Week 4 HW: Protein Design Part I
A. Conceptual Question
- Molecules of Amino Acids in 500g Meat
A typical 500 g serving of lean meat contains about 125–150 g of protein (≈25–30% by weight). Taking 130 g as an average and assuming an average amino acid residue mass of ~110 g/mol, this corresponds to:
130 g ÷ 110 g/mol ≈ 1.18 mol of amino acid residues.
Multiplying by Avogadro’s number (6.022 × 10²³ mol⁻¹) gives:
≈ 7 × 10²³ amino acid molecules.
Therefore, a 500 g serving of lean meat provides on the order of 10²⁴ amino acid molecules
- Why Humans Eat Beef/Fish Without Becoming Them
Humans do not become cows or fish after eating them because dietary proteins are first degraded by proteolysis in the gastrointestinal tract into free amino acids and small peptides. The original protein sequence information is destroyed during digestion.These amino acids enter the bloodstream and are reused by ribosomes to synthesize human proteins according to mRNA transcribed from the human genome, following the central dogma (DNA → RNA → protein).Protein synthesis is directed exclusively by endogenous gene expression, and dietary nucleic acids are also degraded into nucleotides before absorption. Therefore, body structure and identity are determined by the host genome, not by consumed biomolecules.
- Why Only 20 Natural Amino Acids
The 20 canonical amino acids were likely selected through evolutionary processes because they provided sufficient chemical diversity while maintaining translational efficiency and genetic stability. Some, such as glycine and alanine, were probably abundant under prebiotic conditions.Together, the 20 amino acids span a broad range of chemical properties — including size, charge, hydrophobicity, and reactivity — enabling the formation of stable and functionally diverse protein structures.
They are encoded within a 64-codon triplet genetic system that provides redundancy and mutational robustness. Expanding the amino acid repertoire beyond 20 may not have offered sufficient selective advantage to offset the increased complexity of translation machinery. Additionally, once the genetic code became established, it likely became evolutionarily constrained (“frozen”), limiting further expansion. Rare additions such as selenocysteine require specialized recoding mechanisms and do not fundamentally alter the canonical set.
- Prebiotic Origin of Amino Acids
Before the emergence of life and enzymatic pathways, amino acids likely formed abiotically through several prebiotic chemical processes. Laboratory simulations such as the Miller–Urey experiment demonstrated that amino acids can be synthesized in reducing gas mixtures (e.g., CH₄, NH₃, H₂O, H₂) subjected to electrical discharges.Additional mechanisms include UV-driven photochemistry in atmospheric gases and geochemical reactions in hydrothermal environments, where mineral surfaces may have facilitated Strecker-type synthesis from aldehydes, hydrogen cyanide (HCN), and ammonia.
Extraterrestrial delivery also contributed to the prebiotic pool, as carbonaceous meteorites such as the Murchison meteorite contain more than 70 amino acids.Under certain environmental conditions, such as drying–wetting cycles or mineral-catalyzed reactions, these amino acids may have undergone condensation reactions to form short peptides, providing building blocks for early biochemical evolution.
- Handedness of D-Amino Acid α-Helix
An α-helix composed entirely of D-amino acids would adopt a left-handed helical conformation, which is the mirror image of the right-handed α-helix formed by L-amino acids.This occurs because chirality determines the energetically allowed φ and ψ backbone dihedral angles. In D-amino acids, the Ramachandran plot is effectively mirrored relative to L-amino acids, shifting the energy minimum to the opposite quadrant.As a result, the helical geometry is inverted while preserving the characteristic i→i+4 hydrogen bonding pattern of the α-helix. Experimental studies of synthetic D-peptides confirm the formation of stable left-handed α-helices.
- Additional Helices in Proteins
Beyond the canonical α-helix (3.6 residues per turn, i→i+4 hydrogen bonding), proteins can adopt additional helical conformations.The 3₁₀-helix contains approximately 3 residues per turn with i→i+3 hydrogen bonding and is often observed as short segments, frequently at α-helix termini. The π-helix has about 4.4 residues per turn with i→i+5 hydrogen bonding and typically appears as local insertions (π-bulges) within α-helices, sometimes contributing to functional or ligand-binding sites.
Additionally, the polyproline II (PPII) helix is an extended, left-handed helix lacking classical intrahelical hydrogen bonds. It is common in intrinsically disordered regions, and individual collagen chains adopt a PPII-like conformation before assembling into a triple helix. These examples demonstrate that protein secondary structure includes multiple helical geometries beyond the classical α-helix.
- Why Most Molecular Helices Right-Handed
Most biological helices are right-handed because proteins are composed almost exclusively of L-amino acids. The L-configuration at the α-carbon restricts the backbone φ and ψ dihedral angles such that the right-handed α-helix occupies the lowest-energy region of the Ramachandran plot (approximately φ ≈ −60°, ψ ≈ −45°). In contrast, a left-handed α-helix for L-amino acids falls into a higher-energy region due to unfavorable steric and torsional interactions. Thus, right-handed helices are energetically preferred. This molecular asymmetry arises from biological homochirality and contributes to the predominance of right-handed helices in proteins and many nucleic acid structures, although rare left-handed forms such as Z-DNA also exist.
- Why β-Sheets Aggregate
β-sheets aggregate as unsatisfied H-bond donors/acceptors at strand edges pair intermolecularly with adjacent sheets, propagating fibrils; flat sheet geometry exposes hydrophobic faces, driving lateral association in water. This is entropically favored by releasing structured water.
Driving Force
Primary force is hydrophobic burial of nonpolar side chains (e.g., valine, leucine) between sheets, supplemented by van der Waals packing and H-bonds; electrostatics (salt bridges) fine-tune. Thermodynamic stability (low free energy) from these non-covalent interactions exceeds soluble states.
- Why Amyloid Diseases Form β-Sheets
Many amyloid diseases arise because certain proteins can misfold into β-sheet–rich conformations under conditions that destabilize their native structure or impair proteostasis. For example, amyloid-β in Alzheimer’s disease and misfolded prion protein undergo nucleation-dependent polymerization, in which exposed aggregation-prone segments template further misfolding. The resulting cross-β architecture—where β-strands run perpendicular to the fibril axis and sheets stack along it—forms extensive intermolecular hydrogen-bonding networks. This structure confers high thermodynamic stability, protease resistance, and kinetic persistence.
