Week 5 HW: Protein Design Part II
PART 1 Generate Binders with PepMLM
The human SOD1 protein sequence was retrieved from the UniProt database (P00441). To model a disease-associated variant, the A4V mutation was introduced by substituting alanine with valine at residue position 4 of the protein sequence. This mutation is known to be associated with amyotrophic lateral sclerosis (ALS). The resulting mutant SOD1 sequence was then used as the input for subsequent peptide binder generation using the PepMLM model.

In bioinformatics, the letter “X” in peptide sequences denotes an unknown or ambiguous amino acid, often arising from prediction model uncertainties during protein structure analysis. For instance, sequences like WRYYAYAIRWKX, HLVPAVAIEHKX, and WRYPAAAARLKX feature an unclear final residue, which disrupts processing in tools like AlphaFold3 as they do not support non-standard characters. To address this, X is typically replaced with the most contextually reasonable residue, such as K (lysine). This substitution is favored because lysine enhances electrostatic binding—via its positive charge that attracts negatively charged molecules—and improves overall peptide solubility, ensuring greater stability and compatibility for downstream simulations.
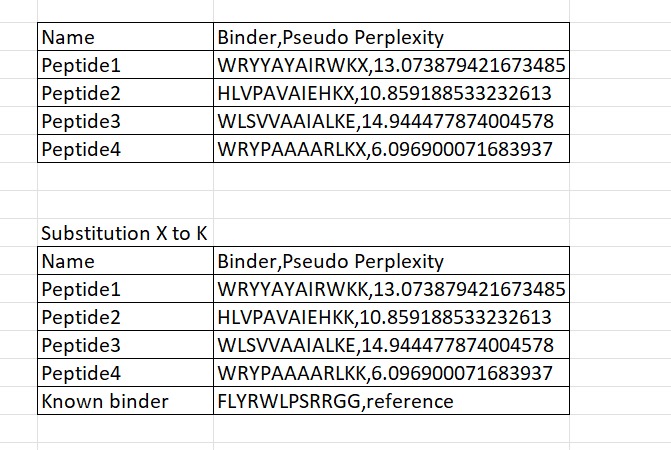

Four peptides of length 12 amino acids were generated using the PepMLM model conditioned on the mutant SOD1 A4V sequence. The generated peptides showed pseudo-perplexity scores ranging from 6.10 to 14.94, reflecting varying levels of model confidence. Among the generated candidates, peptide WRYPAAAARLKK displayed the lowest perplexity score (6.10), suggesting the highest likelihood of being a plausible binder according to the model. For