Week 3 HW : Lab Automation


Published Paper on Opentrons Automation
The paper entitled AssemblyTron: flexible automation of DNA assembly with Opentrons OT-2 lab robots, published in the journal Synthetic Biology (2023), reports the development of AssemblyTron, a software platform that automates DNA assembly workflows using the Opentrons OT-2 liquid-handling robot. The system integrates DNA construct design (for example, from design software such as j5) with automated execution on the OT-2 to perform molecular biology procedures in a precise and standardized manner. AssemblyTron is designed to support the Design–Build–Test–Learn (DBTL) cycle in synthetic biology by automating PCR optimization, Golden Gate assembly, and in vivo assembly (IVA).
In the study, the authors demonstrated the system’s capabilities by constructing four-fragment chromoprotein plasmids and performing site-directed mutagenesis. The results showed that the assembly fidelity achieved by the automated system was comparable to that of manual methods. In addition, the use of AssemblyTron reduced the likelihood of human error, minimized the need for extensive technical training, and decreased reagent waste, while improving experimental reproducibility and throughput. Overall, this platform enables researchers to accelerate the “build” phase of the DBTL cycle and allocate more time to the design and analysis stages in synthetic biology research.
Final Project Automation Plan
I plan to automate cell-free protein synthesis (CFPS) screening for engineered fluorescent biosensors detecting environmental analytes like heavy metals, using Opentrons OT-2 integrated with Ginkgo Nebula for remote design and deployment.
Core Workflow :
A. Design Phase: Use Ginkgo Nebula to design and order biosensor genetic constructs (e.g., GFP variants fused to metal-binding domains), targeting 96 variants with randomized promoter strengths.
B. Prep and Deposition: 3D-print custom PCR tube racks (inspired by Opentrons directory) to hold construct templates; Opentrons pipets DNA/cofactors into a 96-well plate.
C. Reaction Setup: Echo transfers templates; dispense CFPS master mix (lysate, T7 RNA pol, NTPs); Multiflo adds lysate to initiate expression; PlateLoc seals; Inheco incubates at 30°C for 16h.
D. Readout: XPeel removes seal; PHERAstar measures fluorescence/excitation spectra to score sensor performance.
Example Python Pseudocode (Opentrons API) :
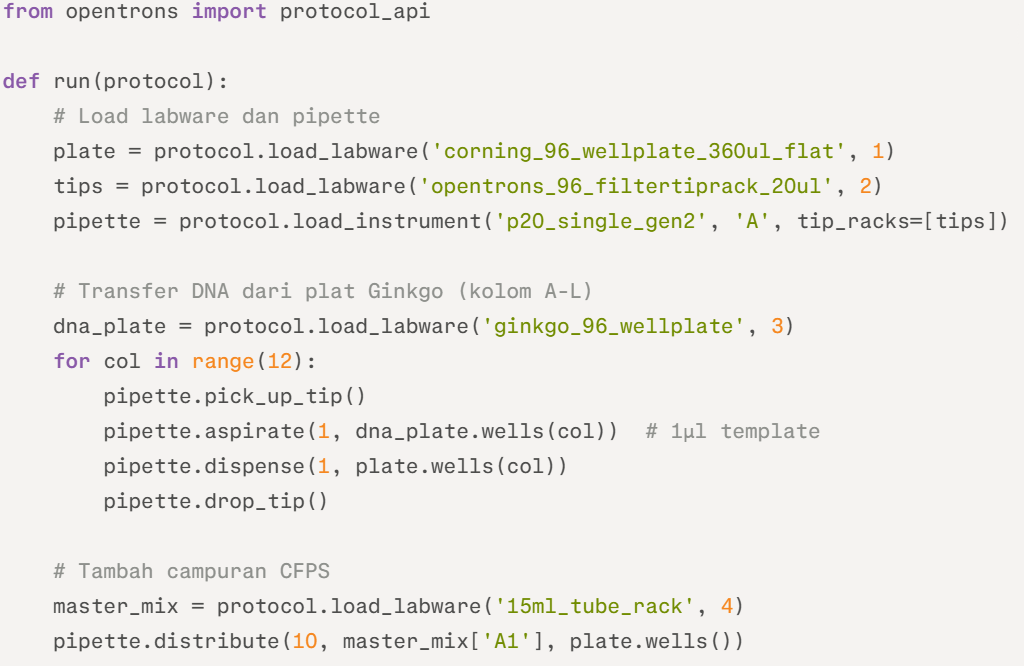

This script handles initial deposition; full protocol chains with temperature modules per recitation slides.
Hardware Additions : A. 3D-printed biosensor plate holder for precise alignment during imaging.
B. Remote monitoring via Opentrons App, synced to Ginkgo for iterative DBTL (e.g., top 10% sensors resynthesized).
This setup enables 100+ parallel reactions weekly, identifying hits for in vivo validation, all deployable remotely without intervention.