As materials, amyloid fibrils self-assemble into highly ordered nanofibers with mechanical stiffness in the gigapascal range. Their nanoscale organization and robustness have inspired applications in biomaterials, hydrogels, tissue scaffolds, and bioelectronic nanowires. Engineered peptide variants can be designed to retain self-assembly properties while minimizing cytotoxicity, enabling safe material applications.
Part B: Protein Analysis and Visualization
The selected protein is the human Hemoglobin subunit beta, which plays a crucial role in transporting oxygen through the bloodstream as a key component of hemoglobin. This protein was chosen for its vital importance in human physiology, its availability of a high-quality 3D structure, and its frequent use as a model in protein structure studies.
Amino Acid Sequence
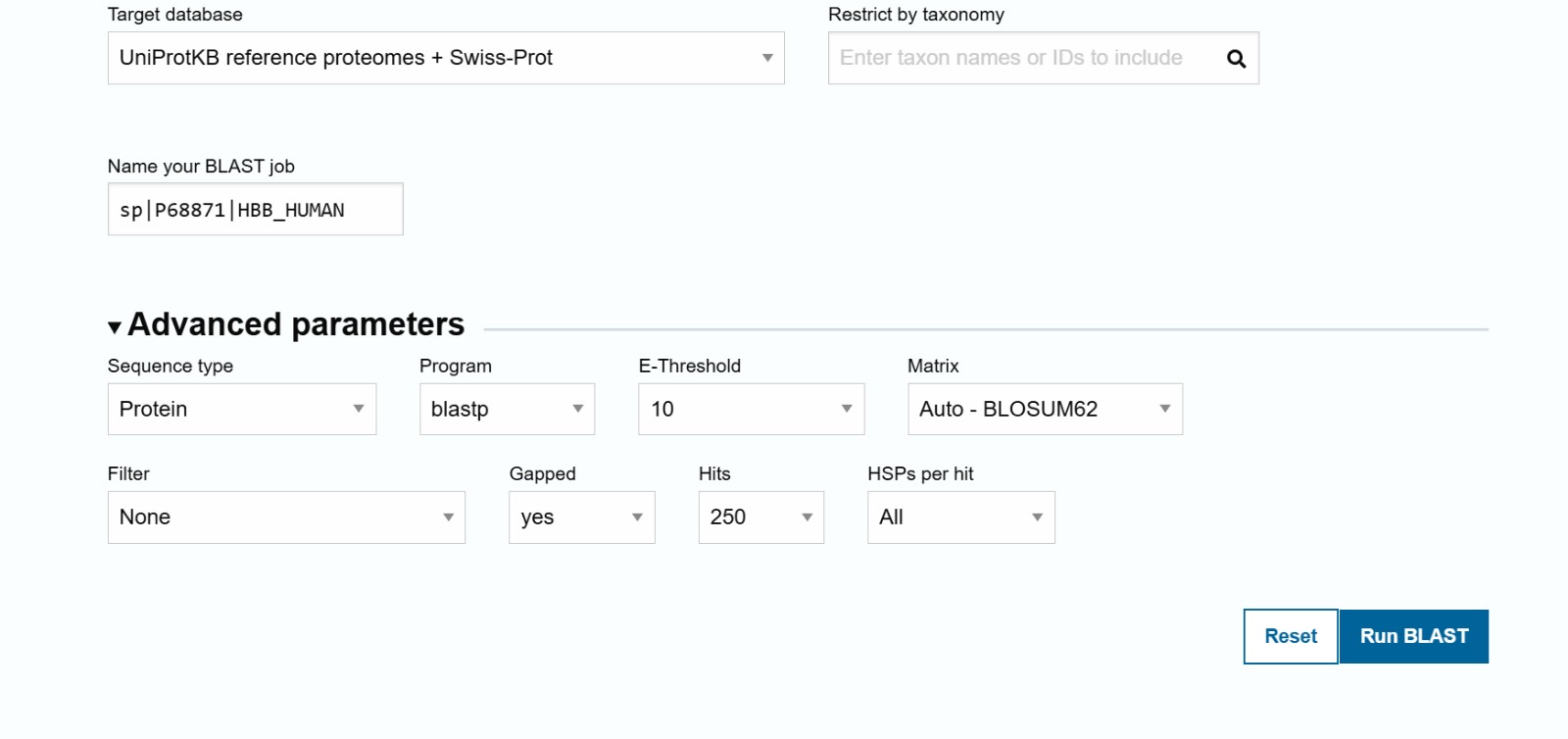
The sequence was obtained from UniProt entry P68871
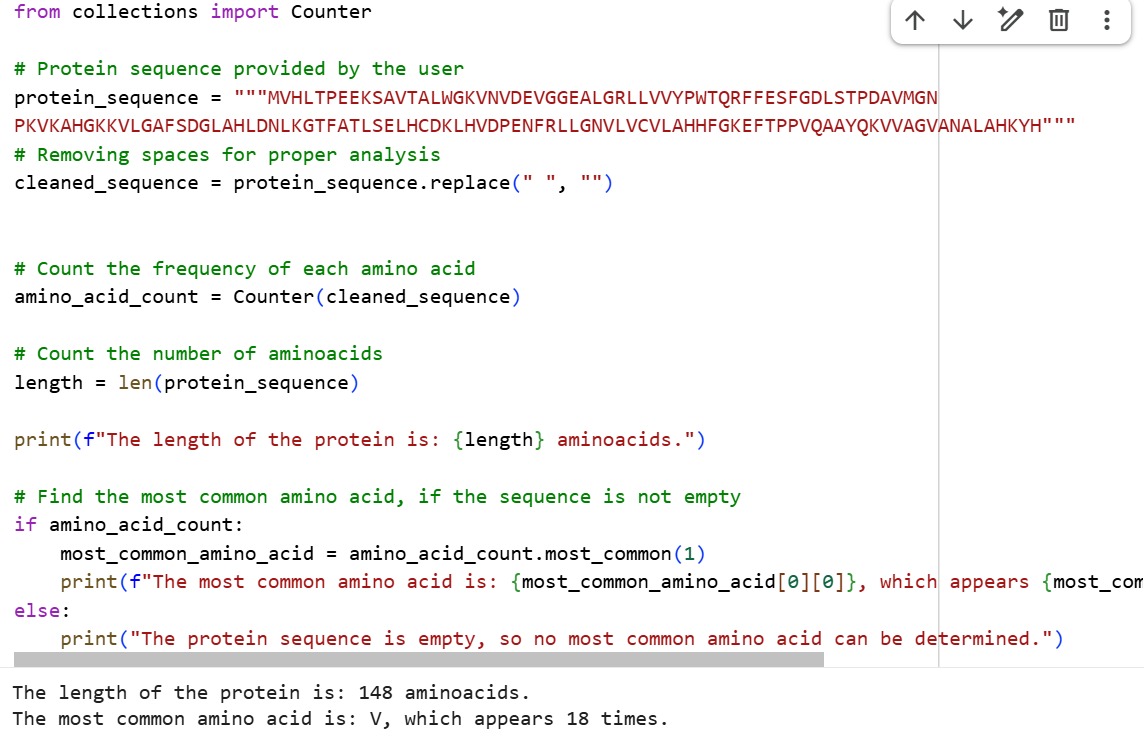
MVHLTPEEKSAVTALWGKVNVDEVGGEALGRLLVVYPWTQRFFESFGDLSTPDAVMGNPKVKAHGKKVLGAFSDGLAHL DNLKGTFATLSELHCDKLHVDPENFRLLGNVLVCVLAHHFGKEFTPPVQAAYQKVVAGVANALAHKYH
The length of the protein is: 148 aminoacids. The most common amino acid is: V, which appears 18 times.


