson et al. Academic Press, 2014)
8. Why do β-sheets tend to aggregate? What is the driving force for β-sheet aggregation?
- β-sheets tend to aggregate due to 2 major reasons. The first, the hydrophobic effect (non-covalent interactions), which leads to face-to-face aggregation. The second, Van der Waals force (weak inter-molecular interactions), which lead to a sandwich-like, layered structure (Pham et al. Pub. Med., 2014).
9. Why do many amyloid diseases form β-sheets? Can you use amyloid β-sheets as materials?
- Amyloids consist of a β-sheet motif that repeats almost indefinitely. This and other unique properties bestow on these aggregates many biological activities like template assistance, membrane binding, and infectivity. This means that the β-sheet structure allows stability to the amyloid and helps it perform many vital activities, making it a staple among amyloid diseases (Riek, Pub. Med., 2017).
- The rigidity, chemical stability, high aspect ratio, and sequence programmability of amyloid fibrils have made them attractive candidates for functional materials with applications in environmental sciences, material engineering, and translational medicines (Zhang et al. Pub. Med., 2021).
Part B
1. Amino Acid Description
- For this assignment, I have chosen Cryptochrome. This is because I find it very interesting that a protein is responsible for circadian rhythms across different organisms. I have obtained the sequence and information about the CRY1 (Cryptochrome) in Homo sapiens from UniProt.
2. Amino Acid Identification
This is what I obtained from the Seqeunce section on the UniProt website:
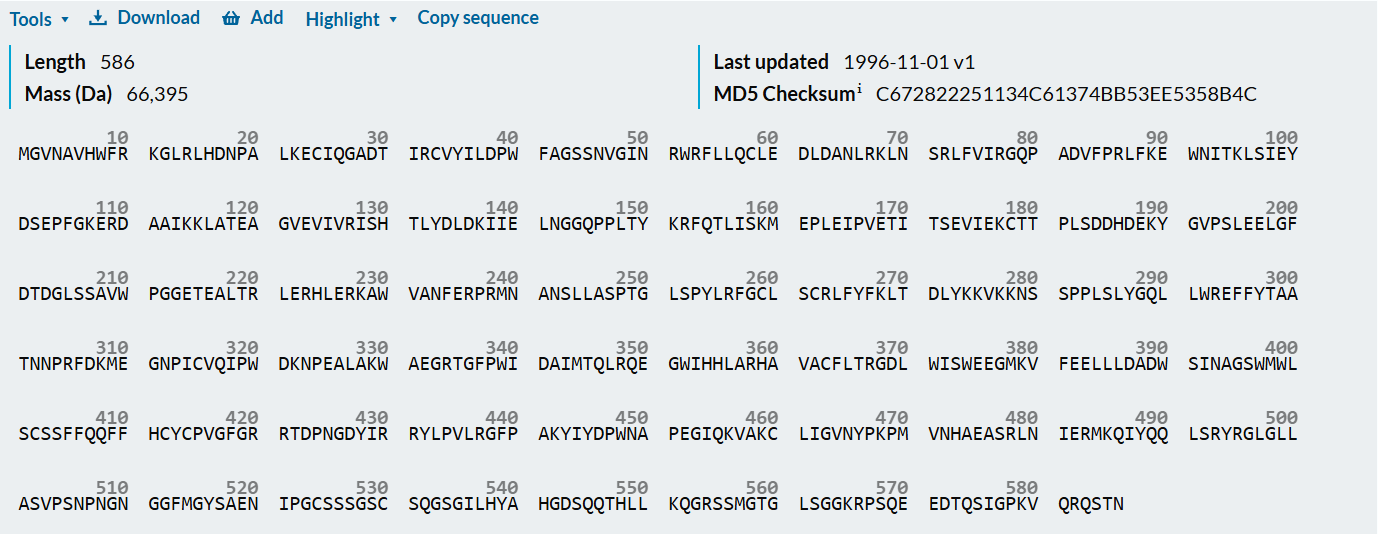

After using the Colab code, I obtained the following results:


- Length = 586 Amino Acids
- Most Frequent Amino Acid = L (60 times)
Number of Protein Sequence Homologs: BLAST found 967 results (I had increased the maximum limit to 1000).