comparison, the previously reported SOD1-binding peptide FLYRWLPSRRGG was included as a reference. These peptides were then used for further structural evaluation using AlphaFold3.
Part 2 Evaluate Binders with AlphaFold3
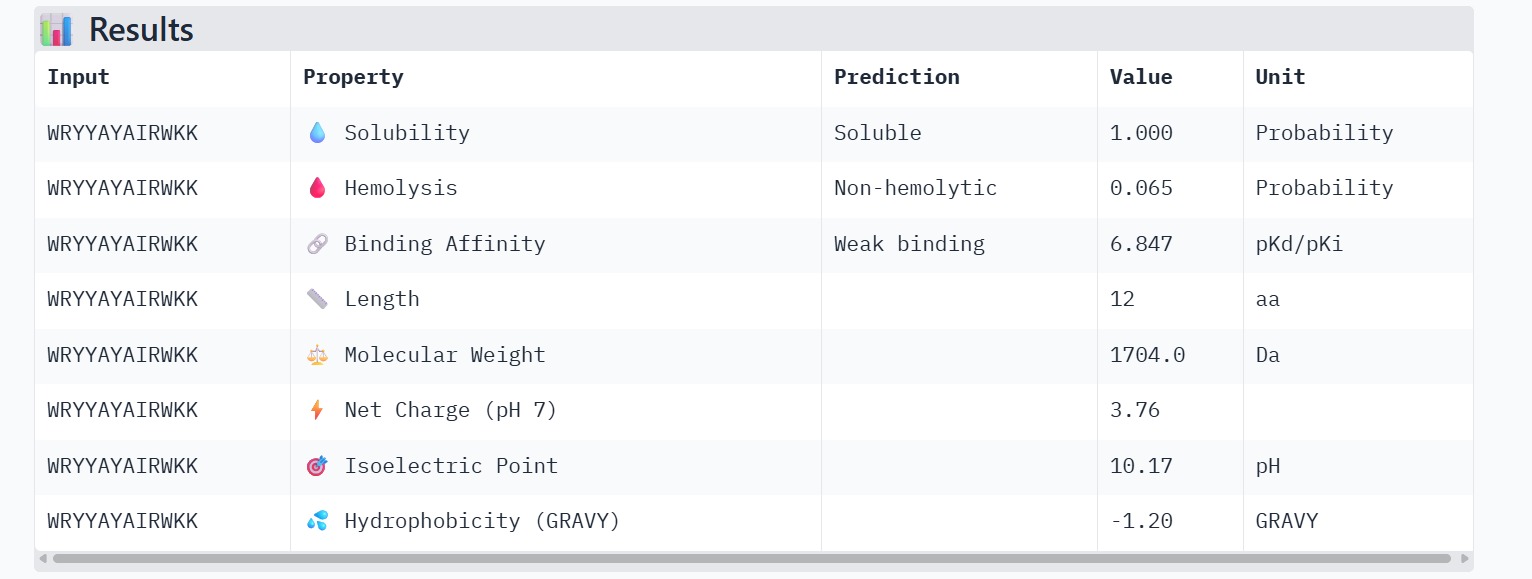
Peptide 1 : WRYYAYAIRWKK
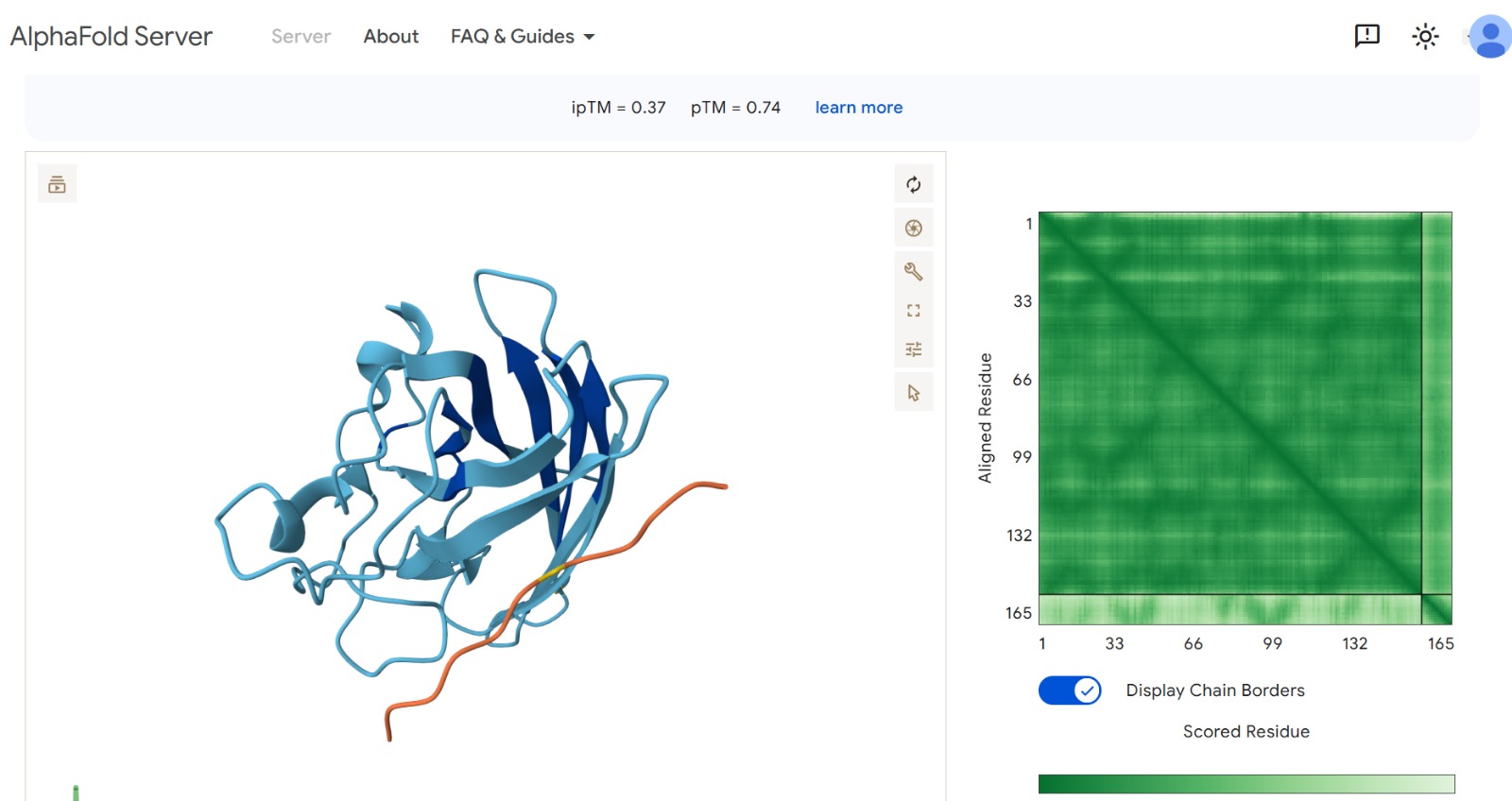

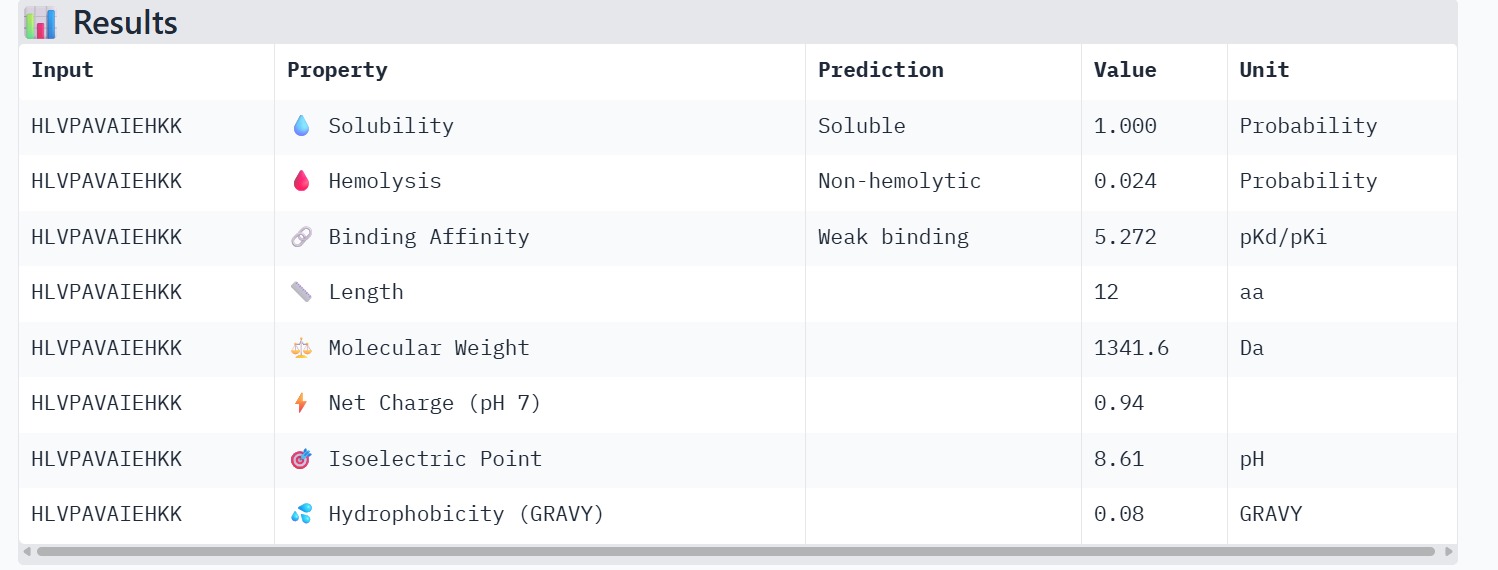
Peptide 2 : HLVPAVAIEHKK
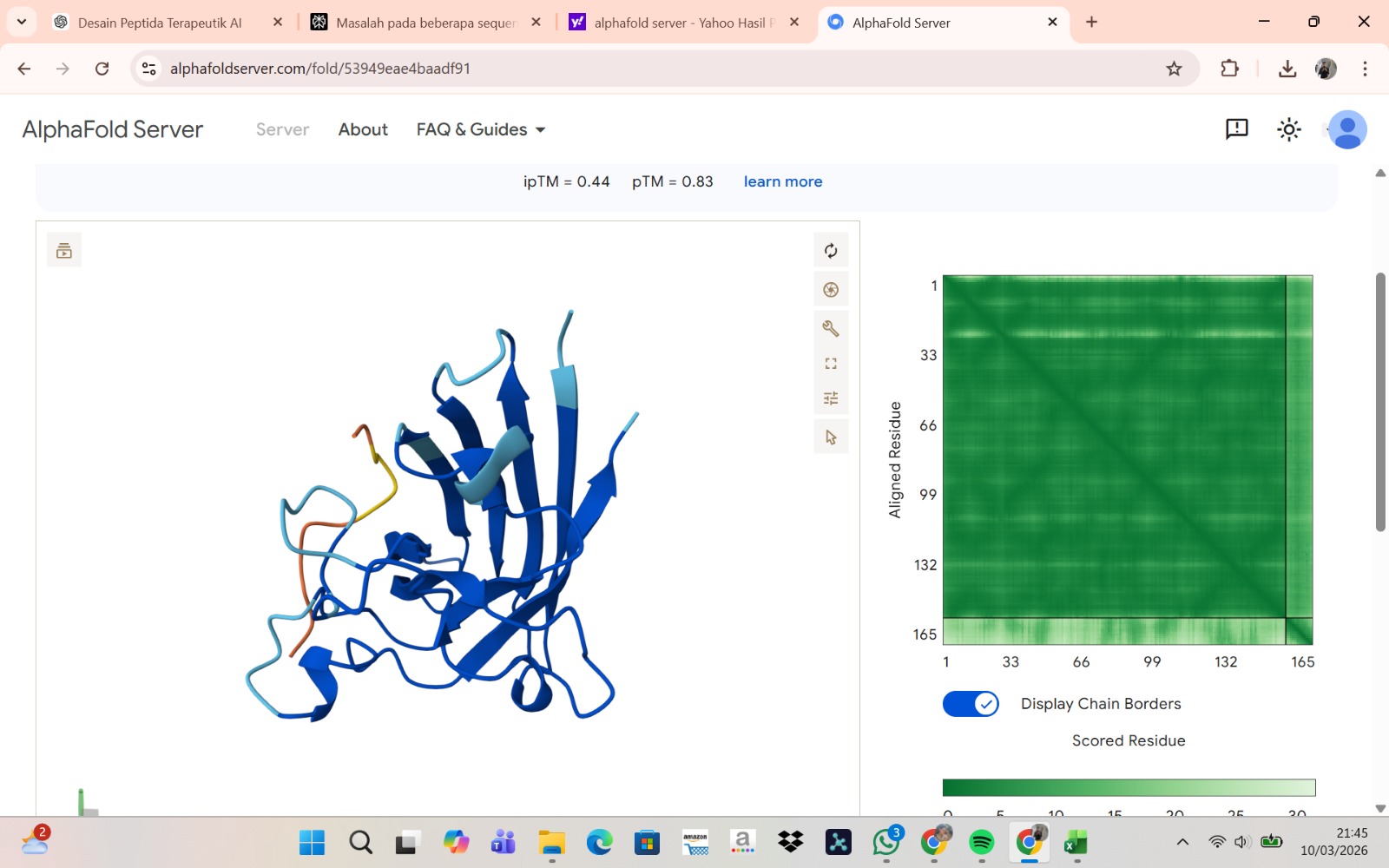

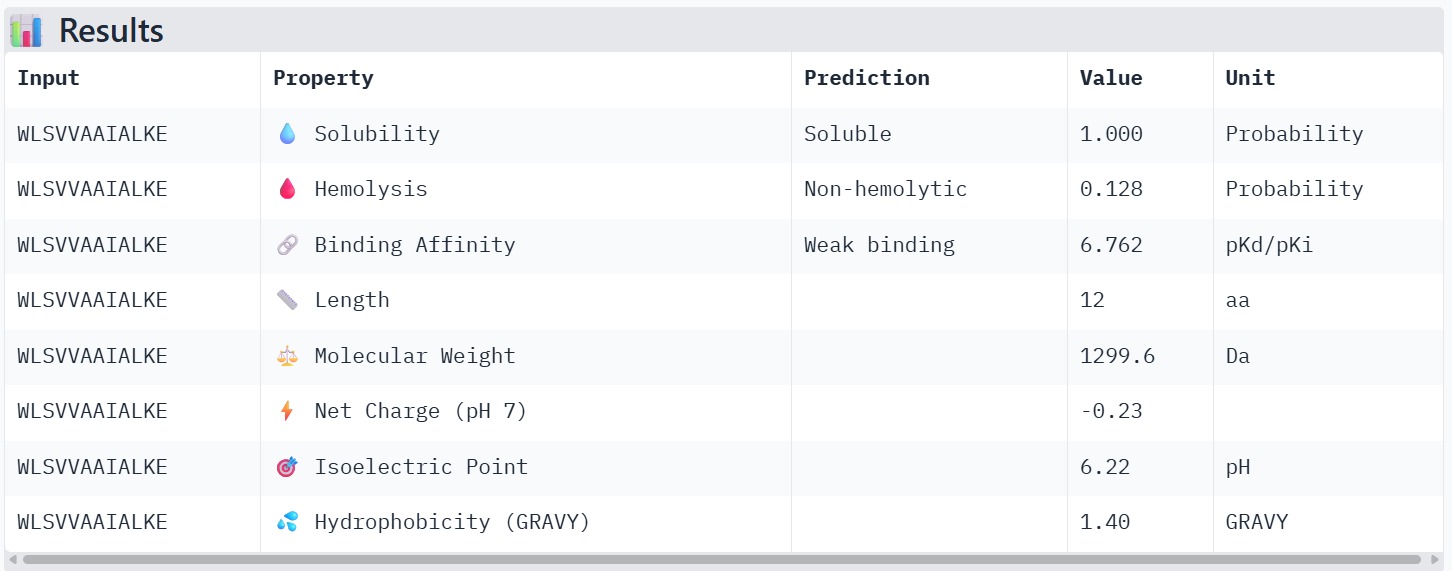
Peptide 3 : WLSVVAAIALKE
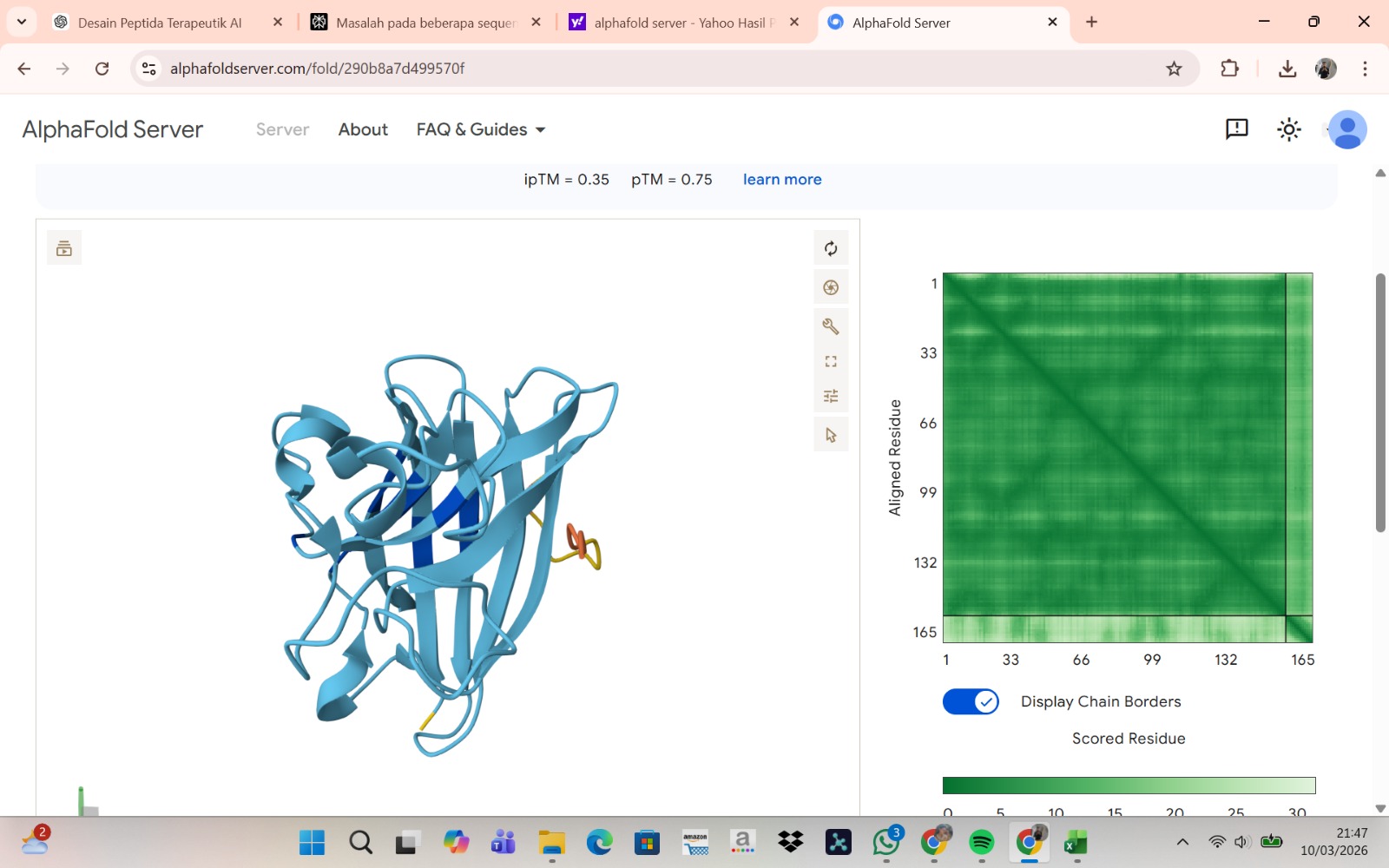

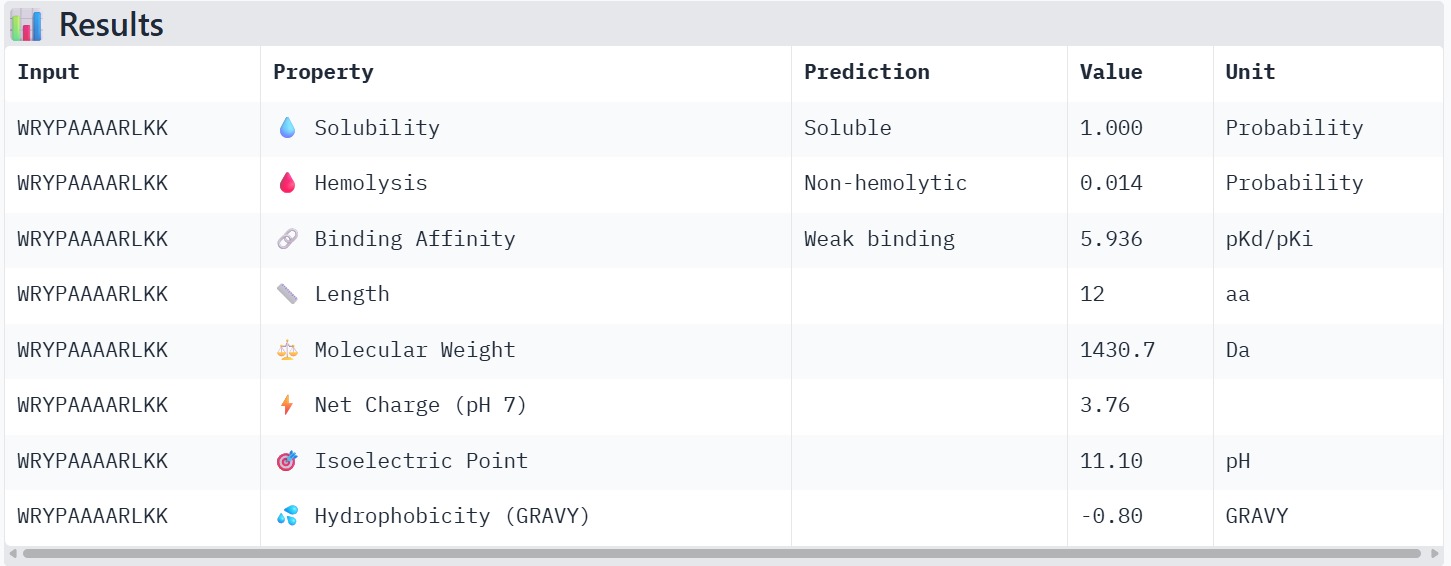