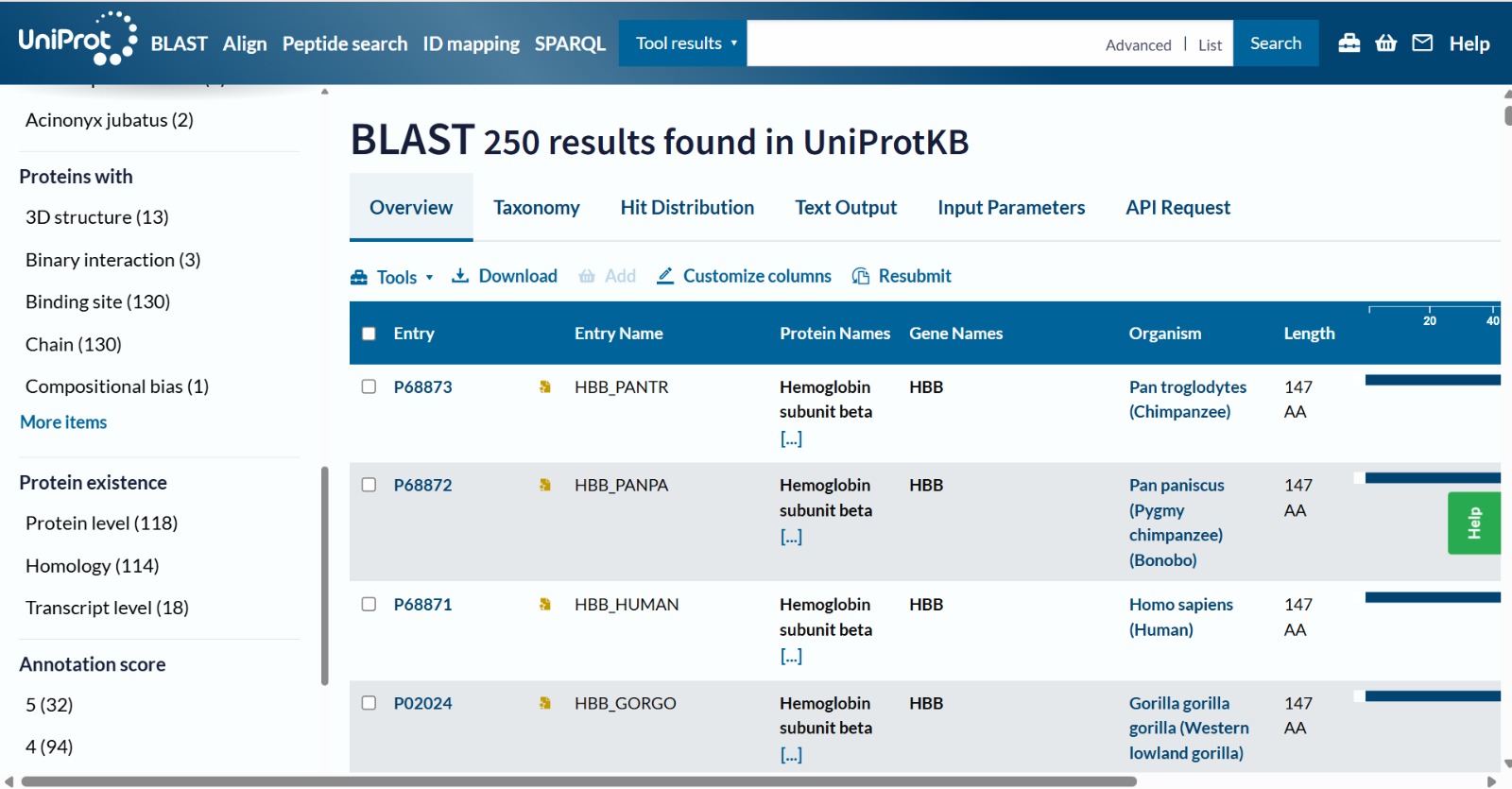

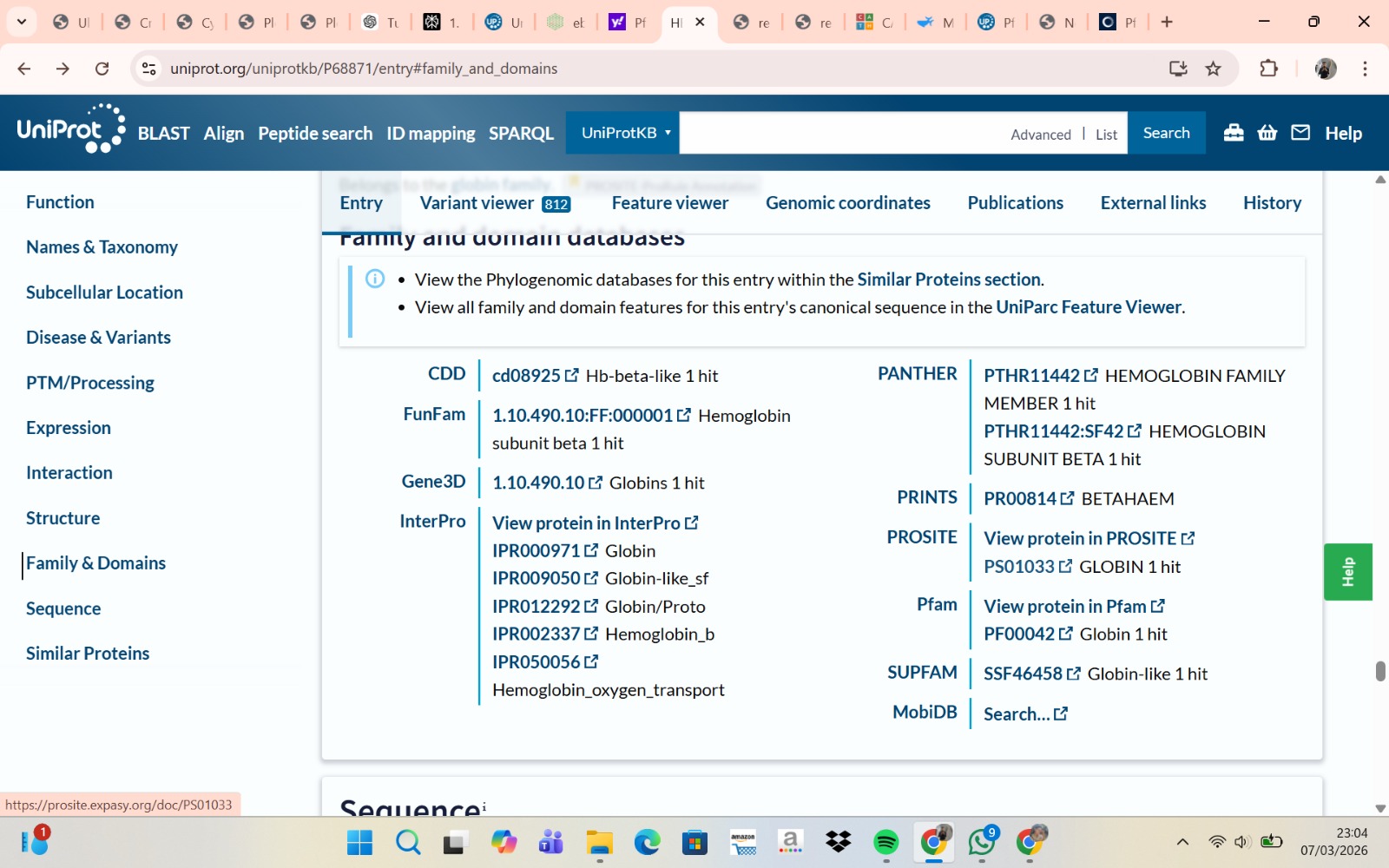
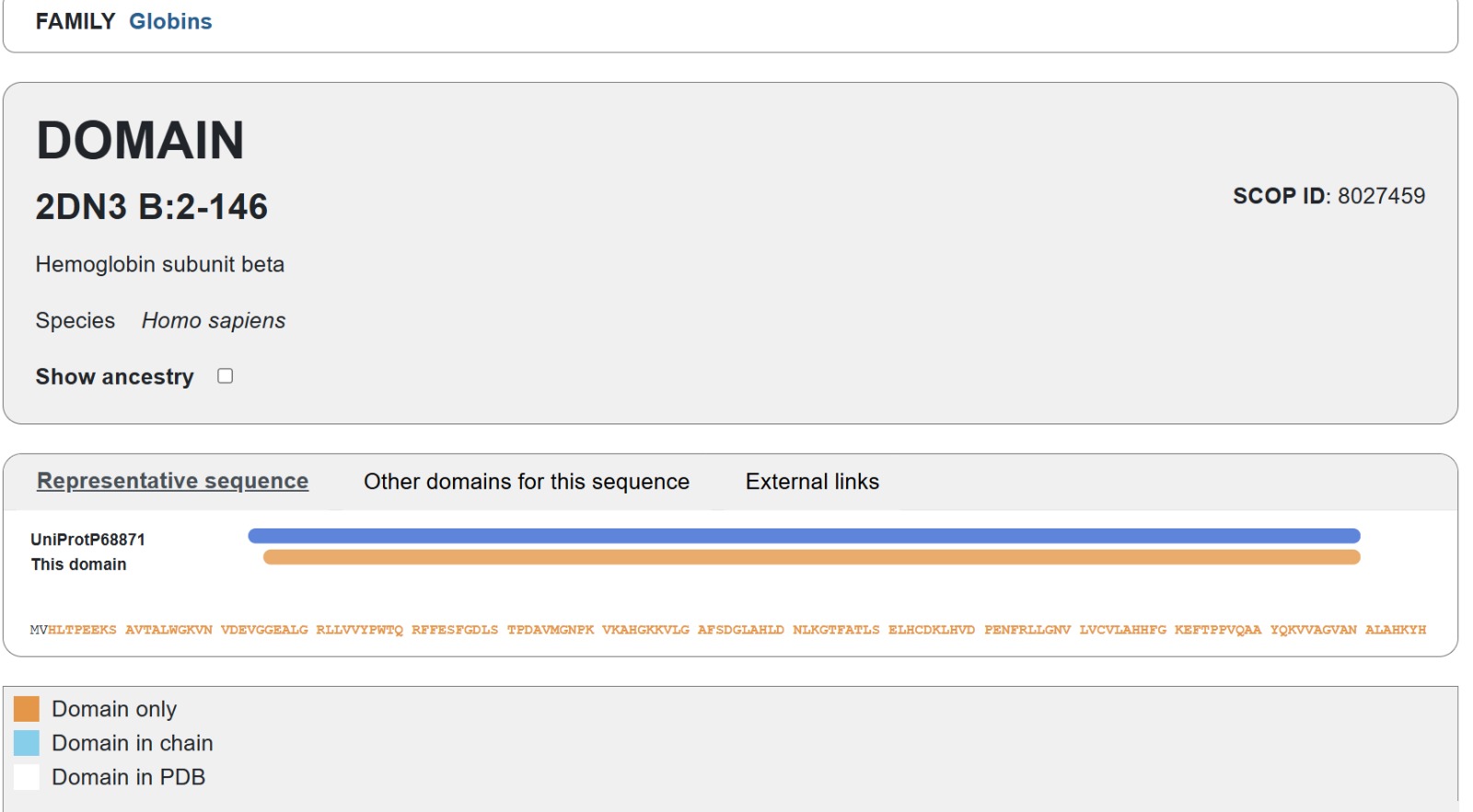
The human Hemoglobin subunit beta belongs to the hemoglobin protein family within the globin superfamily. Domain analysis from the UniProt protein database indicates that the protein contains a conserved globin domain spanning residues 1–147. Multiple protein classification databases consistently identify this protein as a member of the globin family, including Pfam (PF00042), InterPro (IPR000971), and Gene3D (Globins). InterPro further classifies it within the globin-like superfamily (IPR009050) and the hemoglobin beta subfamily (IPR002337). Additional databases such as PANTHER (PTHR11442) and SUPFAM (SSF46458) also support its classification within the globin-like structural superfamily. These annotations indicate that the protein adopts the characteristic globin fold, which enables oxygen binding through a heme prosthetic group.