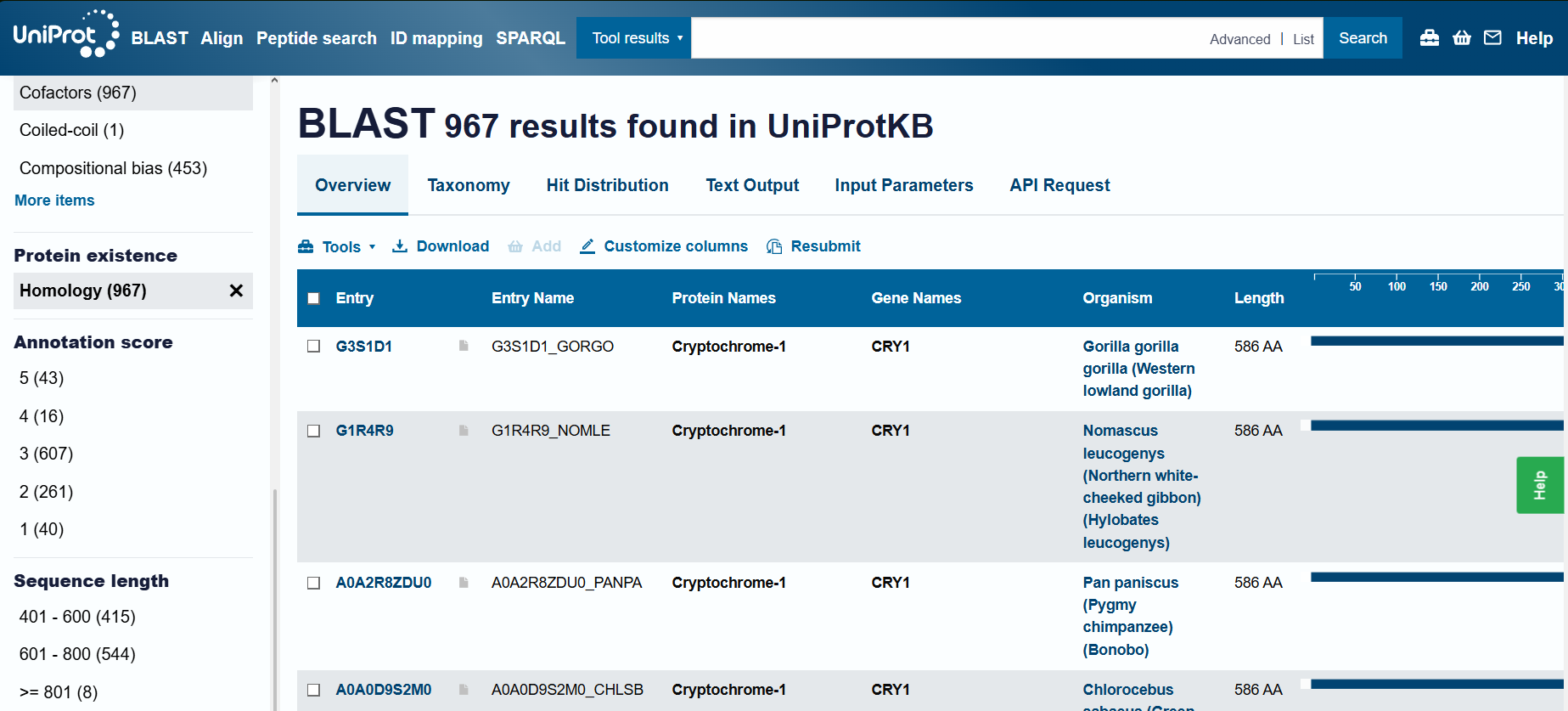

Cryptochrome belongs to the DNA photolyase class-1 family, according to UniProt
3. Protein Structure
- On RCSB, I found 19 results. However, only one was the Cryptochrome in Homo sapiens. The other results included Mus musculus (16) and 3 other organisms (1 each). I have listed 3 of these 19:
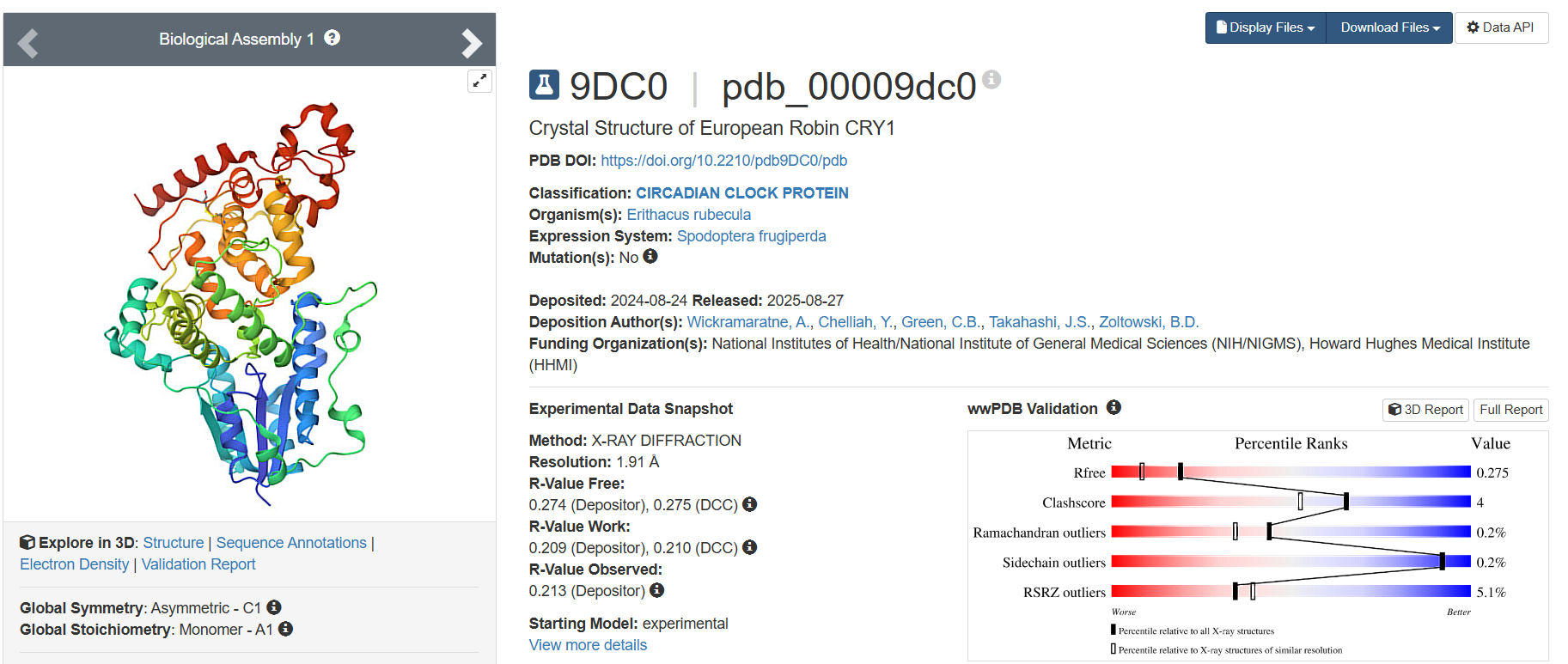
- The latest one, deposited on 2024-08-24, having 1.91 Å resolution, is one of the finer results.

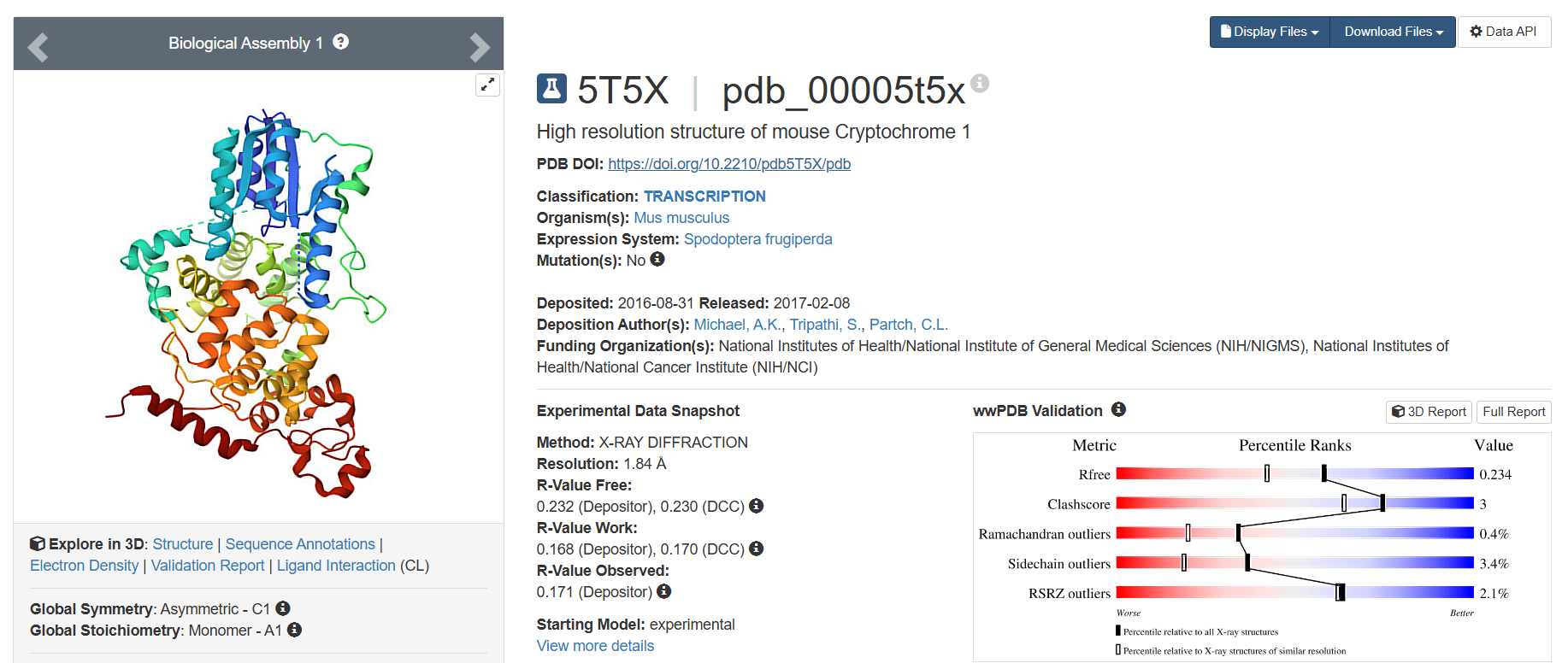
- The one with best resolution, deposited on 2017-02-08, having 1.84 Å resolution, has the lowest resolution and therefore, is the best quality structure.