Peptide 4 : WRYPAAAARLKK
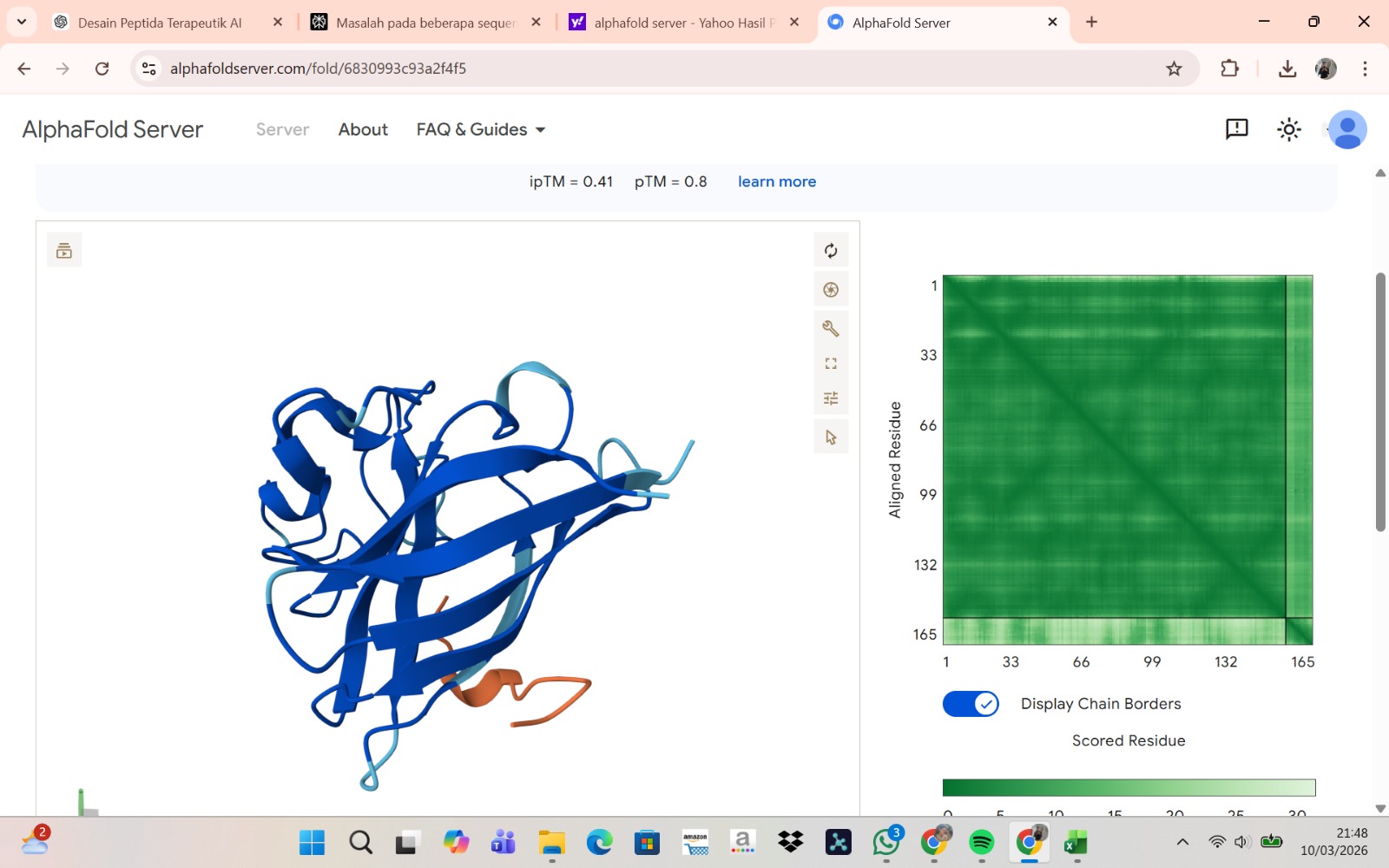

The peptide–protein complexes were modeled using AlphaFold3 by submitting the mutant SOD1 sequence as chain A and each peptide sequence as chain B. The predicted interface scores (ipTM) ranged from 0.35 to 0.44, indicating relatively weak but detectable interactions between the peptides and the SOD1 protein. Among the generated candidates, the peptide HLVPAVAIEHKK showed the highest ipTM value (0.44) and the highest pTM score (0.83), suggesting a more stable predicted complex compared to the other peptides. The remaining