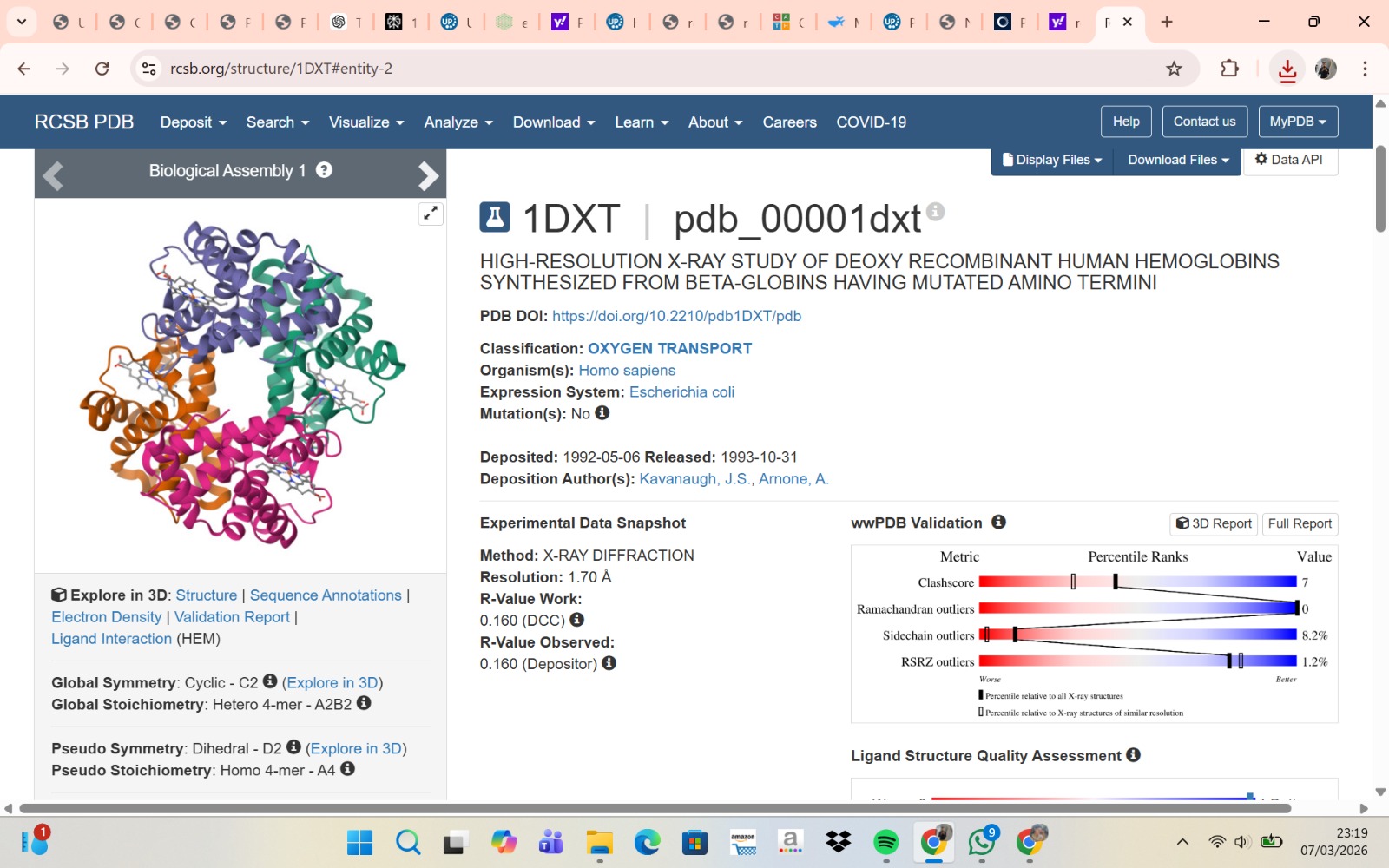
- The structure of Hemoglobin subunit beta from humans is available in the RCSB Protein Data Bank under the PDB ID 1DXT. The structure was determined using X-ray diffraction and deposited in 1992 and released in 1993. It has a resolution of 1.70 Å, indicating a high-quality structure with detailed atomic information. The biological assembly forms a hemoglobin tetramer (A2B2), consisting of two alpha and two beta subunits. In addition to the protein chains, the structure also contains a heme prosthetic group (HEM) that serves as the oxygen-binding site. Structurally, the protein belongs to the globin-like structural superfamily, characterized by a predominantly alpha-helical globin fold and the presence of a heme-binding pocket responsible for oxygen transport.