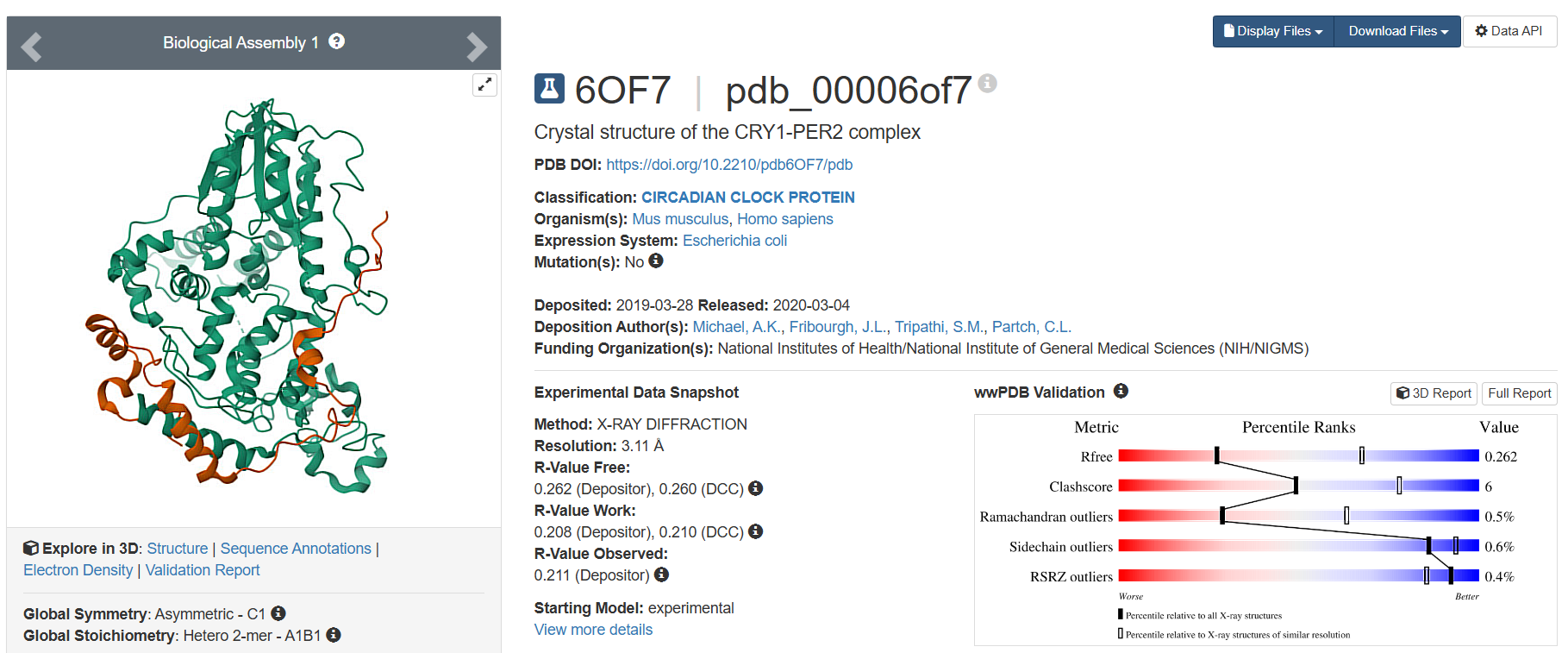
- The only one belonging to Homo sapiens, deposited on 2019-03-28, having 3.11 Å resolution, has the highest resolution and therefore, is not a very good quality structure.

- The latest one, deposited on 2024-08-24, having 1.91 Å resolution, is one of the finer results.
- According to the 3D Structures, only water molecules and polymer molecules are present.
- I was unable to find anything on SCOP related to the structural family of this protein.
4. 3D Molecule Visualization Software
- I chose to use UCSF Chimera.
- I downloaded and opened the latest deposited structure of Cryptochrome on RCSB.
- I tried visualization according to the tools and actions in Chimera.
- The visualization using both ‘Ribbon’ and ‘Ball and Stick.’