peptides displayed slightly lower ipTM scores, indicating weaker predicted interactions. Based on the predicted structures, the peptides appear to bind primarily to the surface of the SOD1 protein rather than being deeply buried within the structure. The interactions likely occur near exposed regions of the β-barrel or close to the N-terminal region where the A4V mutation is located. Although the binding confidence is modest, these results suggest that some of the PepMLM-generated peptides may interact with the mutant SOD1 surface and could serve as starting points for further optimization.
Part 3 Evaluate Properties with PeptiVerse
The therapeutic properties of the generated peptides were evaluated using PeptiVerse. All four peptides were predicted to be highly soluble with a probability of 1.000, indicating favorable physicochemical characteristics for biological applications. Hemolysis prediction suggested that all peptides are non-hemolytic, with WRYPAAAARLKK showing the lowest hemolysis probability (0.014). Binding affinity predictions ranged from pKd/pKi values of 5.27 to 6.85. Among the candidates, WRYYAYAIRWKK displayed the highest predicted binding affinity (6.847), suggesting stronger interaction with the A4V mutant SOD1 target. This peptide also exhibits a positive net charge (+3.76), which may enhance electrostatic interactions with the protein surface. Hydrophobicity analysis indicates that WRYYAYAIRWKK is moderately hydrophilic (GRAVY = −1.20), which supports solubility and structural stability. Overall, WRYYAYAIRWKK appears to provide the best balance between predicted binding affinity, solubility, and safety properties, making it the most promising candidate for further structural and experimental validation.