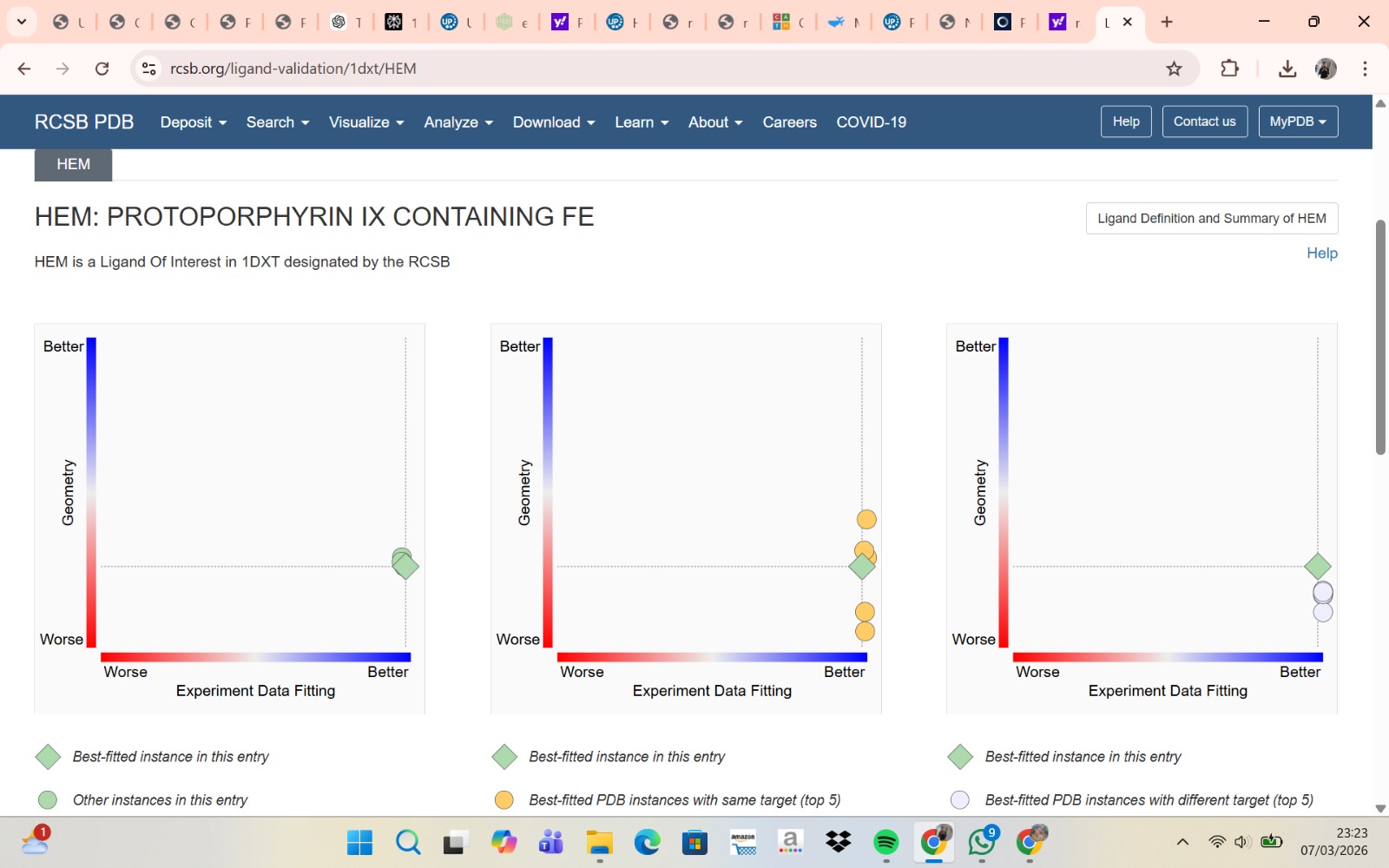

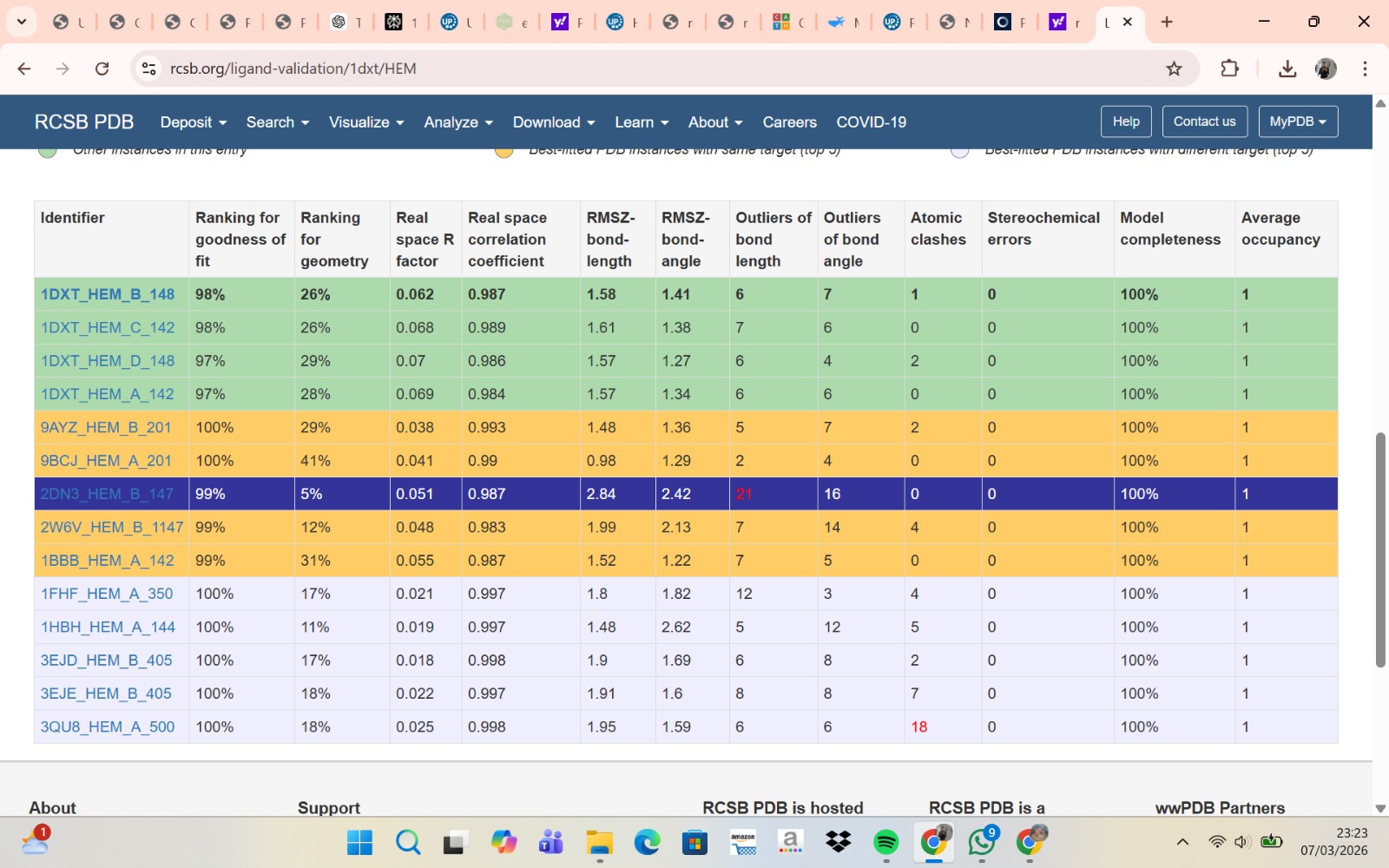


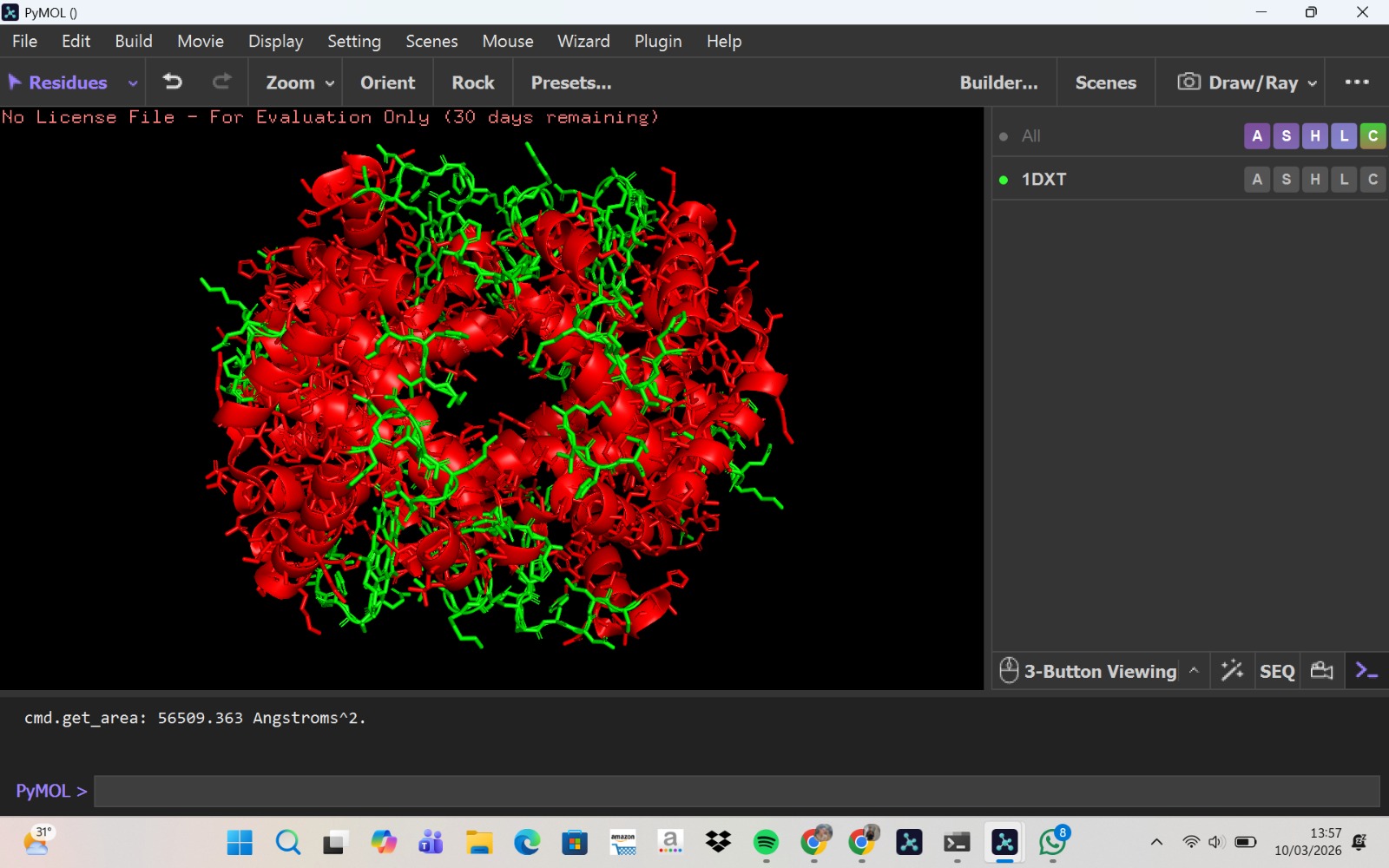
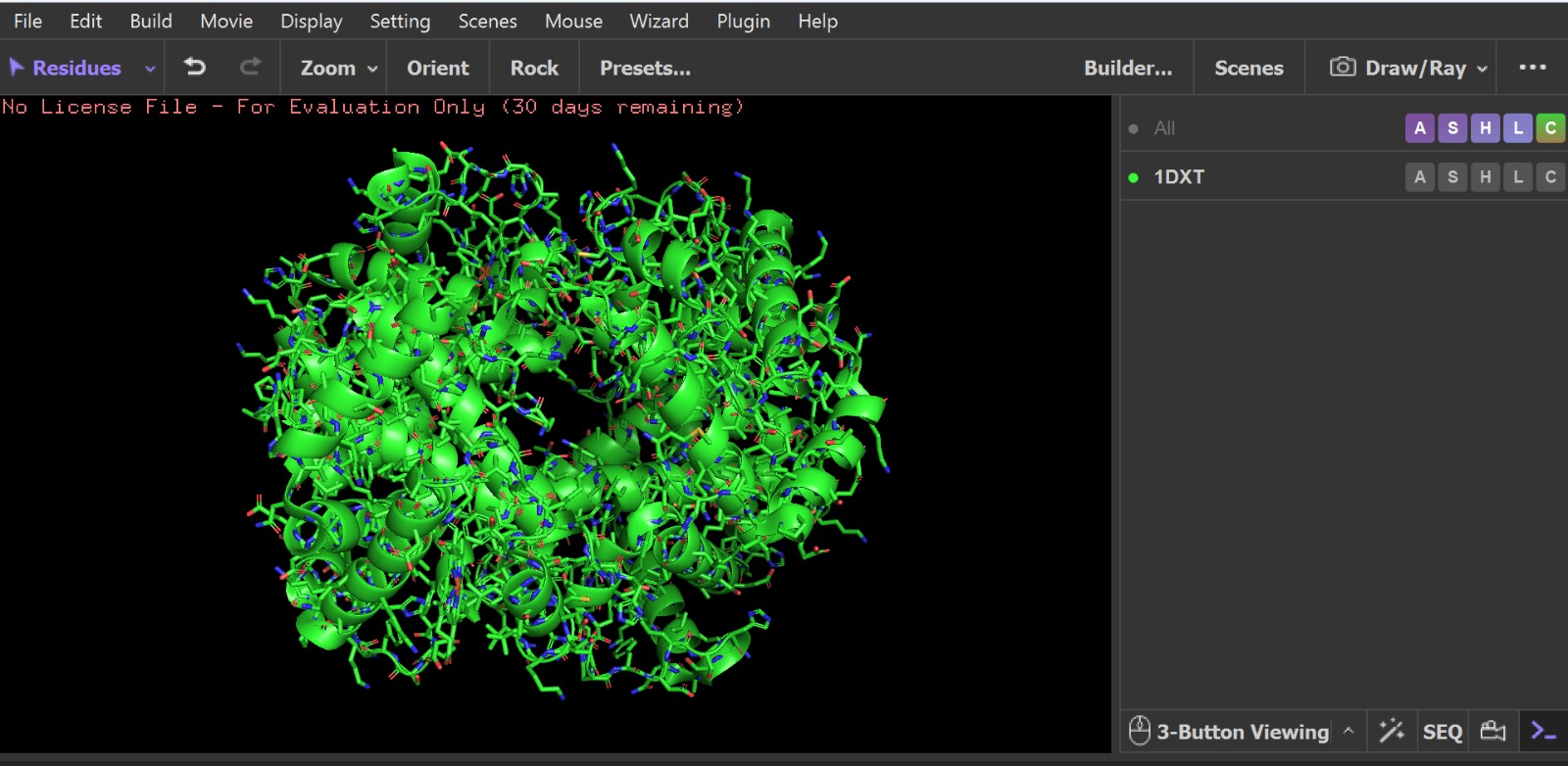
- PyMol Visualization
Visualization of the human Hemoglobin subunit beta protein using PyMOL software with stick representation and water removal.