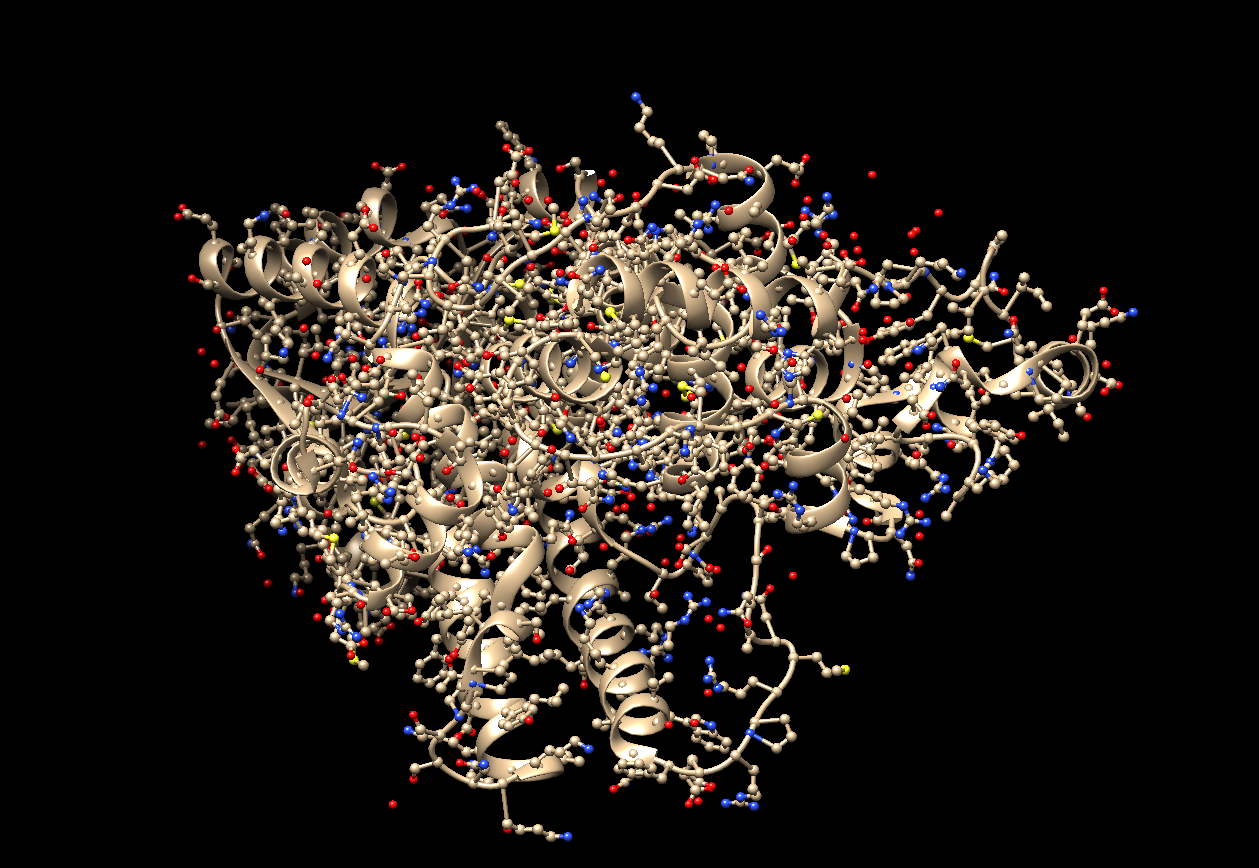

- The visualization using only ‘Ribbon.’
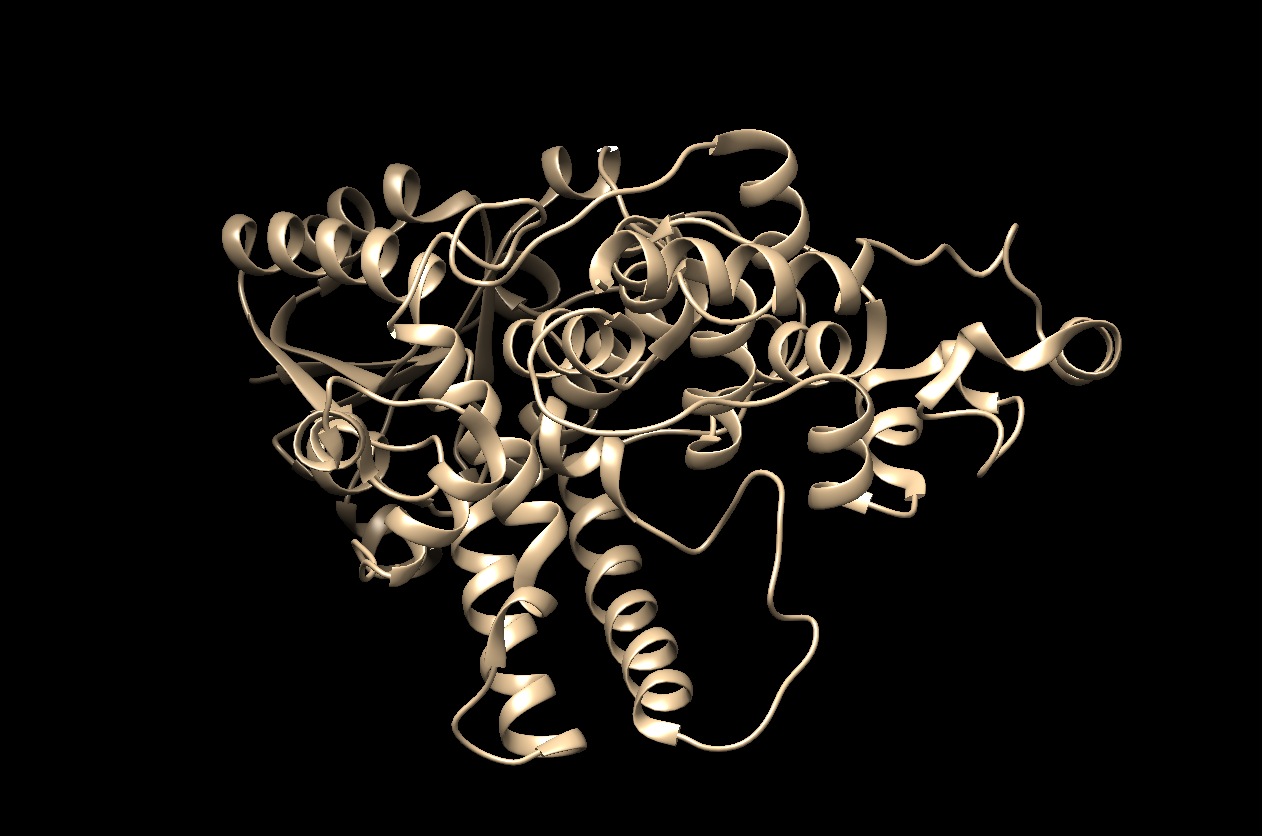

- The visualization using only ‘Ball and Stick.’
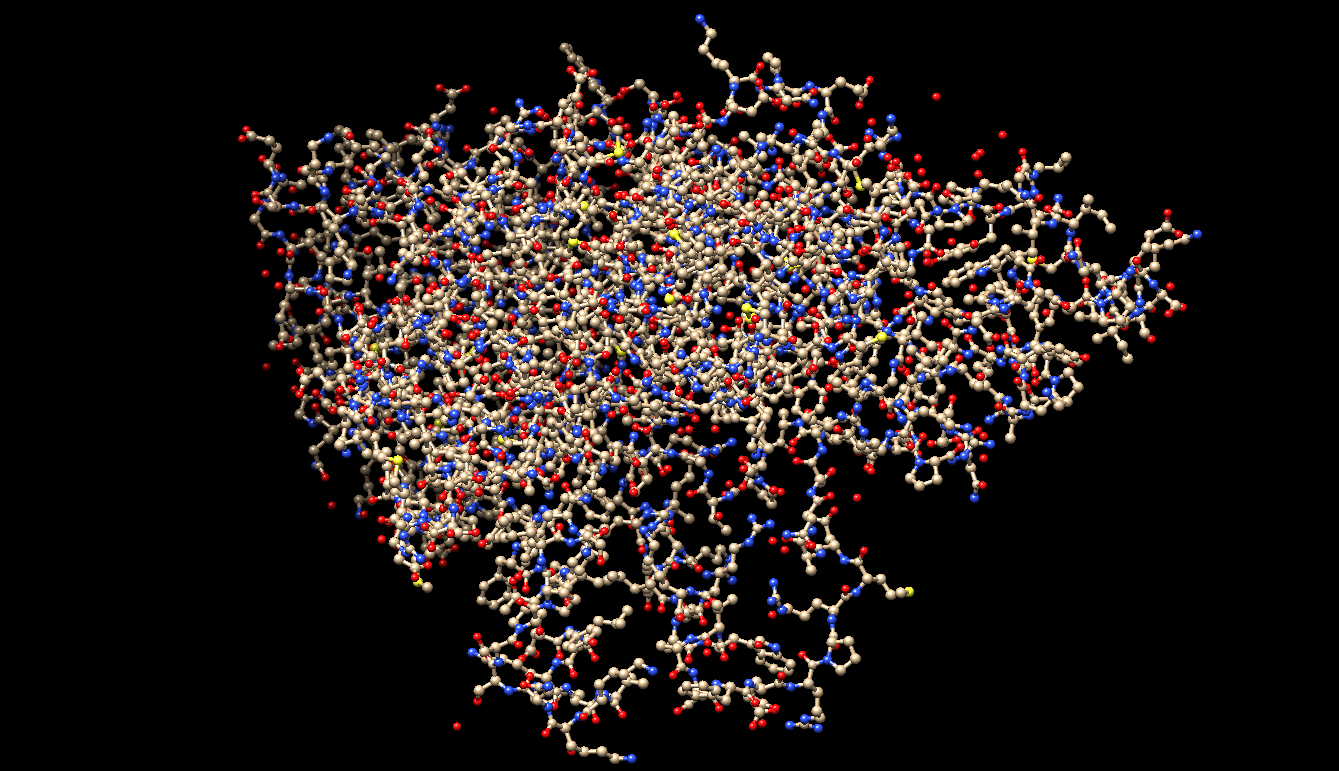

- The visualization by colored Secondary Structure. I chose Green for the Helix, Purple for the Coil, and Gray for the Strand. There are around 4 sheets, and many more numerous helices.