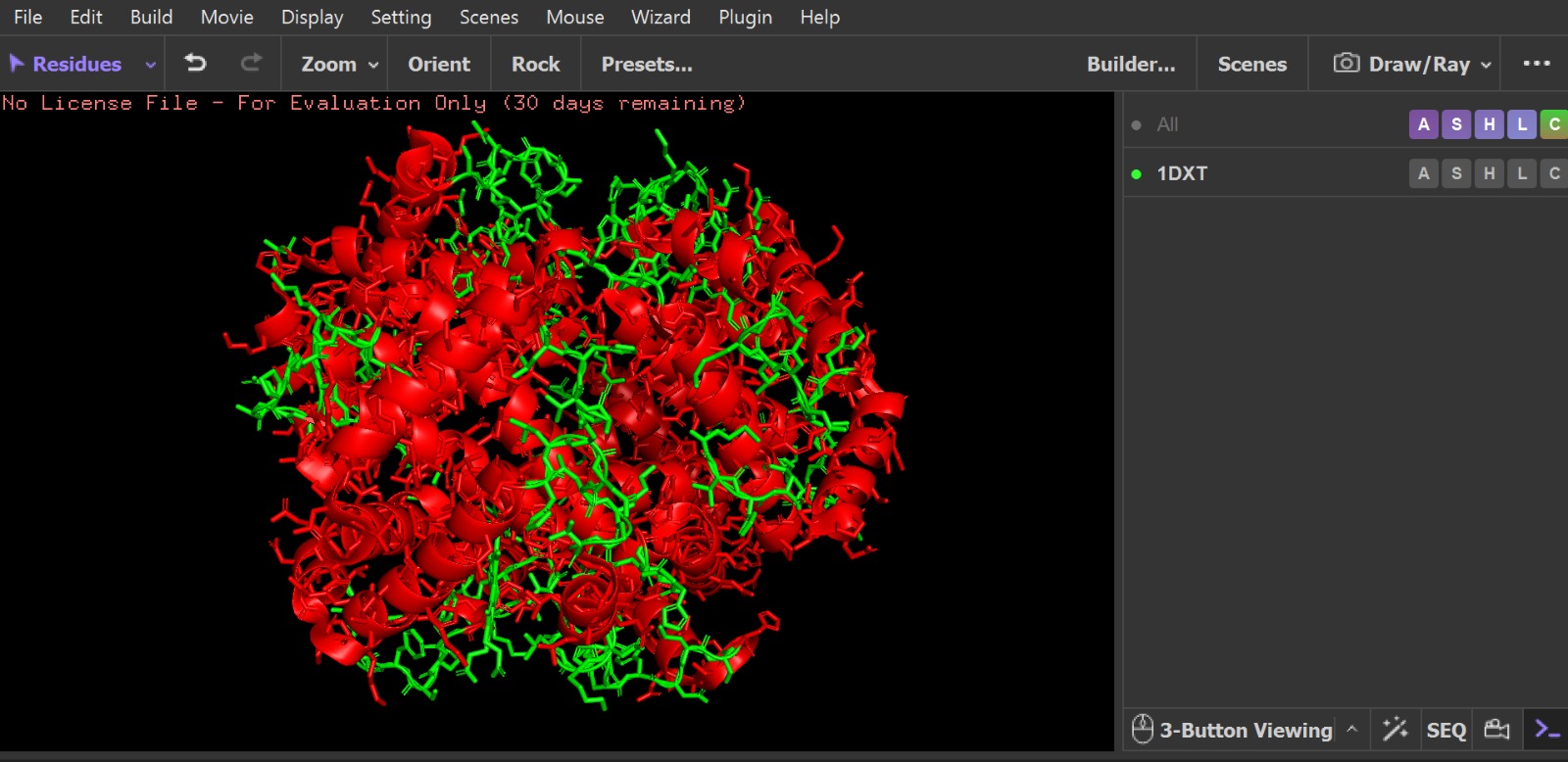
Visualization of the protein structure using PyMOL molecular visualization system colored by secondary structure shows that the structure is dominated by α-helices (red), while only a few regions correspond to loops or coils (green). No significant β-sheet structures are observed. Therefore, the protein contains many more α-helices than β-sheets. This observation is consistent with the typical globin fold of Hemoglobin subunit beta, where each subunit is primarily composed of multiple α-helices that form the structural framework around the heme-binding pocket.

Visualization of the molecular structure of Human hemoglobin using PyMOL molecular visualization system reveals the presence of a distinct binding pocket within the protein. This pocket corresponds to the heme-binding site, where the heme prosthetic group containing an Fe²⁺ ion is located. The pocket is formed by several surrounding α-helices of the globin fold. This structural cavity allows oxygen molecules to bind to the Fe²⁺ ion in the heme group, enabling hemoglobin to perform its biological function in oxygen transport. Therefore, the protein clearly contains functional binding pockets on its surface.