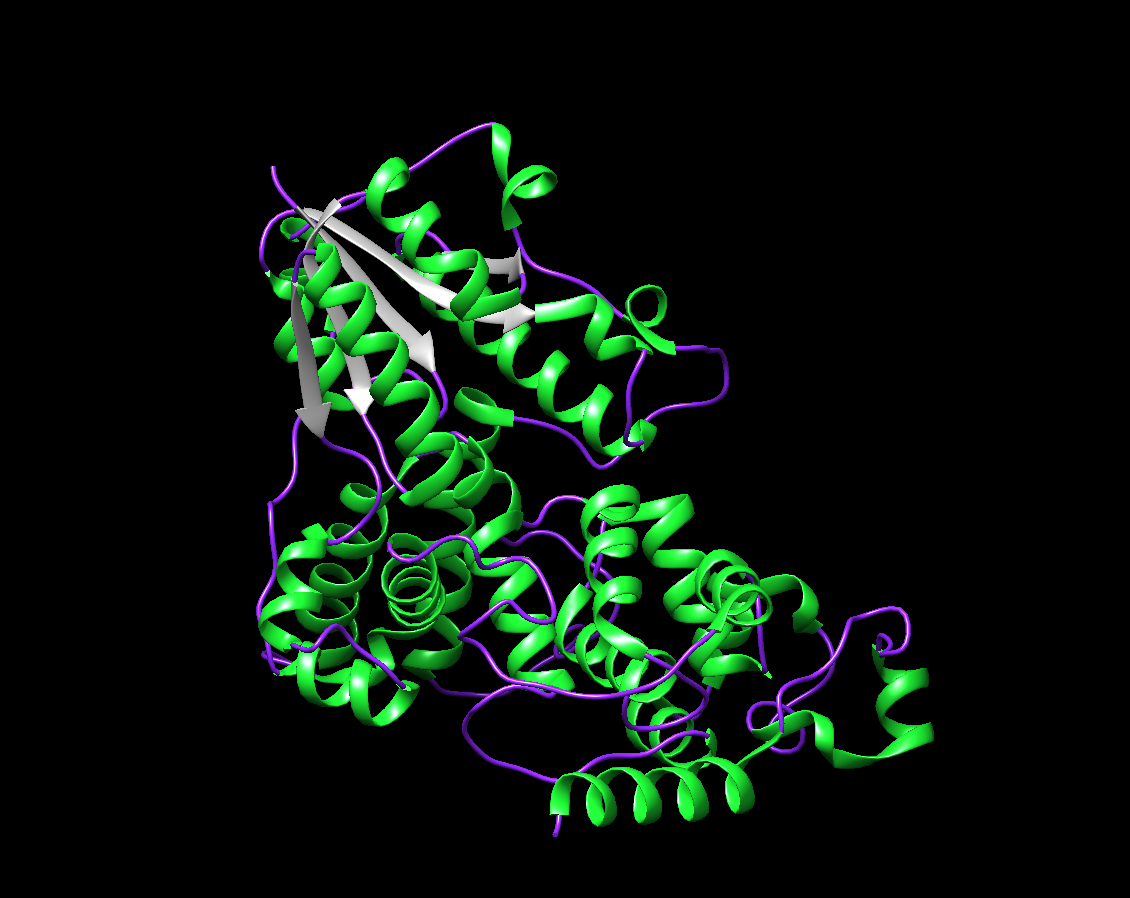

- The visualization by colored Residue Type. I chose Salmon for hydrophobic residues and Dark Cyan for the remaining (hydrophillic residues).
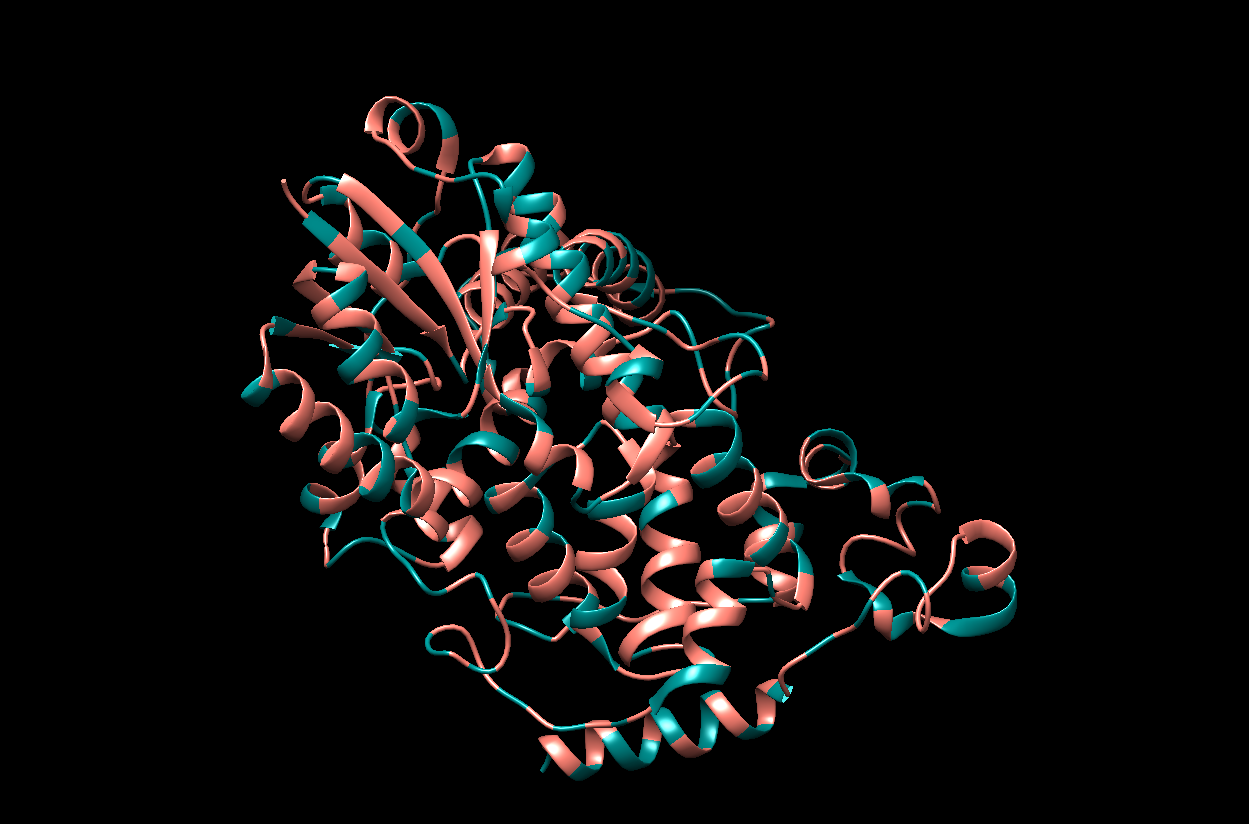

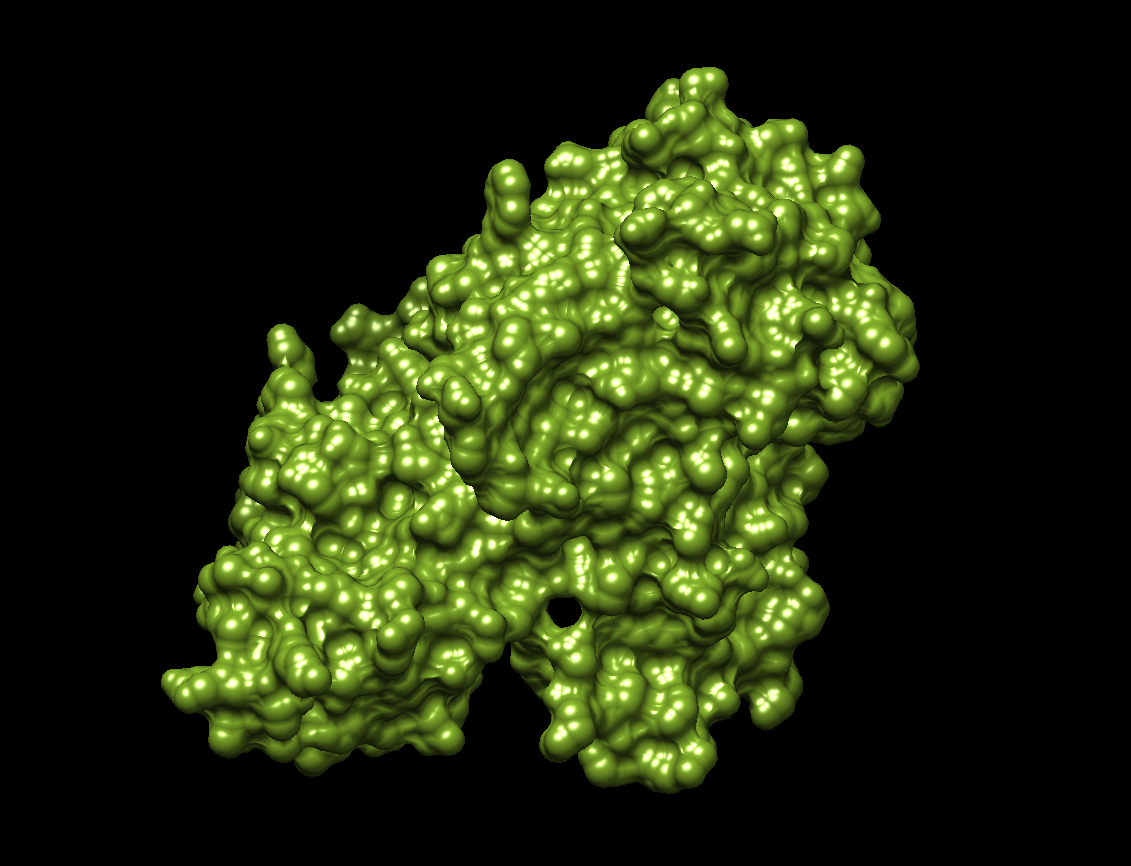
- The visualization of the surface. The protein surface seems to have multiple holes, grooves and depressions, meaning it should have multiple binding sites.